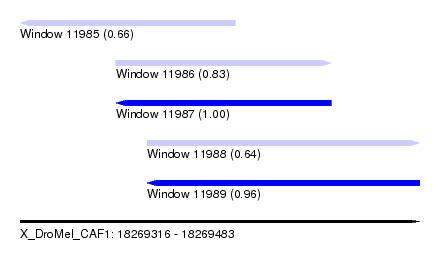

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,269,316 – 18,269,483 |
| Length | 167 |
| Max. P | 0.996316 |

| Location | 18,269,316 – 18,269,406 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 73.47 |
| Mean single sequence MFE | -23.70 |
| Consensus MFE | -14.64 |
| Energy contribution | -15.39 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.664446 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18269316 90 - 22224390 ACCAGAUUCAUCAUGGC---UAAUGAUGGGUCAUUCCUAAUCGGC---------------------------AACGCUCAUCUUUAACACGCCGCCGAGCUCACGUGCAAUAAAUGCAUU ....((((((((((...---..))))))))))........(((((---------------------------...((.............)).)))))......(((((.....))))). ( -23.32) >DroSim_CAF1 12287 93 - 1 UUCAGAUUCAUCACGAUUCAUAAUGAUGGGUCAUUCCUAAUCGGC---------------------------ACCGCUCAUCUUCAACAAGCCGCCGAGCUCACGUGCAAUAAAUGCAUU ....(((((((((..........)))))))))........(((((---------------------------...(((...........))).)))))......(((((.....))))). ( -21.10) >DroEre_CAF1 42214 99 - 1 GCCAGAUUCAUCAUAG---CUAAUGAUGGGUCAUUCCUGCUCGG------------------AAUGACUAUCUCCGCUCAUCUUCAACAAGUCGCCGUGCUCACGUGCAAUUAAUGCAUU ((..((((((((((..---...))))))(((((((((.....))------------------)))))))....................))))(((((....))).)).......))... ( -26.00) >DroYak_CAF1 34234 117 - 1 UUCAGAUUCAUCAUUG---CUAAUGAUAGGUCAUGCCUAAUCGGCGUAUAAAUACUUAUGGCAGUGACUAUCUCCGUUCAUCUUCAACUAGCCGCCGUGCUCACGUGCAAUUAAUGCAUU ....(((..(((((..---...)))))..)))((((.....(((((((....)))....((((((.....................))).)))))))(((......)))......)))). ( -24.40) >consensus UCCAGAUUCAUCAUGG___CUAAUGAUGGGUCAUUCCUAAUCGGC___________________________ACCGCUCAUCUUCAACAAGCCGCCGAGCUCACGUGCAAUAAAUGCAUU ....((((((((((........)))))))))).........((((................................................)))).......(((((.....))))). (-14.64 = -15.39 + 0.75)



| Location | 18,269,356 – 18,269,446 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 68.90 |
| Mean single sequence MFE | -28.56 |
| Consensus MFE | -16.68 |
| Energy contribution | -16.52 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.833201 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18269356 90 + 22224390 UGAGCGUU---------------------------GCCGAUUAGGAAUGACCCAUCAUUA---GCCAUGAUGAAUCUGGUAAGUGGGGCCAGUUAACUGACCAGCUAGAUUCCACGGAGC ...((...---------------------------.(((((((((.((((....))))..---.)).))))(((((((((...(((...(((....))).))))))))))))..))).)) ( -24.00) >DroSec_CAF1 37726 93 + 1 UGAGCGGU---------------------------GCCGAUUAGGAAUGACCCAUCAUUAUGAAUCGUGAUGAAUCUCAAAAGUGGGGCCAGUUAACUGGCCAGCUAGAUUCCACGGAAC ........---------------------------.((((((...(((((....)))))...))))(((..((((((....(((..((((((....)))))).))))))))))))))... ( -29.90) >DroSim_CAF1 12327 93 + 1 UGAGCGGU---------------------------GCCGAUUAGGAAUGACCCAUCAUUAUGAAUCGUGAUGAAUCUGAAAAGUGGGGCCAGUUAACUGGGCAGCUAGAUUCCACGGAGC ...((...---------------------------.((((((...(((((....)))))...))))(((..((((((....(((..(.((((....)))).).)))))))))))))).)) ( -26.50) >DroEre_CAF1 42254 92 + 1 UGAGCGGAGAUAGUCAUU------------------CCGAGCAGGAAUGACCCAUCAUUAG---CUAUGAUGAAUCUGGCAAAUGUGGCCAGUAAGCUGGUUAACUAGAUGCU------- ..(((.......((((((------------------((.....)))))))).((((((...---..)))))).((((((......(((((((....))))))).)))))))))------- ( -32.50) >DroYak_CAF1 34274 117 + 1 UGAACGGAGAUAGUCACUGCCAUAAGUAUUUAUACGCCGAUUAGGCAUGACCUAUCAUUAG---CAAUGAUGAAUCUGAAUAAUGGGGCCAGUAAGCUGGUUAACUGGAUUCCAUCAAAC .....((((.((((.....((((...((((((...(((.....)))........((((((.---...))))))...))))))))))((((((....)))))).))))..))))....... ( -29.90) >consensus UGAGCGGU___________________________GCCGAUUAGGAAUGACCCAUCAUUAG___CCAUGAUGAAUCUGAAAAGUGGGGCCAGUUAACUGGCCAGCUAGAUUCCACGGAAC ...........................................(((((((..((((((........))))))..))..........((((((....))))))......)))))....... (-16.68 = -16.52 + -0.16)

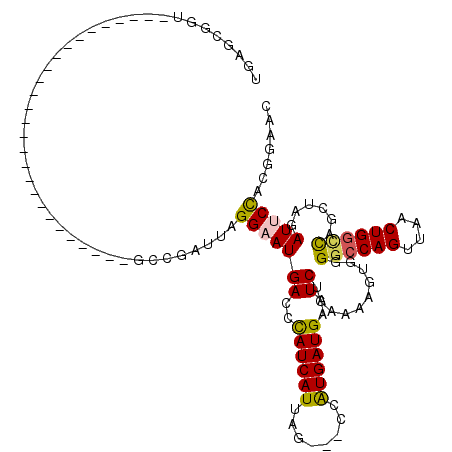
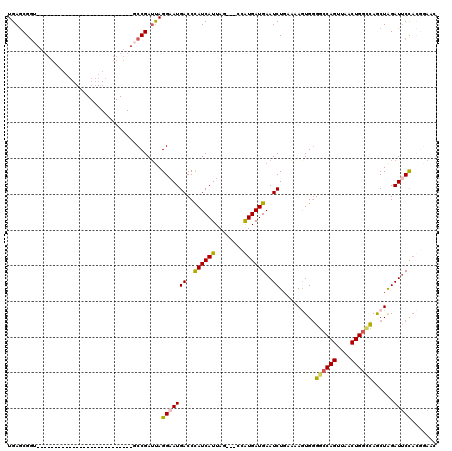
| Location | 18,269,356 – 18,269,446 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 68.90 |
| Mean single sequence MFE | -27.64 |
| Consensus MFE | -14.92 |
| Energy contribution | -15.92 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.54 |
| SVM decision value | 2.68 |
| SVM RNA-class probability | 0.996316 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18269356 90 - 22224390 GCUCCGUGGAAUCUAGCUGGUCAGUUAACUGGCCCCACUUACCAGAUUCAUCAUGGC---UAAUGAUGGGUCAUUCCUAAUCGGC---------------------------AACGCUCA ((.(((.(((((......((((((....))))))..........((((((((((...---..)))))))))))))))....))).---------------------------...))... ( -29.00) >DroSec_CAF1 37726 93 - 1 GUUCCGUGGAAUCUAGCUGGCCAGUUAACUGGCCCCACUUUUGAGAUUCAUCACGAUUCAUAAUGAUGGGUCAUUCCUAAUCGGC---------------------------ACCGCUCA ((.(((.(((((......((((((....))))))..........(((((((((..........))))))))))))))....))).---------------------------))...... ( -29.00) >DroSim_CAF1 12327 93 - 1 GCUCCGUGGAAUCUAGCUGCCCAGUUAACUGGCCCCACUUUUCAGAUUCAUCACGAUUCAUAAUGAUGGGUCAUUCCUAAUCGGC---------------------------ACCGCUCA ((.(((.(((((........((((....))))............(((((((((..........))))))))))))))....))).---------------------------...))... ( -21.60) >DroEre_CAF1 42254 92 - 1 -------AGCAUCUAGUUAACCAGCUUACUGGCCACAUUUGCCAGAUUCAUCAUAG---CUAAUGAUGGGUCAUUCCUGCUCGG------------------AAUGACUAUCUCCGCUCA -------(((..................(((((.......)))))...((((((..---...))))))(((((((((.....))------------------)))))))......))).. ( -27.50) >DroYak_CAF1 34274 117 - 1 GUUUGAUGGAAUCCAGUUAACCAGCUUACUGGCCCCAUUAUUCAGAUUCAUCAUUG---CUAAUGAUAGGUCAUGCCUAAUCGGCGUAUAAAUACUUAUGGCAGUGACUAUCUCCGUUCA (..((((((...(((((..........)))))..))))))..)((((...((((((---(((((((....))))(((.....))).............)))))))))..))))....... ( -31.10) >consensus GCUCCGUGGAAUCUAGCUGACCAGUUAACUGGCCCCACUUUUCAGAUUCAUCAUGG___AUAAUGAUGGGUCAUUCCUAAUCGGC___________________________ACCGCUCA .....((((...........((((....))))............((((((((((........)))))))))).........................................))))... (-14.92 = -15.92 + 1.00)


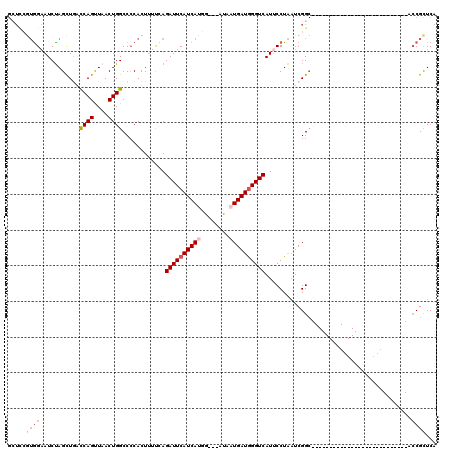
| Location | 18,269,369 – 18,269,483 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.25 |
| Mean single sequence MFE | -34.95 |
| Consensus MFE | -22.83 |
| Energy contribution | -23.82 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.640103 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18269369 114 + 22224390 UUAGGAAUGACCCAUCAUUA---GCCAUGAUGAAUCUGGUAAGUGGGGCCAGUUAACUGACCAGCUAGAUUCCACGGAGCACGUGCCGAUGGUU-UAUUAGUAAUGAAAUGCUGUUGG-- ...((.((((....))))..---.)).....(((((((((...(((...(((....))).))))))))))))((((.....))))((((((((.-..(((....)))...))))))))-- ( -31.10) >DroSec_CAF1 37739 117 + 1 UUAGGAAUGACCCAUCAUUAUGAAUCGUGAUGAAUCUCAAAAGUGGGGCCAGUUAACUGGCCAGCUAGAUUCCACGGAACACGUGCCGAUGGGU-UGUUGGUAAUGAAAUGCUGUUGG-- .....((..(((((((..((((..(((((..((((((....(((..((((((....)))))).))))))))))))))....))))..)))))))-..))((((......)))).....-- ( -40.90) >DroSim_CAF1 12340 117 + 1 UUAGGAAUGACCCAUCAUUAUGAAUCGUGAUGAAUCUGAAAAGUGGGGCCAGUUAACUGGGCAGCUAGAUUCCACGGAGCACGUGCCGAUGGGU-UGUUGGUAAUGAAAUGCUGAUGG-- ........((((((((........(((((..((((((....(((..(.((((....)))).).)))))))))))))).((....)).)))))))-)((..(((......)))..))..-- ( -37.40) >DroYak_CAF1 34314 117 + 1 UUAGGCAUGACCUAUCAUUAG---CAAUGAUGAAUCUGAAUAAUGGGGCCAGUAAGCUGGUUAACUGGAUUCCAUCAAACACGUGCUGCUGAUUACACAAGUAAUGCUAUGCCGUAGGAU ...((((((.....(((.(((---((.((((((((((.........((((((....))))))....))))).)))))......))))).)))((((....))))...))))))....... ( -30.42) >consensus UUAGGAAUGACCCAUCAUUAU__ACCAUGAUGAAUCUGAAAAGUGGGGCCAGUUAACUGGCCAGCUAGAUUCCACGGAACACGUGCCGAUGGGU_UAUUAGUAAUGAAAUGCUGUUGG__ (((..(((((((((((...............((((((....(((..((((((....)))))).)))))))))((((.....))))..))))))).))))..)))................ (-22.83 = -23.82 + 1.00)

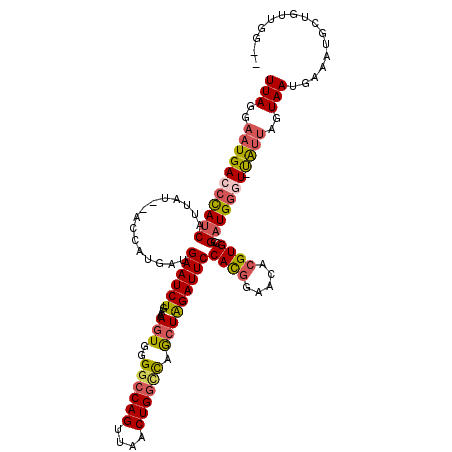
| Location | 18,269,369 – 18,269,483 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.25 |
| Mean single sequence MFE | -29.70 |
| Consensus MFE | -20.14 |
| Energy contribution | -19.58 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.68 |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.957060 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18269369 114 - 22224390 --CCAACAGCAUUUCAUUACUAAUA-AACCAUCGGCACGUGCUCCGUGGAAUCUAGCUGGUCAGUUAACUGGCCCCACUUACCAGAUUCAUCAUGGC---UAAUGAUGGGUCAUUCCUAA --....(((((((........))).-..((((.(((....)).).))))......))))(((((....)))))...........((((((((((...---..))))))))))........ ( -28.40) >DroSec_CAF1 37739 117 - 1 --CCAACAGCAUUUCAUUACCAACA-ACCCAUCGGCACGUGUUCCGUGGAAUCUAGCUGGCCAGUUAACUGGCCCCACUUUUGAGAUUCAUCACGAUUCAUAAUGAUGGGUCAUUCCUAA --.......................-((((((((....(((...((((((((((((..((((((....))))))...))....))))))..))))...)))..))))))))......... ( -33.10) >DroSim_CAF1 12340 117 - 1 --CCAUCAGCAUUUCAUUACCAACA-ACCCAUCGGCACGUGCUCCGUGGAAUCUAGCUGCCCAGUUAACUGGCCCCACUUUUCAGAUUCAUCACGAUUCAUAAUGAUGGGUCAUUCCUAA --.......................-((((((((((....))).((((((((((((.((.((((....))))...))))....))))))..)))).........)))))))......... ( -26.30) >DroYak_CAF1 34314 117 - 1 AUCCUACGGCAUAGCAUUACUUGUGUAAUCAGCAGCACGUGUUUGAUGGAAUCCAGUUAACCAGCUUACUGGCCCCAUUAUUCAGAUUCAUCAUUG---CUAAUGAUAGGUCAUGCCUAA .......(((((.(((((((....)))))..))......((..((((((...(((((..........)))))..))))))..))(((..(((((..---...)))))..))))))))... ( -31.00) >consensus __CCAACAGCAUUUCAUUACCAACA_AACCAUCGGCACGUGCUCCGUGGAAUCUAGCUGGCCAGUUAACUGGCCCCACUUUUCAGAUUCAUCACGAU__AUAAUGAUGGGUCAUUCCUAA .............((((((..............(((....))).((((((((((((..((((((....))))))...))....))))))..)))).....))))))((((.....)))). (-20.14 = -19.58 + -0.56)

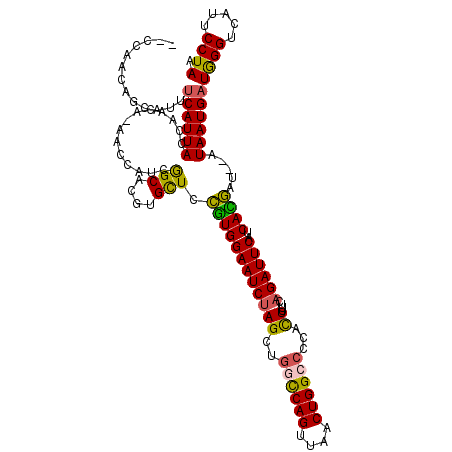

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:35:55 2006