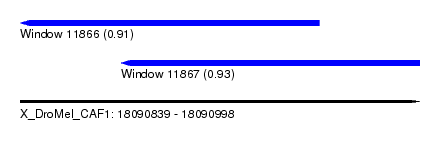

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,090,839 – 18,090,998 |
| Length | 159 |
| Max. P | 0.929747 |

| Location | 18,090,839 – 18,090,958 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.83 |
| Mean single sequence MFE | -39.35 |
| Consensus MFE | -36.08 |
| Energy contribution | -37.32 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.912303 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18090839 119 - 22224390 AACAAUAAAAAACAAUCCGCUGCC-CCACCGCCCCCUCGCGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUGCGCACAAGACGCACCAGAGAUGGAUAUGGCAGCGAUCCCA .................(((((((-...((((......)))).(((((((......((((.(((.......)))))))(((((....).)))).....)))))))..)))))))...... ( -42.60) >DroSec_CAF1 13186 120 - 1 AACAAUAAAAAACAAUCCGCUGCCCCCACCGCCCCCUCGAGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUGCGCACAAGACGCACCAGAGAUGGAUAUGGCAGCGAUCCCA .................(((((((..........((....)).(((((((......((((.(((.......)))))))(((((....).)))).....)))))))..)))))))...... ( -37.20) >DroEre_CAF1 10350 119 - 1 AACAAUAAAAAACAAUCCGCUGCCCCCACCGCCCCCUCGCGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUGCGCACAAGACGCACCAGAGAUGGAUAUGGCAGCGAUCCC- .................(((((((....((((......)))).(((((((......((((.(((.......)))))))(((((....).)))).....)))))))..))))))).....- ( -42.60) >DroYak_CAF1 12348 114 - 1 AACAAUAAAAAACAAUCCGCCGCCGC-----CCCCCUCGCGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUGCGCACAAGACGCACCAGAGAUGGAUAUGGCAGCGAUCCC- .............(((((((.(((((-----.......)).((((.......)))))))..)))))))...(((.((((((((....).))))(((....)))....))).))).....- ( -35.00) >consensus AACAAUAAAAAACAAUCCGCUGCCCCCACCGCCCCCUCGCGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUGCGCACAAGACGCACCAGAGAUGGAUAUGGCAGCGAUCCC_ .................(((((((....((((......)))).(((((((......((((.(((.......)))))))(((((....).)))).....)))))))..)))))))...... (-36.08 = -37.32 + 1.25)

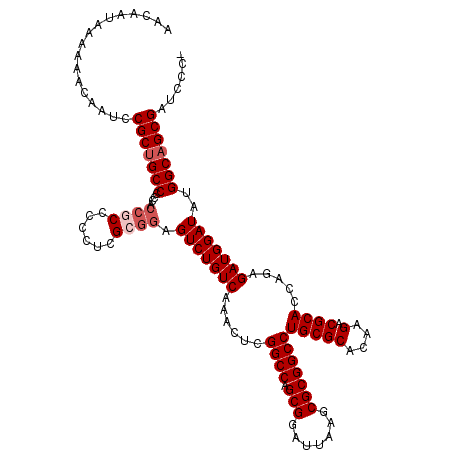
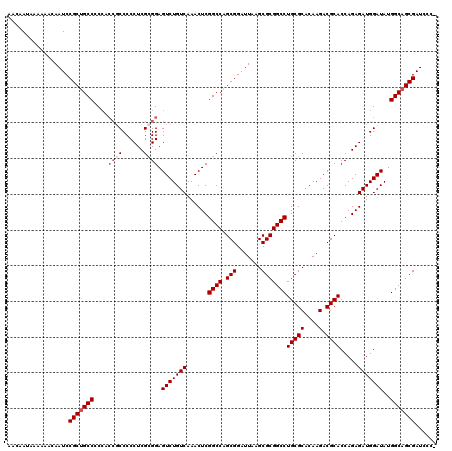
| Location | 18,090,879 – 18,090,998 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.39 |
| Mean single sequence MFE | -37.75 |
| Consensus MFE | -34.35 |
| Energy contribution | -35.35 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.929747 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18090879 119 - 22224390 GCGCGGACAGCGCGAAAUAAAAGCGCAGAAAUGUUAAAUAAACAAUAAAAAACAAUCCGCUGCC-CCACCGCCCCCUCGCGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUG ((((.....)))).........((((.....((((.....)))).........(((((((((((-....(((......)))((((.......)))))).)))))))))..))))...... ( -39.80) >DroSec_CAF1 13226 120 - 1 GCGCGGACAGCGCGAAAUAAAAGCGCAGAAAUGUUAAAUAAACAAUAAAAAACAAUCCGCUGCCCCCACCGCCCCCUCGAGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUG ((((.....)))).........((((.....((((.....)))).........(((((((((((.....((......))..((((.......)))))).)))))))))..))))...... ( -35.80) >DroEre_CAF1 10389 120 - 1 GCGCGGACAGCGCGAAAUAAAAGCGCAGAAAUGUUAAAUAAACAAUAAAAAACAAUCCGCUGCCCCCACCGCCCCCUCGCGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUG ((((.....)))).........((((.....((((.....)))).........(((((((((((.....(((......)))((((.......)))))).)))))))))..))))...... ( -39.80) >DroYak_CAF1 12387 115 - 1 GCGCGGACAGCGCGAAAUAAAAGCGCAGAAAUGUUAAAUAAACAAUAAAAAACAAUCCGCCGCCGC-----CCCCCUCGCGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUG ((((.....)))).........((((.....((((.....)))).........(((((((.(((((-----.......)).((((.......)))))))..)))))))..))))...... ( -35.60) >consensus GCGCGGACAGCGCGAAAUAAAAGCGCAGAAAUGUUAAAUAAACAAUAAAAAACAAUCCGCUGCCCCCACCGCCCCCUCGCGGAGUCUGUCAAACUCGGCCAGCGGAUUAAGCGCGGCCUG ((((.....)))).........((((.....((((.....)))).........(((((((((((.....(((......)))((((.......)))))).)))))))))..))))...... (-34.35 = -35.35 + 1.00)

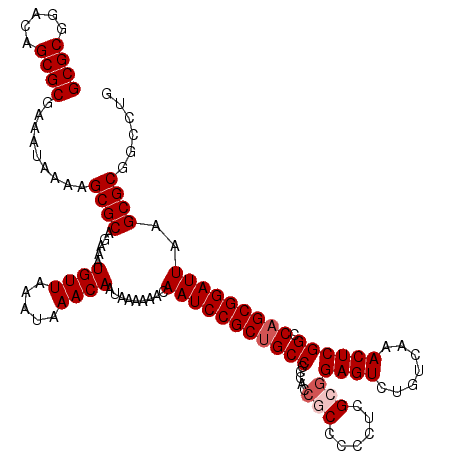

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:34:09 2006