
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,538,628 – 18,538,771 |
| Length | 143 |
| Max. P | 0.996353 |

| Location | 18,538,628 – 18,538,731 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 96.73 |
| Mean single sequence MFE | -19.67 |
| Consensus MFE | -19.29 |
| Energy contribution | -19.07 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.935817 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18538628 103 + 22224390 AAGAAAAUAAAAUACAAUUAAAUAAUGUUGAAAAAAAAAAGUCGGCAUCACAAUCGAAAUCUGGUCAAGGUCGCACUGCCUUCAAGCAGCCCUAUGUGACCAC ........................(((((((..........)))))))....................(((((((((((......)))).....))))))).. ( -18.20) >DroSec_CAF1 3559 100 + 1 AAGAAAAUAAAAUACAAUUAAAUAAUGUCGAAA---AAAAGUCGGCAACACAAUCGAAAUCUGGUCAAGGUCGCACUGCCUUCAAGCAGCCCUAUGUGACCAC .........................((((((..---.....)))))).....................(((((((((((......)))).....))))))).. ( -20.40) >DroSim_CAF1 3607 100 + 1 AAGAAAAUAAAAUACAAUUAAAUAAUGUCGAAA---AAAAGUCGGCAACACAAUCGAAAUCUGGUCAAGGUCGCACUGCCUUCAAGCAGCCCUAUGUGACCAC .........................((((((..---.....)))))).....................(((((((((((......)))).....))))))).. ( -20.40) >consensus AAGAAAAUAAAAUACAAUUAAAUAAUGUCGAAA___AAAAGUCGGCAACACAAUCGAAAUCUGGUCAAGGUCGCACUGCCUUCAAGCAGCCCUAUGUGACCAC .........................((((((..........)))))).....................(((((((((((......)))).....))))))).. (-19.29 = -19.07 + -0.22)


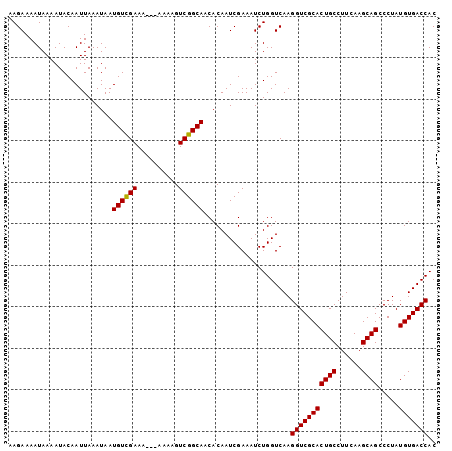
| Location | 18,538,651 – 18,538,771 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.19 |
| Mean single sequence MFE | -26.93 |
| Consensus MFE | -26.09 |
| Energy contribution | -25.87 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.69 |
| SVM RNA-class probability | 0.996353 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18538651 120 + 22224390 AAUGUUGAAAAAAAAAAGUCGGCAUCACAAUCGAAAUCUGGUCAAGGUCGCACUGCCUUCAAGCAGCCCUAUGUGACCACCAGACACCCAAUUACAGAUACAGAUACAGAUACAGAUACA .(((((((..........)))))))...........((((((...(((((((((((......)))).....))))))))))))).................................... ( -25.00) >DroSec_CAF1 3582 97 + 1 AAUGUCGAAA---AAAAGUCGGCAACACAAUCGAAAUCUGGUCAAGGUCGCACUGCCUUCAAGCAGCCCUAUGUGACCACCAGACACCCAAUUACAGGUA-------------------- ..((((((..---.....))))))............((((((...(((((((((((......)))).....))))))))))))).(((........))).-------------------- ( -28.60) >DroSim_CAF1 3630 111 + 1 AAUGUCGAAA---AAAAGUCGGCAACACAAUCGAAAUCUGGUCAAGGUCGCACUGCCUUCAAGCAGCCCUAUGUGACCACCAGACACCCAAUUACAGAUACAGAUACA------GAUACA ..((((((..---.....))))))............((((((...(((((((((((......)))).....)))))))))))))........................------...... ( -27.20) >consensus AAUGUCGAAA___AAAAGUCGGCAACACAAUCGAAAUCUGGUCAAGGUCGCACUGCCUUCAAGCAGCCCUAUGUGACCACCAGACACCCAAUUACAGAUACAGAUACA______GAUACA ..((((((..........))))))............((((((...(((((((((((......)))).....))))))))))))).................................... (-26.09 = -25.87 + -0.22)


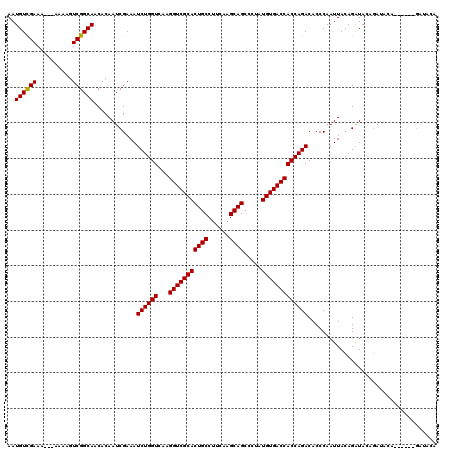
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:38:41 2006