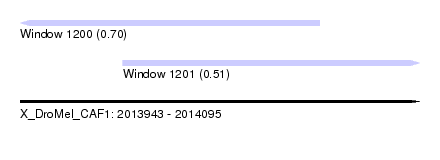

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,013,943 – 2,014,095 |
| Length | 152 |
| Max. P | 0.696314 |

| Location | 2,013,943 – 2,014,057 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.94 |
| Mean single sequence MFE | -37.15 |
| Consensus MFE | -22.53 |
| Energy contribution | -21.22 |
| Covariance contribution | -1.31 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.696314 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
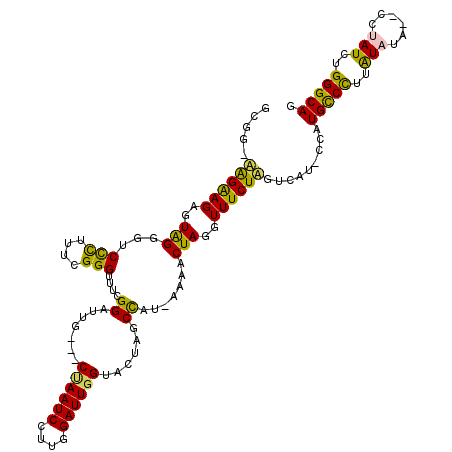
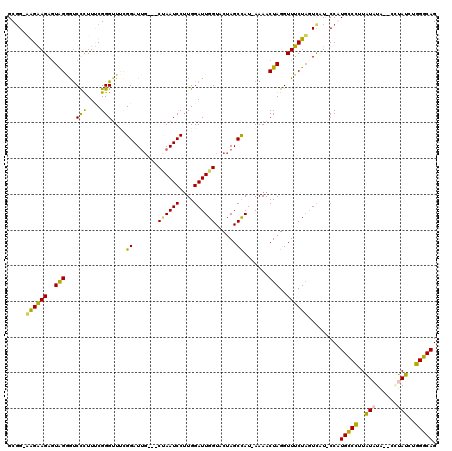
>X_DroMel_CAF1 2013943 114 - 22224390 GCGG-AGGGAGAGUAGGGUCUCUUUCGGGUUUUGGAUUC---CUAAUCCUUGGAUUGGUACUAGCUAU-AAAACUAGGUUUCUUGUCAU-CCGUGCCCUUAUCUGGCCCUAUCUGGGCAG .(((-((..((((......))))..)((((..(((((..---((((((....))))))..((((....-....))))..........))-))).))))...))))((((.....)))).. ( -39.50) >DroSec_CAF1 6863 112 - 1 GCGG-AGGAAGAGUAGGGUCUCUUUCGGGGUUCGGAUUG---CUAAUCCUUGGAUUGGUACUAGCCAU-AAAACUAGGUUUCUAGUCAU-CCGUGUCCUUAUCUA--CCUAUCUGGGCAG ((((-((((((((......)))))))((((..(((((((---((((((....))))))))........-...(((((....))))).))-)))..))))..))).--((.....)))).. ( -35.50) >DroEre_CAF1 7438 114 - 1 GCUG-UAGGAGAAUAGAGUCCUUUCCGGGUUACGGAUGG---CCAAUCCAUGGAUUGGUACUAGCCAUAAAAACUAGGUUUCUAUUUAUGCCAUGCCUUUGUAUA--CAUAUCUGGGCAG ....-..((.((((((((.(((.((((.....))))..(---((((((....)))))))................))).))))))))...)).(((((..(((..--..)))..))))). ( -34.80) >DroYak_CAF1 6961 116 - 1 GCGGAUAGAAGAGUGGCACCCUUU-CGGGUUCAGGAUUGUUCCGAAUCCUUGGAUUGGUACUAGCCAUUAAAACUAGGUUUCUAUUUAU-CCAUGCCCUUAUAUA--CGUAUCGGGGCAG ..(((((((..(((((((((..((-(((((((.(((....)))))))))..)))..)))....)))))).....(((....))))))))-)).((((((.(((..--..))).)))))). ( -38.80) >consensus GCGG_AAGAAGAGUAGGGUCCCUUUCGGGUUUCGGAUUG___CUAAUCCUUGGAUUGGUACUAGCCAU_AAAACUAGGUUUCUAGUCAU_CCAUGCCCUUAUAUA__CCUAUCUGGGCAG .....((((((..(((...(((....)))....((.......((((((....))))))......)).......)))..)))))).........(((((..(((......)))..))))). (-22.53 = -21.22 + -1.31)

| Location | 2,013,982 – 2,014,095 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 81.29 |
| Mean single sequence MFE | -36.13 |
| Consensus MFE | -22.14 |
| Energy contribution | -22.07 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.61 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.514467 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
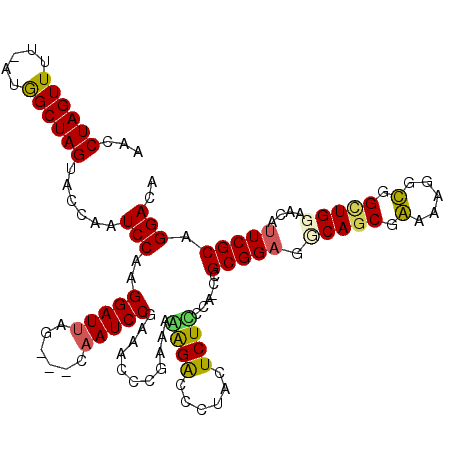
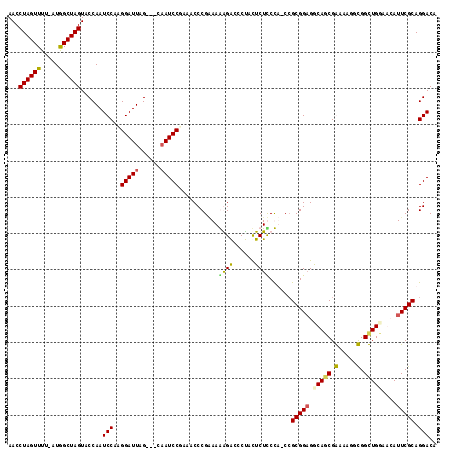
>X_DroMel_CAF1 2013982 113 + 22224390 AACCUAGUUUU-AUAGCUAGUACCAAUCCAAGGAUUAG---GAAUCCAAAACCCGAAAGAGACCCUACUCUCCCU-CCGCGGAGGCAGCGAAAAGGUGGUUGGAGGAUUCGCAGGACA ...((((((..-..))))))..((((((....))))..---((((((..(((((....)..(((...(.((.(((-(....)))).)).)....)))))))...))))))...))... ( -32.10) >DroSec_CAF1 6900 113 + 1 AACCUAGUUUU-AUGGCUAGUACCAAUCCAAGGAUUAG---CAAUCCGAACCCCGAAAGAGACCCUACUCUUCCU-CCGCGGAGGCAUCGAAAGGGUGGUUGGAGGAUUCGCAGGACA ...((((((..-..))))))..((((((....)))).(---((((((.((((.(....)..(((((......(((-(....)))).......)))))))))...))))).)).))... ( -34.22) >DroEre_CAF1 7476 114 + 1 AACCUAGUUUUUAUGGCUAGUACCAAUCCAUGGAUUGG---CCAUCCGUAACCCGGAAAGGACUCUAUUCUCCUA-CAGCGGUGGCAGCGGAAUGGCGGCUGUAACAUUCGCAGGACA ..((((((((((.((((((((.(((.....))))))))---)))((((.....)))))))))))...........-..((((..(((((.(.....).))))).....)))))))... ( -36.40) >DroYak_CAF1 6998 117 + 1 AACCUAGUUUUAAUGGCUAGUACCAAUCCAAGGAUUCGGAACAAUCCUGAACCCG-AAAGGGUGCCACUCUUCUAUCCGCGGAGACAGCGGAAAGGCGGCUGUAACAUUCGCAGGACA ...((((((.....))))))..((((((....)))).)).....(((((.((((.-...))))(((.(.(((...((((((....).))))))))).)))...........))))).. ( -41.80) >consensus AACCUAGUUUU_AUGGCUAGUACCAAUCCAAGGAUUAG___CAAUCCGAAACCCGAAAAAGACCCUACUCUCCCA_CCGCGGAGGCAGCGAAAAGGCGGCUGGAACAUUCGCAGGACA ...((((((.....))))))......(((..(((((......)))))...........((((......))))......(((((.(((((.(.....).)))))....))))).))).. (-22.14 = -22.07 + -0.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:52:40 2006