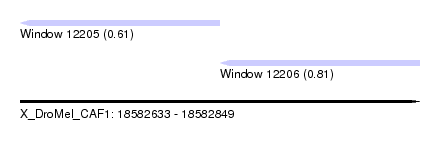
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,582,633 – 18,582,849 |
| Length | 216 |
| Max. P | 0.813149 |

| Location | 18,582,633 – 18,582,741 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.70 |
| Mean single sequence MFE | -26.58 |
| Consensus MFE | -18.26 |
| Energy contribution | -18.89 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.606519 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18582633 108 - 22224390 AAAAAAAAAAAAGGAUAAACCGGCAGCACUUGCGAGCUUCCUUCUGAGCUUUGUUGUCGUUGAAGUGCCAUGUCCCUU-CGUA-UAUCCAUGCCCCAGC----------CCACCACCACC ............(((((.((((((((((.....((((((......))))))))))))))..((((.(......).)))-))).-)))))..........----------........... ( -23.10) >DroSec_CAF1 47459 107 - 1 AA--AAAAAAAUGGAUAAACCGGCAGCACUUGCGAGCUUCCUUCUGAGCUUUGUUGUCGUUUAAGUGCCAUGUCCCUU-CAUGGCAUACAUGGCCCAAC----------CCACCACCACC ..--.......(((.(((((.(((((((.....((((((......)))))))))))))))))).((((((((......-))))))))...(((......----------))))))..... ( -31.00) >DroSim_CAF1 45232 109 - 1 AAAAAAAAAAAUGGAUAAACCGGCAGCACUUGCGAGCUUCCUUCUGAGCUUUGUUGUCGUUGAAGUGCCAUGUCCCUU-CAUGGCAUACAUGGCCCAAC----------CCACCACCACC ...........(((......((((((((.....((((((......)))))))))))))).....((((((((......-))))))))...(((......----------))))))..... ( -29.80) >DroYak_CAF1 39443 110 - 1 ----AACGAAAAGGAUAAACUGGCAGCACUUGCGAGAGUCCUUCUGAGCUCGGUUGUCGCUCAAGUGGCAUGCCCCAUUCC------UCAUUCCCCAACCCAACAACACCCACCACCACC ----...(((.((((......(((((((((((((((((..(....)..))).....))))..)))).)).))))....)))------)..)))........................... ( -22.40) >consensus AA__AAAAAAAAGGAUAAACCGGCAGCACUUGCGAGCUUCCUUCUGAGCUUUGUUGUCGUUGAAGUGCCAUGUCCCUU_CAUG_CAUACAUGCCCCAAC__________CCACCACCACC ............((......((((((((.....((((((......)))))))))))))).....(.((((((................)))))).)................))...... (-18.26 = -18.89 + 0.63)


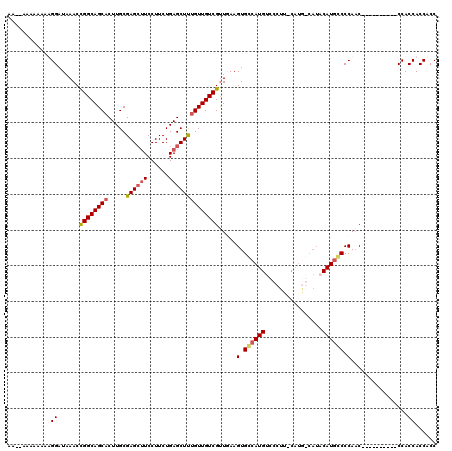
| Location | 18,582,741 – 18,582,849 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 86.41 |
| Mean single sequence MFE | -23.80 |
| Consensus MFE | -17.14 |
| Energy contribution | -17.07 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.813149 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
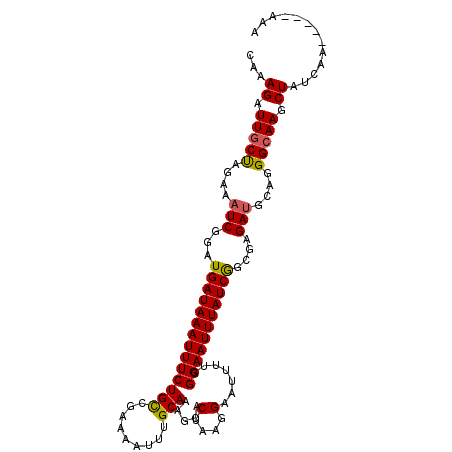
>X_DroMel_CAF1 18582741 108 - 22224390 CAAAGAUUGCUAGAAAUCGGAUGAUAAAUUUCUGCCGAAAAUUUGCAAAACACUAAGGAAUUUUGGGAAUUUAUCGGCGAGAUGCAGGGCAAGCUAUACAAAAAAAAA ...((.(((((....((((((........)))((((((((((((.(((((..(....)..))))).)))))).)))))).)))....))))).))............. ( -24.70) >DroSec_CAF1 47566 103 - 1 CAAAGAUUGCUAGAAAUCGGAUGAUAAAUUUCUGCCGAAAAUUGGCAAAGCACUAAGGAAUUUUUGGAAUUUAUCGGCGAGAUGCAGGGCAAGCUAUCAA-----AAA ...((.(((((....(((...(((((((((((((((((...)))))).....(....).......)))))))))))....)))....))))).)).....-----... ( -23.70) >DroSim_CAF1 45341 103 - 1 CAAAGAUUGCUAGAAAUCGGAUGAUAAAUUUCUGCCGAAAAUUUGCAAAGCACUAAGGAAUUUUUGGAAUUUAUCAGCGAGAUGCAGGGCAAGCUAUCAA-----AAA ...((.(((((....(((...(((((((((....((((((((((.............)))))))))))))))))))....)))....))))).)).....-----... ( -22.92) >DroYak_CAF1 39553 96 - 1 CAAAGAUUCCCAGAAAUCGGAUGAUAAAUUUCUGUCGACAAUUUGCAAAGCACUUUGGAGUUUCGGGAAUUUAUCGAAGAGAGGCAGGGCAAGCUA------------ ...((.(((((.(...((...(((((((((((((.(..(((..(((...)))..)))..)...)))))))))))))....))..).))).)).)).------------ ( -23.90) >consensus CAAAGAUUGCUAGAAAUCGGAUGAUAAAUUUCUGCCGAAAAUUUGCAAAGCACUAAGGAAUUUUGGGAAUUUAUCGGCGAGAUGCAGGGCAAGCUAUCAA_____AAA ...((.(((((....(((...((((((((((((((.........))).....(....).......)))))))))))....)))....))))).))............. (-17.14 = -17.07 + -0.06)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:39:02 2006