
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,822,606 – 18,822,747 |
| Length | 141 |
| Max. P | 0.998666 |

| Location | 18,822,606 – 18,822,726 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.11 |
| Mean single sequence MFE | -35.03 |
| Consensus MFE | -32.76 |
| Energy contribution | -32.39 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.09 |
| Structure conservation index | 0.94 |
| SVM decision value | 2.48 |
| SVM RNA-class probability | 0.994483 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18822606 120 + 22224390 UCACUGUUUUCAUUAUCUGUGAUUUGCCCUUAUAGUCGCCUUGCCGGCAGCUCAUCCUUAAUUGCGAUUGUUGCAAGGACUUGCAGAAUUUUGCAUAUUUGAUGAGCCUGGCUCUUUCUC ..................((((((.........))))))...(((((..(((((((......(((((...((((((....))))))....))))).....))))))))))))........ ( -33.10) >DroSec_CAF1 59657 118 + 1 UCACUGUUUUCAUUAUCUGUGAUUUGC-CUUAUAGUCGCCUCGCCGGCAGCUCAUCCUUAAUUGCGAUUGUUGCAAGGACUUGCAGAAU-UUGCAUAUUUGAUGAGCCUGGCUCUCCCUC ..................((((((...-.....))))))...(((((..(((((((......(((((...((((((....))))))...-))))).....))))))))))))........ ( -34.90) >DroEre_CAF1 57588 115 + 1 UCACUGUUUUCAUUAUCUGUGAUUUGC-CUUAUUGCCGCCUCGCCGGCAGCUCAUCCUUAAUUGCGAUUGUUGCAAGGACUUGCAGAAUUUUGCAUAUUUGAUGAGCCUGGCUCU----C ((((..............))))...((-(....(((((......)))))(((((((......(((((...((((((....))))))....))))).....)))))))..)))...----. ( -35.44) >DroYak_CAF1 58566 115 + 1 UCACUGUUUUCAUUAUCUGUGAUUUGC-CUUGUAGUCGCCUCGCCGGCAGCUCAUCCUUAAUUGCAAUUGCUGCAAGGACUUGCAGAAUUUUGCAUAUUUGAUGAGCCUGGCUCU----C ..................((((((...-.....))))))...(((((..(((((((......(((((...((((((....))))))....))))).....))))))))))))...----. ( -36.70) >consensus UCACUGUUUUCAUUAUCUGUGAUUUGC_CUUAUAGUCGCCUCGCCGGCAGCUCAUCCUUAAUUGCGAUUGUUGCAAGGACUUGCAGAAUUUUGCAUAUUUGAUGAGCCUGGCUCU____C ((((..............))))....................(((((..(((((((......(((((...((((((....))))))....))))).....))))))))))))........ (-32.76 = -32.39 + -0.37)



| Location | 18,822,606 – 18,822,726 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.11 |
| Mean single sequence MFE | -33.02 |
| Consensus MFE | -27.26 |
| Energy contribution | -27.26 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.918035 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18822606 120 - 22224390 GAGAAAGAGCCAGGCUCAUCAAAUAUGCAAAAUUCUGCAAGUCCUUGCAACAAUCGCAAUUAAGGAUGAGCUGCCGGCAAGGCGACUAUAAGGGCAAAUCACAGAUAAUGAAAACAGUGA ........(((.((((((((.....(((.......(((((....)))))......)))......))))))))(((.....))).........)))...((((..............)))) ( -30.86) >DroSec_CAF1 59657 118 - 1 GAGGGAGAGCCAGGCUCAUCAAAUAUGCAA-AUUCUGCAAGUCCUUGCAACAAUCGCAAUUAAGGAUGAGCUGCCGGCGAGGCGACUAUAAG-GCAAAUCACAGAUAAUGAAAACAGUGA ........(((.((((((((.....(((..-....(((((....)))))......)))......))))))))(((.....)))........)-))...((((..............)))) ( -31.64) >DroEre_CAF1 57588 115 - 1 G----AGAGCCAGGCUCAUCAAAUAUGCAAAAUUCUGCAAGUCCUUGCAACAAUCGCAAUUAAGGAUGAGCUGCCGGCGAGGCGGCAAUAAG-GCAAAUCACAGAUAAUGAAAACAGUGA .----...(((.((((((((.....(((.......(((((....)))))......)))......))))))))((((......)))).....)-))...((((..............)))) ( -32.86) >DroYak_CAF1 58566 115 - 1 G----AGAGCCAGGCUCAUCAAAUAUGCAAAAUUCUGCAAGUCCUUGCAGCAAUUGCAAUUAAGGAUGAGCUGCCGGCGAGGCGACUACAAG-GCAAAUCACAGAUAAUGAAAACAGUGA .----...(((.((((((((.....(((((....((((((....))))))...)))))......))))))))(((.....)))........)-))...((((..............)))) ( -36.74) >consensus G____AGAGCCAGGCUCAUCAAAUAUGCAAAAUUCUGCAAGUCCUUGCAACAAUCGCAAUUAAGGAUGAGCUGCCGGCGAGGCGACUAUAAG_GCAAAUCACAGAUAAUGAAAACAGUGA ........(((.((((((((.....(((.......(((((....)))))......)))......))))))))(((.....)))........).))...((((..............)))) (-27.26 = -27.26 + -0.00)

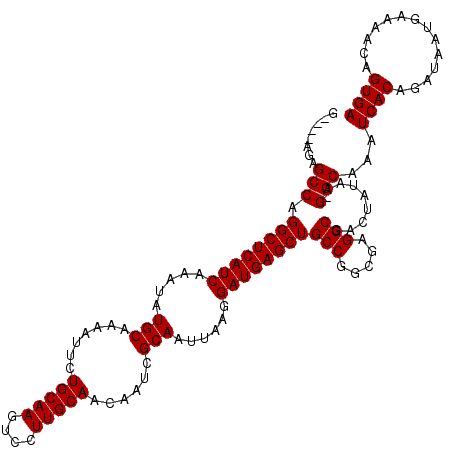
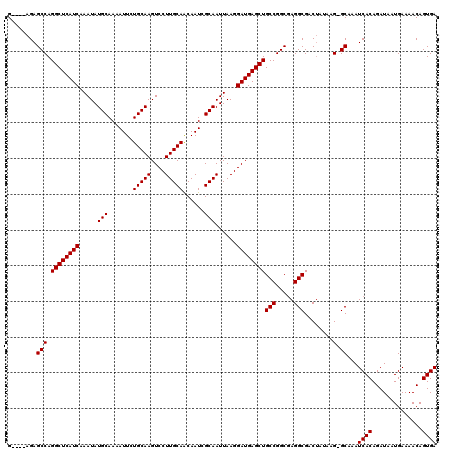
| Location | 18,822,646 – 18,822,747 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 90.83 |
| Mean single sequence MFE | -33.45 |
| Consensus MFE | -32.22 |
| Energy contribution | -31.79 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.15 |
| Mean z-score | -3.71 |
| Structure conservation index | 0.96 |
| SVM decision value | 3.18 |
| SVM RNA-class probability | 0.998666 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18822646 101 + 22224390 UUGCCGGCAGCUCAUCCUUAAUUGCGAUUGUUGCAAGGACUUGCAGAAUUUUGCAUAUUUGAUGAGCCUGGCUCUUUCUC---ACUCCUCUGCCUGCCUGCAUA .(((.(((((((((((......(((((...((((((....))))))....))))).....)))))))..(((........---........))))))).))).. ( -34.89) >DroSec_CAF1 59696 103 + 1 UCGCCGGCAGCUCAUCCUUAAUUGCGAUUGUUGCAAGGACUUGCAGAAU-UUGCAUAUUUGAUGAGCCUGGCUCUCCCUCUCAACUCCUCCGCCUGCCUGCAUA ..((.(((((((((((......(((((...((((((....))))))...-))))).....)))))))..(((...................))))))).))... ( -34.41) >DroEre_CAF1 57627 97 + 1 UCGCCGGCAGCUCAUCCUUAAUUGCGAUUGUUGCAAGGACUUGCAGAAUUUUGCAUAUUUGAUGAGCCUGGCUCU----C---AAUCCUCUGCCCGCAUGCAUA ..((.(((((((((((......(((((...((((((....))))))....))))).....)))))))..((....----.---...))..)))).))....... ( -31.00) >DroYak_CAF1 58605 97 + 1 UCGCCGGCAGCUCAUCCUUAAUUGCAAUUGCUGCAAGGACUUGCAGAAUUUUGCAUAUUUGAUGAGCCUGGCUCU----C---ACUCCUCUGCCUGUUUGCAUA ..(((((..(((((((......(((((...((((((....))))))....))))).....))))))))))))...----.---.......(((......))).. ( -33.50) >consensus UCGCCGGCAGCUCAUCCUUAAUUGCGAUUGUUGCAAGGACUUGCAGAAUUUUGCAUAUUUGAUGAGCCUGGCUCU____C___ACUCCUCUGCCUGCCUGCAUA ..((.(((.(((((((......(((((...((((((....))))))....))))).....)))))))..(((...................))).))).))... (-32.22 = -31.79 + -0.44)



| Location | 18,822,646 – 18,822,747 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 90.83 |
| Mean single sequence MFE | -32.18 |
| Consensus MFE | -27.62 |
| Energy contribution | -28.18 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.89 |
| SVM RNA-class probability | 0.981714 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18822646 101 - 22224390 UAUGCAGGCAGGCAGAGGAGU---GAGAAAGAGCCAGGCUCAUCAAAUAUGCAAAAUUCUGCAAGUCCUUGCAACAAUCGCAAUUAAGGAUGAGCUGCCGGCAA ..(((.(((.(((........---........))).((((((((.....(((.......(((((....)))))......)))......))))))))))).))). ( -31.61) >DroSec_CAF1 59696 103 - 1 UAUGCAGGCAGGCGGAGGAGUUGAGAGGGAGAGCCAGGCUCAUCAAAUAUGCAA-AUUCUGCAAGUCCUUGCAACAAUCGCAAUUAAGGAUGAGCUGCCGGCGA ..(((.(((((((((((...(((.....((((((...)))).)).......)))-.))))))..((((((((.......)))....)))))...))))).))). ( -33.10) >DroEre_CAF1 57627 97 - 1 UAUGCAUGCGGGCAGAGGAUU---G----AGAGCCAGGCUCAUCAAAUAUGCAAAAUUCUGCAAGUCCUUGCAACAAUCGCAAUUAAGGAUGAGCUGCCGGCGA ......(((.(((...((...---.----....)).((((((((.....(((.......(((((....)))))......)))......))))))))))).))). ( -30.52) >DroYak_CAF1 58605 97 - 1 UAUGCAAACAGGCAGAGGAGU---G----AGAGCCAGGCUCAUCAAAUAUGCAAAAUUCUGCAAGUCCUUGCAGCAAUUGCAAUUAAGGAUGAGCUGCCGGCGA ..(((.....(((....(.((---.----...))).((((((((.....(((((....((((((....))))))...)))))......))))))))))).))). ( -33.50) >consensus UAUGCAGGCAGGCAGAGGAGU___G____AGAGCCAGGCUCAUCAAAUAUGCAAAAUUCUGCAAGUCCUUGCAACAAUCGCAAUUAAGGAUGAGCUGCCGGCGA ..(((.(((.(((...................))).((((((((.....(((.......(((((....)))))......)))......))))))))))).))). (-27.62 = -28.18 + 0.56)

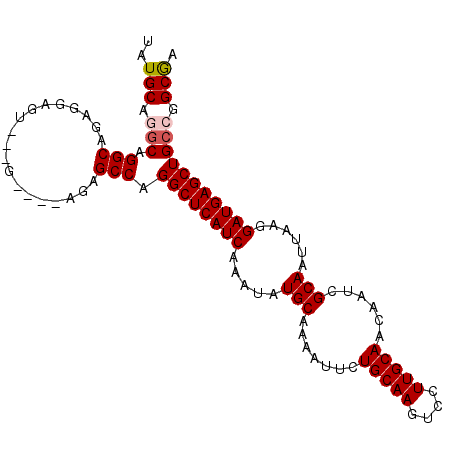

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:41:47 2006