
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,970,254 – 18,970,398 |
| Length | 144 |
| Max. P | 0.878414 |

| Location | 18,970,254 – 18,970,358 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 87.30 |
| Mean single sequence MFE | -24.28 |
| Consensus MFE | -22.08 |
| Energy contribution | -22.20 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.878414 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18970254 104 + 22224390 AAAAAAUAUUUCGUCAUGUUUUUAGGUUGGCCGCAGGCAGUGCUCACUGUACCGCCACAAUGUUUAUCGUUUUGCAUUUUUUUUUUCUUUGUUUUCUUGCGGUU .((((((((......)))))))).....((((((((((.((((.....)))).)))..(((((..(....)..)))))...................))))))) ( -23.30) >DroSec_CAF1 21527 101 + 1 -AAUAAUAUUUCGUCAUGUUGUUAGGUUGGCCGCAGGCAGUGCUCACUGUACCGCCACAAUGUAUAUCGUUUUGCGUUUUUUUG--UUUUGUUUUCUUGCGGUU -((((((((......)))))))).....((((((((((.((((.....)))).)))..((((((........))))))......--...........))))))) ( -25.30) >DroSim_CAF1 6048 100 + 1 -AAUAAUAUUUCGUCAUGUUGUUAGGUUGGCCGCAGGCAGUGCUCACUGUACCGCCACAAUGUAUAUAGUUUUGCGUUUUUUU---UUUUGUUUUCUUGCGGUU -((((((((......)))))))).....((((((((((.((((.....)))).)))..((((((........)))))).....---...........))))))) ( -25.30) >DroEre_CAF1 11596 96 + 1 -AAUAAUAUUUCGUCAUGUUGUUAGGUUGGCCGAAGGCAGUGCUCACUGUACCGCCAAAGAGUUCAUCGUUUUACAUUACGUG-------UUUUUCUUGCGGUU -((((((((......)))))))).....(((((..(((.((((.....)))).))).(((((..((.(((........)))))-------...))))).))))) ( -23.20) >consensus _AAUAAUAUUUCGUCAUGUUGUUAGGUUGGCCGCAGGCAGUGCUCACUGUACCGCCACAAUGUAUAUCGUUUUGCAUUUUUUU___UUUUGUUUUCUUGCGGUU .((((((((......)))))))).....((((((((((.((((.....)))).)))..((((((........))))))...................))))))) (-22.08 = -22.20 + 0.12)


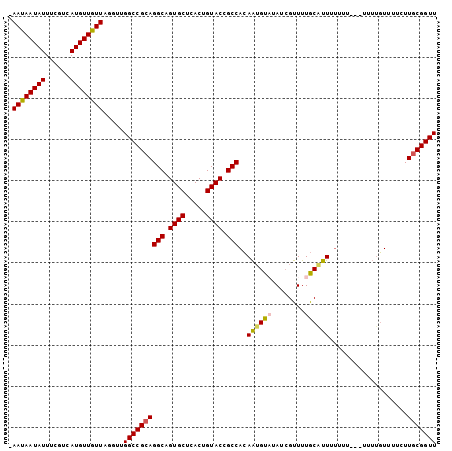
| Location | 18,970,278 – 18,970,398 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.31 |
| Mean single sequence MFE | -27.32 |
| Consensus MFE | -20.70 |
| Energy contribution | -20.66 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.804232 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
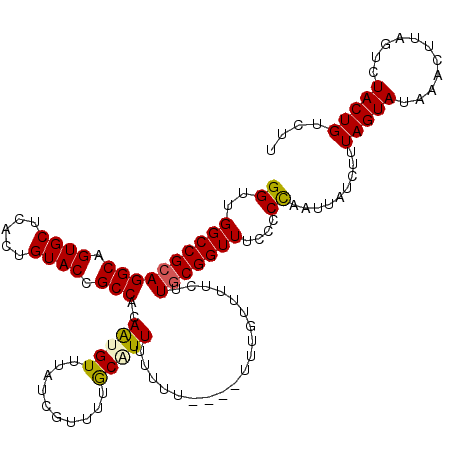
>X_DroMel_CAF1 18970278 120 + 22224390 GGUUGGCCGCAGGCAGUGCUCACUGUACCGCCACAAUGUUUAUCGUUUUGCAUUUUUUUUUUCUUUGUUUUCUUGCGGUUUCCCCUAAUUAUCUUUAGUAUAAACUUAGUCUACUGUCUU ((..((((((((((.((((.....)))).)))..(((((..(....)..)))))...................)))))))..))((((......))))...................... ( -26.30) >DroSec_CAF1 21550 117 + 1 GGUUGGCCGCAGGCAGUGCUCACUGUACCGCCACAAUGUAUAUCGUUUUGCGUUUUUUUG--UUUUGUUUUCUUGCGGUU-CCCCCAAUUAUCUUUAGUAUAAACUUAGUCUACUGUCUU ((..((((((((((.((((.....)))).)))..((((((........))))))......--...........)))))))-..))..........(((((...........))))).... ( -27.10) >DroSim_CAF1 6071 116 + 1 GGUUGGCCGCAGGCAGUGCUCACUGUACCGCCACAAUGUAUAUAGUUUUGCGUUUUUUU---UUUUGUUUUCUUGCGGUU-CACCCAAUUAUCUUUAGUAUAAACUUAGUCUACUGUCUU (((..(((((((((.((((.....)))).)))..((((((........)))))).....---...........)))))).-.)))..........(((((...........))))).... ( -28.40) >DroEre_CAF1 11619 112 + 1 GGUUGGCCGAAGGCAGUGCUCACUGUACCGCCAAAGAGUUCAUCGUUUUACAUUACGUG-------UUUUUCUUGCGGUUUCC-CCAAUUAUCUUUAGUAUAAACUUAGUCUACUGUCUU ((..(((((..(((.((((.....)))).))).(((((..((.(((........)))))-------...))))).)))))..)-)..........(((((...........))))).... ( -25.60) >DroYak_CAF1 13437 113 + 1 GGUUGGCCGCAGGCAGUGCUCACUGUACCGCCACAAAGUUUAUCGUUUUACAUUAUGUU-------CUUUUCGUGCGGUUUCCCCCAGUUAUCUUUAGUAUAAACUUAGUCUACUGUCUU ((..((((((((((.((((.....)))).)))..((((..(((.((.....)).)))..-------))))...)))))))..)).((((...((..(((....))).))...)))).... ( -29.20) >consensus GGUUGGCCGCAGGCAGUGCUCACUGUACCGCCACAAUGUUUAUCGUUUUGCAUUUUUUU____UUUGUUUUCUUGCGGUUUCCCCCAAUUAUCUUUAGUAUAAACUUAGUCUACUGUCUU ((..((((((((((.((((.....)))).)))..(((((..........)))))...................)))))))....)).........(((((...........))))).... (-20.70 = -20.66 + -0.04)

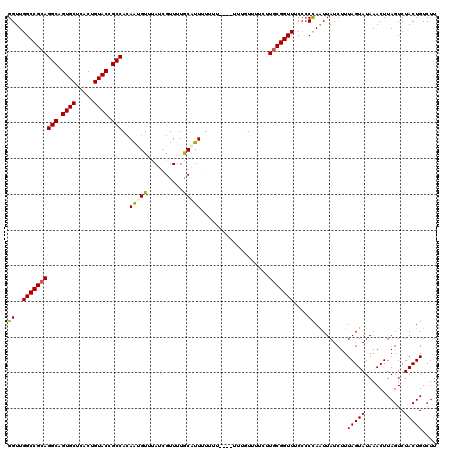
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:43:18 2006