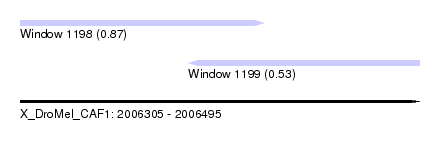
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,006,305 – 2,006,495 |
| Length | 190 |
| Max. P | 0.869589 |

| Location | 2,006,305 – 2,006,421 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 95.69 |
| Mean single sequence MFE | -16.43 |
| Consensus MFE | -15.37 |
| Energy contribution | -15.62 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.869589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
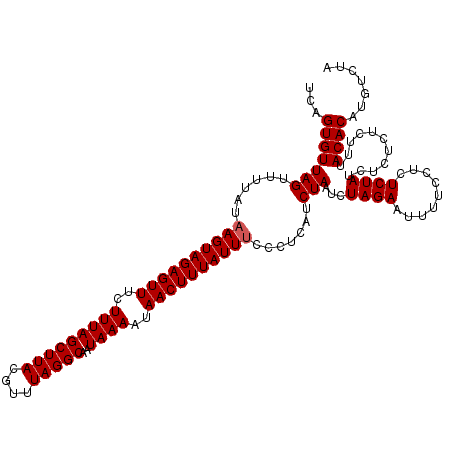
>X_DroMel_CAF1 2006305 116 + 22224390 UCAGUGUUAGUUUUAUAAGUAGAGUUUCUUUAGCUUACGUUUAGGCAAUAAAAUAACUUUAUUUCCCACAUCUAAUCUAGAAUUUUCCUCUCUAACUCUCUCUUUACACAUGUCUA ..((.(((((......((((((((((..(((((((((....)))))..))))..))))))))))......((((...))))..........))))).))................. ( -16.20) >DroSec_CAF1 7127 116 + 1 UCAGUGUUAGUUUUAUAAGUAGAGUUUCUUUAGCUUACGUUUAGGCAAUAAAAUAACUUUAUUUCCCUCAUCUAAUCUAGAAUUUUCCUCUCUAUCUCUCUCUUUACACAUGUCUA ...(((((((......((((((((((..(((((((((....)))))..))))..)))))))))).......)))...((((.........))))...........))))....... ( -15.62) >DroSim_CAF1 6033 116 + 1 UCAGUGUUAGUUUUAUAAGUAGAGUUUCUUUAGCUUACGUUUAGGCAAUAAAAUAACUUUAUUUCCCUCAUCUAAUCUAGAAUUUUCCUCUCUAUCUCUCUCUUUACACAUGUCUA ...(((((((......((((((((((..(((((((((....)))))..))))..)))))))))).......)))...((((.........))))...........))))....... ( -15.62) >DroEre_CAF1 7155 116 + 1 UCAGUGUUAGUUUUAUAAGUAGAGUUUCUUUAGCUUACAUUUAGGCAAUAAAAUAACUUUAUUCUCCUCAUCUAAUCUAGAGUUUCUUUCUCUAUCUCUCUCUUUACACUUGUCUA ..((((((((.......(((((((((..(((((((((....)))))..))))..)))))))))........)))...(((((.......)))))...........)))))...... ( -18.26) >consensus UCAGUGUUAGUUUUAUAAGUAGAGUUUCUUUAGCUUACGUUUAGGCAAUAAAAUAACUUUAUUUCCCUCAUCUAAUCUAGAAUUUUCCUCUCUAUCUCUCUCUUUACACAUGUCUA ...(((((((......((((((((((..(((((((((....)))))..))))..)))))))))).......)))...((((.........))))...........))))....... (-15.37 = -15.62 + 0.25)

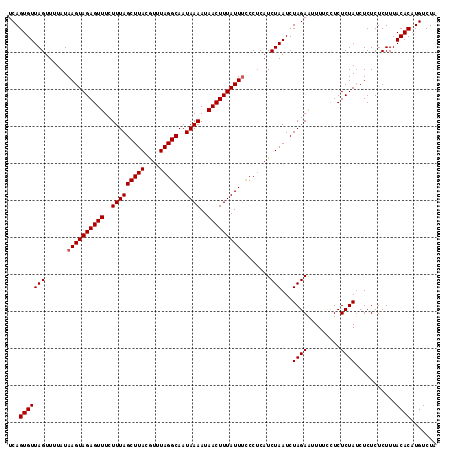
| Location | 2,006,385 – 2,006,495 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 95.45 |
| Mean single sequence MFE | -22.08 |
| Consensus MFE | -20.61 |
| Energy contribution | -20.30 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.93 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.530193 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
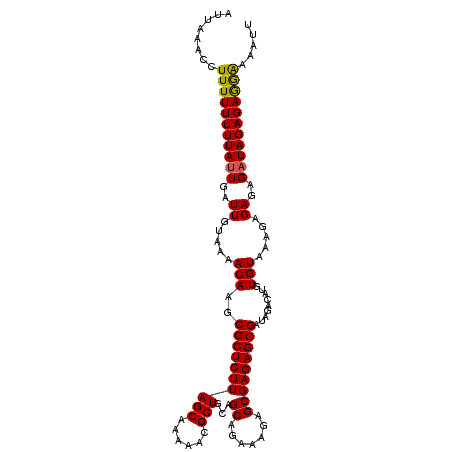
>X_DroMel_CAF1 2006385 110 - 22224390 AUUAAACCUUUUUUUUAUUGAUUGUAAAACAAGCGCUCUUAGCAAAAACGCUGCAGCAGAAAGAGCGAGAGCGAUAGACAUGUGUAAAGAGAGAGUUAGAGAGGAAAAUU ....(((.((((((((((....(((........((((((((((......)))...((.......)))))))))....)))...)))))))))).)))............. ( -22.30) >DroSec_CAF1 7207 110 - 1 AUUAAACCUUUUUUUUAUUGAUUGUAAAACAAGCGCUCUUAGCAAAAACGCUGCAGCAGAAAGAGCGAGAGCGAUAGACAUGUGUAAAGAGAGAGAUAGAGAGGAAAAUU ......(((((((....((..((.....(((..((((((((((......)))...((.......))))))))).......))).....))..))...)))))))...... ( -21.20) >DroSim_CAF1 6113 110 - 1 AUUAAACCUUUUUUUUAUUGAUUGUAAAACAAGCGCUCUUAGCAAAAACGCUGCAGCAGAAAGAGCGAGAGCGAUAGACAUGUGUAAAGAGAGAGAUAGAGAGGAAAAUU ......(((((((....((..((.....(((..((((((((((......)))...((.......))))))))).......))).....))..))...)))))))...... ( -21.20) >DroEre_CAF1 7235 109 - 1 AUUAAA-CUUUUUUUUAUUCAUUGUAAAACAAGCGCUCUUAGCAAAAACGCUGCAGCUGGAAGAGCGAGAGCGAUAGACAAGUGUAAAGAGAGAGAUAGAGAAAGAAACU ......-(((((((((.....((((........((((((((((......)))...(((.....))))))))))....)))).....)))))))))............... ( -23.60) >consensus AUUAAACCUUUUUUUUAUUGAUUGUAAAACAAGCGCUCUUAGCAAAAACGCUGCAGCAGAAAGAGCGAGAGCGAUAGACAUGUGUAAAGAGAGAGAUAGAGAGGAAAAUU ........(((((((((((..((.....(((..((((((((((......)))...((.......))))))))).........))).....))..)))))))))))..... (-20.61 = -20.30 + -0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:52:38 2006