
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,653,478 – 18,653,582 |
| Length | 104 |
| Max. P | 0.847538 |

| Location | 18,653,478 – 18,653,582 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 88.08 |
| Mean single sequence MFE | -29.52 |
| Consensus MFE | -23.81 |
| Energy contribution | -24.12 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.81 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.529399 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18653478 104 + 22224390 UUGUUCAAGUUGGCCCAGAAAAACGUAUUUCCGUUUAAAUUCCGGGUUAAGGUGGGAGUUACCGCCAGCGGCACUUGACUGCUGAAGGCACUUAAAAAAGAUGU ...(((...(((((((.((((((((......)))))...))).)))))))(((((......)))))((((((....).)))))))).((((((....))).))) ( -28.70) >DroSec_CAF1 12490 103 + 1 UUGUUCAAUUUGGCCCAGAAAAACGUAUUUCCGUUUAAAUUCCGGGUUAAGGUGGGAGUUCCCGCCAGCAGCACUUGACUGUUGAAGGCACUUAA-AAAGAUGU .(.(((...(((((((.((((((((......)))))...))).)))))))((((((....))))))((((((....).)))))))).).......-........ ( -29.10) >DroSim_CAF1 12442 103 + 1 UUGUUCAAUUUGGCCCAGAAAAACGUAUUUCCGUUUAAAUUCCGGGUUAAGGUGGGAGUUCCCGCCAGCAGCACUUGACUGCUGAAGGCACUUAA-AAAGAUGU .(.(((...(((((((.((((((((......)))))...))).)))))))((((((....))))))((((((....).)))))))).).......-........ ( -31.50) >DroEre_CAF1 11164 100 + 1 UGGUUCAGCUUGGCCCAGAAAAACGUAUUUCCGCUUAAGUUCCGGGCUAAGGUGGGCGUUACAGACU---GCGCCUGACUGCUAAAAUAACUUAA-AAAGGUGU ..(((((.((((((((.(((....((......)).....))).)))))))).)))))..........---((((((...................-..)))))) ( -28.80) >consensus UUGUUCAAUUUGGCCCAGAAAAACGUAUUUCCGUUUAAAUUCCGGGUUAAGGUGGGAGUUACCGCCAGCAGCACUUGACUGCUGAAGGCACUUAA_AAAGAUGU ...((((.((((((((.((((((((......)))))...))).)))))))).)))).......(((..(((((......)))))..)))............... (-23.81 = -24.12 + 0.31)

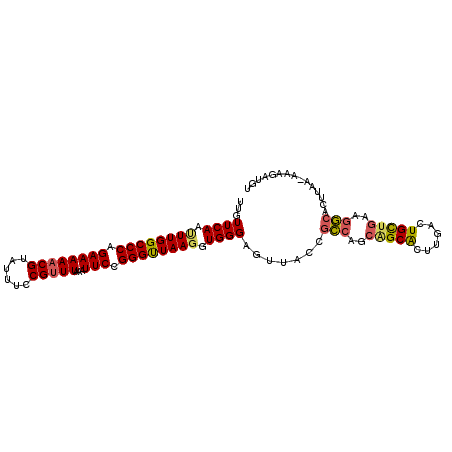

| Location | 18,653,478 – 18,653,582 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 88.08 |
| Mean single sequence MFE | -29.08 |
| Consensus MFE | -20.49 |
| Energy contribution | -21.55 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.847538 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18653478 104 - 22224390 ACAUCUUUUUUAAGUGCCUUCAGCAGUCAAGUGCCGCUGGCGGUAACUCCCACCUUAACCCGGAAUUUAAACGGAAAUACGUUUUUCUGGGCCAACUUGAACAA ........((((((((((.(((((.((.....)).))))).)))..............(((((((...(((((......))))))))))))...)))))))... ( -28.70) >DroSec_CAF1 12490 103 - 1 ACAUCUUU-UUAAGUGCCUUCAACAGUCAAGUGCUGCUGGCGGGAACUCCCACCUUAACCCGGAAUUUAAACGGAAAUACGUUUUUCUGGGCCAAAUUGAACAA ........-(((((.(((.....((((.....))))..)))(((....)))..)))))(((((((...(((((......))))))))))))............. ( -26.80) >DroSim_CAF1 12442 103 - 1 ACAUCUUU-UUAAGUGCCUUCAGCAGUCAAGUGCUGCUGGCGGGAACUCCCACCUUAACCCGGAAUUUAAACGGAAAUACGUUUUUCUGGGCCAAAUUGAACAA ........-(((((.(((..(((((......)))))..)))(((....)))..)))))(((((((...(((((......))))))))))))............. ( -32.90) >DroEre_CAF1 11164 100 - 1 ACACCUUU-UUAAGUUAUUUUAGCAGUCAGGCGC---AGUCUGUAACGCCCACCUUAGCCCGGAACUUAAGCGGAAAUACGUUUUUCUGGGCCAAGCUGAACCA ........-.........((((((.....((((.---.........)))).......((((((((...(((((......)))))))))))))...))))))... ( -27.90) >consensus ACAUCUUU_UUAAGUGCCUUCAGCAGUCAAGUGCUGCUGGCGGGAACUCCCACCUUAACCCGGAAUUUAAACGGAAAUACGUUUUUCUGGGCCAAAUUGAACAA .........(((((.(((..((((........))))..)))(((....)))..)))))(((((((...(((((......))))))))))))............. (-20.49 = -21.55 + 1.06)


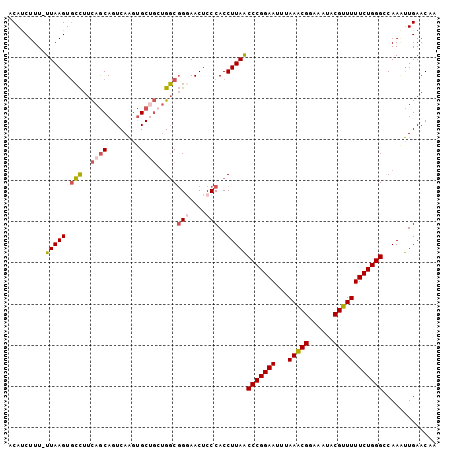
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:39:47 2006