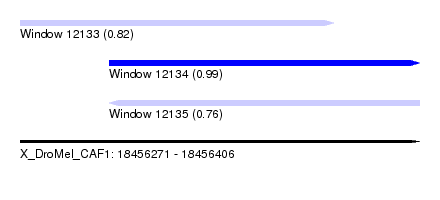
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,456,271 – 18,456,406 |
| Length | 135 |
| Max. P | 0.986789 |

| Location | 18,456,271 – 18,456,377 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 98.74 |
| Mean single sequence MFE | -28.73 |
| Consensus MFE | -28.13 |
| Energy contribution | -28.13 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820446 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18456271 106 + 22224390 CUCGAUUUGUUUCUCGGCCAACGGAAACGGGCCAAAAAUCGCACAAUCGACGGCACUAUGGCCAAAUUCAAAUCGAAAUCGAAGUCCAAAUGCGCCAGCAAAGCGU ..((.((((((....((((..((....))))))......((((.....((((((......))).........(((....))).)))....))))..)))))).)). ( -29.00) >DroSec_CAF1 73 106 + 1 CUCGACUUGUUUCUCGGCCAACGGAAACGGGCCAAAAAUCGCACAAUCGACGGCACUACGGCCAAAUUCAAAUCGAAAUCGAAGUCCAAAUGCGCCAGCAAAGCGU ..((..(((((....((((..((....))))))......((((.....((((((......))).........(((....))).)))....))))..)))))..)). ( -28.60) >DroSim_CAF1 416 106 + 1 CUCGACUUGUUUCUCGGCCAACGGAAACGGGCCAAAAAUCGCACAAUCGACGGCACUAUGGCCAAAUUCAAAUCGAAAUCGAAGUCCAAAUGCGCCAGCAAAGCGU ..((..(((((....((((..((....))))))......((((.....((((((......))).........(((....))).)))....))))..)))))..)). ( -28.60) >consensus CUCGACUUGUUUCUCGGCCAACGGAAACGGGCCAAAAAUCGCACAAUCGACGGCACUAUGGCCAAAUUCAAAUCGAAAUCGAAGUCCAAAUGCGCCAGCAAAGCGU ..((..(((((....((((..((....))))))......((((.....((((((......))).........(((....))).)))....))))..)))))..)). (-28.13 = -28.13 + 0.00)


| Location | 18,456,301 – 18,456,406 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 99.37 |
| Mean single sequence MFE | -33.70 |
| Consensus MFE | -33.70 |
| Energy contribution | -33.70 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -4.25 |
| Structure conservation index | 1.00 |
| SVM decision value | 2.05 |
| SVM RNA-class probability | 0.986789 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
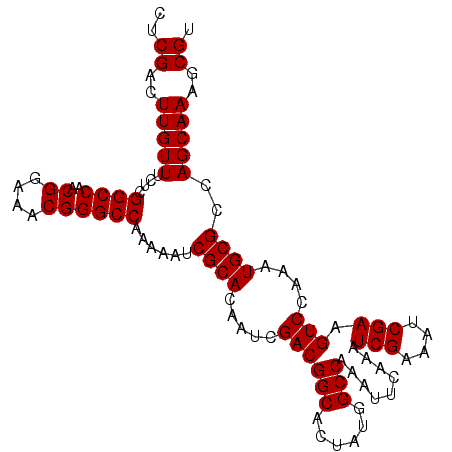
>X_DroMel_CAF1 18456301 105 + 22224390 GCCAAAAAUCGCACAAUCGACGGCACUAUGGCCAAAUUCAAAUCGAAAUCGAAGUCCAAAUGCGCCAGCAAAGCGUAUCAAAUGCCAUGUUGACCGCAUUUGGAC ((........))....((((.(((......)))...(((.....))).)))).(((((((((((.(((((..((((.....))))..)))))..))))))))))) ( -33.70) >DroSec_CAF1 103 105 + 1 GCCAAAAAUCGCACAAUCGACGGCACUACGGCCAAAUUCAAAUCGAAAUCGAAGUCCAAAUGCGCCAGCAAAGCGUAUCAAAUGCCAUGUUGACCGCAUUUGGAC ((........))....((((.(((......)))...(((.....))).)))).(((((((((((.(((((..((((.....))))..)))))..))))))))))) ( -33.70) >DroSim_CAF1 446 105 + 1 GCCAAAAAUCGCACAAUCGACGGCACUAUGGCCAAAUUCAAAUCGAAAUCGAAGUCCAAAUGCGCCAGCAAAGCGUAUCAAAUGCCAUGUUGACCGCAUUUGGAC ((........))....((((.(((......)))...(((.....))).)))).(((((((((((.(((((..((((.....))))..)))))..))))))))))) ( -33.70) >consensus GCCAAAAAUCGCACAAUCGACGGCACUAUGGCCAAAUUCAAAUCGAAAUCGAAGUCCAAAUGCGCCAGCAAAGCGUAUCAAAUGCCAUGUUGACCGCAUUUGGAC ((........))....((((.(((......)))...(((.....))).)))).(((((((((((.(((((..((((.....))))..)))))..))))))))))) (-33.70 = -33.70 + -0.00)

| Location | 18,456,301 – 18,456,406 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 99.37 |
| Mean single sequence MFE | -32.97 |
| Consensus MFE | -32.99 |
| Energy contribution | -32.77 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.65 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.764859 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
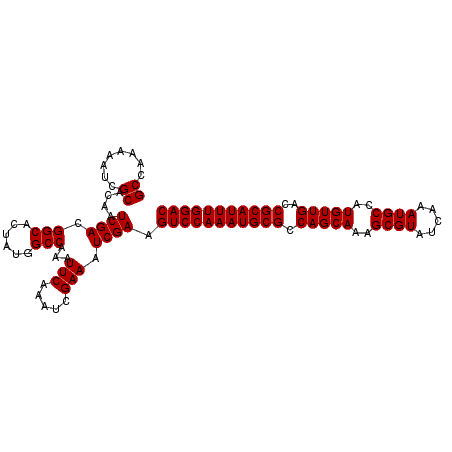
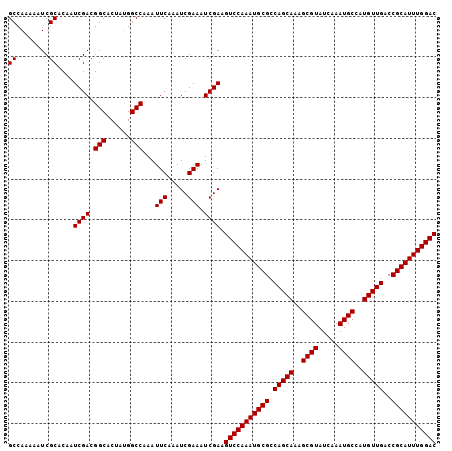
>X_DroMel_CAF1 18456301 105 - 22224390 GUCCAAAUGCGGUCAACAUGGCAUUUGAUACGCUUUGCUGGCGCAUUUGGACUUCGAUUUCGAUUUGAAUUUGGCCAUAGUGCCGUCGAUUGUGCGAUUUUUGGC (((((((((((.(((.((.(((.........))).)).))))))))))))))..(((.(((.....))).)))((((...(((((.....)).))).....)))) ( -33.00) >DroSec_CAF1 103 105 - 1 GUCCAAAUGCGGUCAACAUGGCAUUUGAUACGCUUUGCUGGCGCAUUUGGACUUCGAUUUCGAUUUGAAUUUGGCCGUAGUGCCGUCGAUUGUGCGAUUUUUGGC (((((((((((.(((.((.(((.........))).)).)))))))))))))).((((..(((..((((...((((......)))))))).....)))...)))). ( -32.90) >DroSim_CAF1 446 105 - 1 GUCCAAAUGCGGUCAACAUGGCAUUUGAUACGCUUUGCUGGCGCAUUUGGACUUCGAUUUCGAUUUGAAUUUGGCCAUAGUGCCGUCGAUUGUGCGAUUUUUGGC (((((((((((.(((.((.(((.........))).)).))))))))))))))..(((.(((.....))).)))((((...(((((.....)).))).....)))) ( -33.00) >consensus GUCCAAAUGCGGUCAACAUGGCAUUUGAUACGCUUUGCUGGCGCAUUUGGACUUCGAUUUCGAUUUGAAUUUGGCCAUAGUGCCGUCGAUUGUGCGAUUUUUGGC (((((((((((.(((.((.(((.........))).)).))))))))))))))..(((.(((.....))).)))((((...(((((.....)).))).....)))) (-32.99 = -32.77 + -0.22)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:38:01 2006