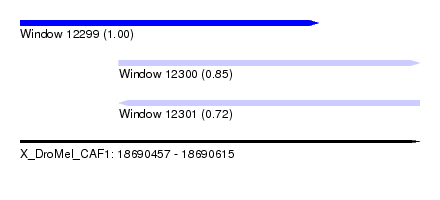
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,690,457 – 18,690,615 |
| Length | 158 |
| Max. P | 0.999970 |

| Location | 18,690,457 – 18,690,575 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.33 |
| Mean single sequence MFE | -30.48 |
| Consensus MFE | -30.52 |
| Energy contribution | -29.92 |
| Covariance contribution | -0.60 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.51 |
| Structure conservation index | 1.00 |
| SVM decision value | 5.04 |
| SVM RNA-class probability | 0.999970 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18690457 118 + 22224390 UAUGCUAAGUUUAAUGCUGAAGGUCAAUUAGAGU-GGGCUACCAGUUGCCCUUUCUUACAAUCUGAUCAUGAUAAUCGUUAUCAUAGUUUAACGAUU-CGCUGAUCGUGAUCUUUCAAAA .......(((.....)))((((((((...((((.-((((........)))).))))........(((((.(..((((((((........))))))))-..)))))).))))))))..... ( -31.20) >DroSec_CAF1 22790 119 + 1 -AUGCUACAUUUAAUGCUGAAGGUCAAUUAGAGUUGGGCUUCCAGUUGCCUUUUCUUAUAAUCUGAUCAUGAUAAUCGUUAUCAUAGUUUAACGAUUCCGGUGAUCGUGAUCUUUUAAAA -..((..........)).((((((((....(((..((((........))))...))).......((((((...((((((((........))))))))...)))))).))))))))..... ( -29.40) >DroSim_CAF1 5058 119 + 1 -AUGCUACAUUUAAUGCUGAAGGUCAAUUAGAGUUGGGCUUCCAGUUGCCUUUUCUUAUAAUCUGAUCAUGAUAAUCGUUAUCAUAGUUUAACGAUUCCGGUGAUCGUGAUCUUUCAAAA -..((..........)).((((((((....(((..((((........))))...))).......((((((...((((((((........))))))))...)))))).))))))))..... ( -29.40) >DroEre_CAF1 11864 117 + 1 -AUGCUGAAAUGAAUACUGAAGGUCAAUUAGAGU-GGGUUUUCAGUUGCCCUUUCUUAUAAUCUGAUCAUAAUAAUCGUUAUCAUAGUUUAACGAUU-CGGUGAUCGUGAUCUUCCAAAA -.................((((((((...((((.-((((........)))).))))........((((((...((((((((........))))))))-..)))))).))))))))..... ( -32.80) >DroYak_CAF1 10069 117 + 1 -AUGCUAAAAUAAAUACUGAAGGUCAAUUAGAGU-GGGUUUUCAGUUGCCCUUUCUUAUAAUUUGAUCAUGAUAGUCGUUAUCACAGUUUAACGAUU-CGGUGAUCGUGAUCUUUCAAAA -.................((((((((...((((.-((((........)))).))))........((((((...((((((((........))))))))-..)))))).))))))))..... ( -29.60) >consensus _AUGCUAAAUUUAAUGCUGAAGGUCAAUUAGAGU_GGGCUUCCAGUUGCCCUUUCUUAUAAUCUGAUCAUGAUAAUCGUUAUCAUAGUUUAACGAUU_CGGUGAUCGUGAUCUUUCAAAA ..................((((((((....(((..((((........))))...))).......((((((...((((((((........))))))))...)))))).))))))))..... (-30.52 = -29.92 + -0.60)



| Location | 18,690,496 – 18,690,615 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.12 |
| Mean single sequence MFE | -27.41 |
| Consensus MFE | -19.50 |
| Energy contribution | -19.98 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.847842 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18690496 119 + 22224390 ACCAGUUGCCCUUUCUUACAAUCUGAUCAUGAUAAUCGUUAUCAUAGUUUAACGAUU-CGCUGAUCGUGAUCUUUCAAAAAGCAAGCAGAACUUGUUUUUGAUAAGAACGAGAGUUCCGC ....(..((.((((((((......(((((((((((((((((........))))))))-.....)))))))))..(((((((.((((.....))))))))))))))))..))).))..).. ( -30.70) >DroSec_CAF1 22829 120 + 1 UCCAGUUGCCUUUUCUUAUAAUCUGAUCAUGAUAAUCGUUAUCAUAGUUUAACGAUUCCGGUGAUCGUGAUCUUUUAAAAACCAAACAGAACUUGUUUUUGAUAAGAACGAGAGUUCCGU ....(..((((((((((((.....(((((((((.((((..(((..........)))..)))).)))))))))...(((((((............)))))))))))))).))).))..).. ( -27.80) >DroSim_CAF1 5097 120 + 1 UCCAGUUGCCUUUUCUUAUAAUCUGAUCAUGAUAAUCGUUAUCAUAGUUUAACGAUUCCGGUGAUCGUGAUCUUUCAAAAACCAAACAGAACUUGUUUUUGAUAAGAACGAGAGUUCCGU ....(..(((((((((((......(((((((((.((((..(((..........)))..)))).)))))))))..((((((((............)))))))))))))).))).))..).. ( -29.80) >DroEre_CAF1 11902 118 + 1 UUCAGUUGCCCUUUCUUAUAAUCUGAUCAUAAUAAUCGUUAUCAUAGUUUAACGAUU-CGGUGAUCGUGAUCUUCCAAAAAGCAA-CAGAACCAACUUUUGAUAAGAACGACAGUAUCGU ....((((....(((((((.((((((((((...((((((((........))))))))-..))))))).)))..............-(((((.....))))))))))))))))........ ( -25.10) >DroYak_CAF1 10107 106 + 1 UUCAGUUGCCCUUUCUUAUAAUUUGAUCAUGAUAGUCGUUAUCACAGUUUAACGAUU-CGGUGAUCGUGAUCUUUCAAAAAACAA-GGGACCUAAUUUUUGAUAAGCA------------ ...((...(((((...........(((((((((((((((((........))))))))-.....)))))))))...........))-)))..))...............------------ ( -23.65) >consensus UCCAGUUGCCCUUUCUUAUAAUCUGAUCAUGAUAAUCGUUAUCAUAGUUUAACGAUU_CGGUGAUCGUGAUCUUUCAAAAACCAA_CAGAACUUGUUUUUGAUAAGAACGAGAGUUCCGU ............(((((((.....(((((((((((((((((........))))))))......)))))))))...((((((..............)))))))))))))............ (-19.50 = -19.98 + 0.48)


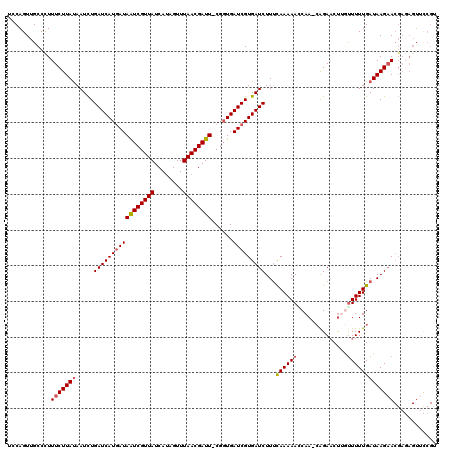
| Location | 18,690,496 – 18,690,615 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.12 |
| Mean single sequence MFE | -26.56 |
| Consensus MFE | -20.16 |
| Energy contribution | -20.52 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724667 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18690496 119 - 22224390 GCGGAACUCUCGUUCUUAUCAAAAACAAGUUCUGCUUGCUUUUUGAAAGAUCACGAUCAGCG-AAUCGUUAAACUAUGAUAACGAUUAUCAUGAUCAGAUUGUAAGAAAGGGCAACUGGU ((((((((...(((.........))).))))))))(((((((((....((((..(((((...-((((((((........))))))))....))))).)))).....)))))))))..... ( -32.10) >DroSec_CAF1 22829 120 - 1 ACGGAACUCUCGUUCUUAUCAAAAACAAGUUCUGUUUGGUUUUUAAAAGAUCACGAUCACCGGAAUCGUUAAACUAUGAUAACGAUUAUCAUGAUCAGAUUAUAAGAAAAGGCAACUGGA ((((((((...(((.........))).))))))))(((.(((((....((((..(((((....((((((((........))))))))....))))).)))).....))))).)))..... ( -24.60) >DroSim_CAF1 5097 120 - 1 ACGGAACUCUCGUUCUUAUCAAAAACAAGUUCUGUUUGGUUUUUGAAAGAUCACGAUCACCGGAAUCGUUAAACUAUGAUAACGAUUAUCAUGAUCAGAUUAUAAGAAAAGGCAACUGGA .(((....((..((((((((((((((.((......)).))))))))..((((..(((((....((((((((........))))))))....))))).)))).))))))..))...))).. ( -27.50) >DroEre_CAF1 11902 118 - 1 ACGAUACUGUCGUUCUUAUCAAAAGUUGGUUCUG-UUGCUUUUUGGAAGAUCACGAUCACCG-AAUCGUUAAACUAUGAUAACGAUUAUUAUGAUCAGAUUAUAAGAAAGGGCAACUGAA (((((...)))))................(((.(-(((((((((..(.((((..(((((...-((((((((........))))))))....))))).)))).)...)))))))))).))) ( -27.20) >DroYak_CAF1 10107 106 - 1 ------------UGCUUAUCAAAAAUUAGGUCCC-UUGUUUUUUGAAAGAUCACGAUCACCG-AAUCGUUAAACUGUGAUAACGACUAUCAUGAUCAAAUUAUAAGAAAGGGCAACUGAA ------------(((((.(((((((..((....)-)...)))))))..(((((((((.....-.)))))......((((((.....)))))))))).............)))))...... ( -21.40) >consensus ACGGAACUCUCGUUCUUAUCAAAAACAAGUUCUG_UUGCUUUUUGAAAGAUCACGAUCACCG_AAUCGUUAAACUAUGAUAACGAUUAUCAUGAUCAGAUUAUAAGAAAGGGCAACUGGA ............(((((((((((((((((.....)))).)))))))........(((((....((((((((........))))))))....)))))......)))))).((....))... (-20.16 = -20.52 + 0.36)


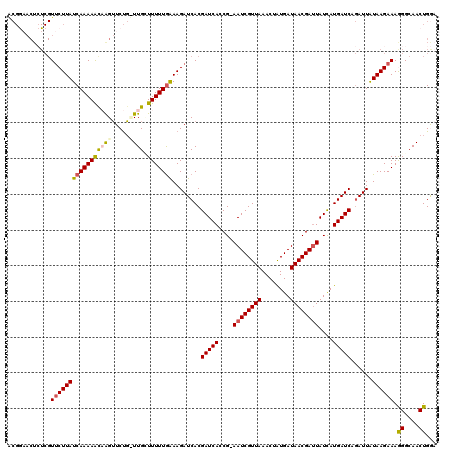
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:40:26 2006