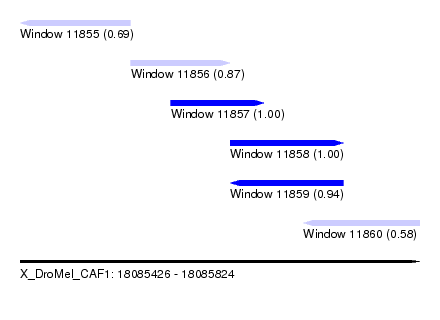
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,085,426 – 18,085,824 |
| Length | 398 |
| Max. P | 0.998312 |

| Location | 18,085,426 – 18,085,536 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.17 |
| Mean single sequence MFE | -33.90 |
| Consensus MFE | -20.32 |
| Energy contribution | -21.24 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.686371 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18085426 110 - 22224390 UAACAACAAGAAGCUGCUCCCGAUGUUGCAGCUACGCCAGUGGCAGUGUUGCA------GUUGCAAUUGCAGUCACUGCAGCAGCA---UAAAAAAAAUAAGGAGA-AAACCAGUAGAAA .............(((((....(((((((.(((((....)))))(((((((((------((....))))))).))))...))))))---)...........((...-...)))))))... ( -32.60) >DroSec_CAF1 7858 109 - 1 UAACAACAAGAAGCUGCUCCCGAUGUUGCUGCUGCGCCAGUGGCAGUGUUGCA------GUUGCAAUUGCAGUCGCUGCAGCCGCA---GAAA-AAAAUAAGGCGA-AGACCAGUAGAAA .............(((((.......((((.((((((((...)))(((((((((------((....))))))).))))))))).)))---)...-.......((...-...)))))))... ( -32.60) >DroSim_CAF1 7634 115 - 1 UAACAACAAGAAGCUGCUCCCGAUGUUGCUGCUGCGCCAGUGGCAGUGUUGCAGUUGCAGUUGCAAUUGCAGUCGCUGCAGCCGCA---GAAA-AAAAUAAGGCGA-AGACCAGUAGAAA .............(((((.......((((.((((((((...)))((((((((((((((....)))))))))).))))))))).)))---)...-.......((...-...)))))))... ( -40.50) >DroEre_CAF1 5454 101 - 1 UAACAACAAGAAGCUGCUCCCGAUGUUGCUGCUGCGCCAGUGGCAGUGUUGCA------GUUGCAGUUGCAGUUGC------------AGAAA-AAGACAAGGAGAGAAACCUGUGGAAA ......((...((...(((((..(((((((((..(....)..))))).(((((------..(((....)))..)))------------))...-..)))).)).)))....)).)).... ( -31.50) >DroYak_CAF1 6426 112 - 1 UAACAACAAGUAUCUGCUCCCGAUGUUGCUGCUGCCGCUGUGGCAGUGUUGCA------GUUGCAAUUGUGGUUGCUGUAGUCGCAGUAAAAA-AAAAUAAGGAGA-AAACCAGUAGAAA ............((((((.(((..(((((.((((((((((...)))))..)))------)).)))))..)))(((((((....)))))))...-.......((...-...)))))))).. ( -32.30) >consensus UAACAACAAGAAGCUGCUCCCGAUGUUGCUGCUGCGCCAGUGGCAGUGUUGCA______GUUGCAAUUGCAGUCGCUGCAGCCGCA___GAAA_AAAAUAAGGAGA_AAACCAGUAGAAA ............(((((...((((...((((((((....))))))))((((((........))))))....))))..)))))...................((.......))........ (-20.32 = -21.24 + 0.92)



| Location | 18,085,536 – 18,085,635 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.01 |
| Mean single sequence MFE | -33.08 |
| Consensus MFE | -28.14 |
| Energy contribution | -28.14 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873393 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18085536 99 + 22224390 AUUUGAUGUGGCACAUGUAAUAAGGCUAAUGAGGUGGGCCACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGCUACCA---------------------CACAC ...((.(((((....((((....((((.........))))..)))).(((.((((((((((((....)))...)))))).))).)))....)))---------------------)))). ( -30.50) >DroSec_CAF1 7967 99 + 1 AUUUGAUGUGGCACAUGUAAUAAGGCUAAUGAGGUGGGCCACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGCUACCA---------------------CACGC .......(((((...((((....((((.........))))..)))).(((.((((((((((((....)))...)))))).))).))))))))..---------------------..... ( -30.40) >DroSim_CAF1 7749 99 + 1 AUUUGAUGUGGCACAUGUAAUAAGGCUAAUGAGGUGGGCCACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGCUACCA---------------------CACGC .......(((((...((((....((((.........))))..)))).(((.((((((((((((....)))...)))))).))).))))))))..---------------------..... ( -30.40) >DroEre_CAF1 5555 99 + 1 AUUUGAUGUGGCACAUGUAAUGAGGCUAAUGAGGUGGGCCACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGAAUAUCC---------------------CACAC ......(((((....((((.((.(.(((......))).))).)))).(((.((((((((((((....)))...)))))).))).)))......)---------------------)))). ( -30.40) >DroYak_CAF1 6538 120 + 1 AUUUGAUGUGGCACAUGUAAUAAGGCUAAUGAGGUGGGCCACUGCCACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGAUACCCCAAGCCCCACAUCCCACAUCGCACAC ....(((((((...((((.....((((..((.(((((((....))).(((.((((((((((((....)))...)))))).))).)))..)))).))))))..)))).)))))))...... ( -43.70) >consensus AUUUGAUGUGGCACAUGUAAUAAGGCUAAUGAGGUGGGCCACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGCUACCA_____________________CACAC .......(((((.(((.(.((.......)).).))).))))).....(((.((((((((((((....)))...)))))).))).)))................................. (-28.14 = -28.14 + 0.00)


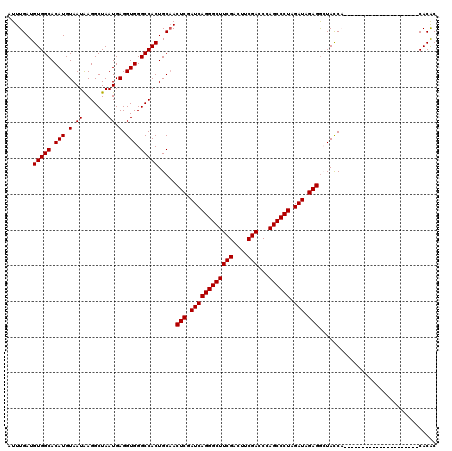
| Location | 18,085,576 – 18,085,669 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 72.85 |
| Mean single sequence MFE | -22.75 |
| Consensus MFE | -18.26 |
| Energy contribution | -18.26 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.80 |
| SVM decision value | 3.06 |
| SVM RNA-class probability | 0.998312 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18085576 93 + 22224390 ACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGCUACCA---------------------CACACCCCAUGCCUCCAUGCCCCCAUGCCGCCAUGCCAC------ ...(((...((((((((((((((....)))...)))))).))).(((((.....---------------------..........)))))((((....)))).))...)))...------ ( -24.46) >DroSec_CAF1 8007 83 + 1 ACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGCUACCA---------------------CACGCCCCAUGCCCCCAU--------GCCACCAUGCC-------- ...(((..((.((((((((((((....)))...)))))).))).))(((.....---------------------...)))...)))...(((--------(....))))..-------- ( -21.70) >DroSim_CAF1 7789 72 + 1 ACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGCUACCA---------------------CACGCCCCAUGCCACCAU--------------------------- ...(((..((.((((((((((((....)))...)))))).))).))(((.....---------------------...)))...)))......--------------------------- ( -20.60) >DroEre_CAF1 5595 99 + 1 ACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGAAUAUCC---------------------CACACCCCAUGCCACCAUGCCACCAUGCAACCAUGCCUCCAUGCC ...(((.(((.((((((((((((....)))...)))))).))).))).......---------------------.......((((....((((....))))....))))......))). ( -24.50) >DroYak_CAF1 6578 120 + 1 ACUGCCACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGAUACCCCAAGCCCCACAUCCCACAUCGCACACUCCAUGCCCCCAAGCCCCCAAGCUACCAUGCCACCAUGCC .......(((.((((((((((((....)))...)))))).))).)))...........................(((.....((((......(((......)))..))))......))). ( -22.50) >consensus ACUGCAACUCGAUCAGGGCUUCGACUUCGACCCAGCCCUAGAUAGAGGCUACCA_____________________CACACCCCAUGCCACCAUGCC_CCA_GCCACCAUGCC_C______ .......(((.((((((((((((....)))...)))))).))).)))......................................................................... (-18.26 = -18.26 + -0.00)


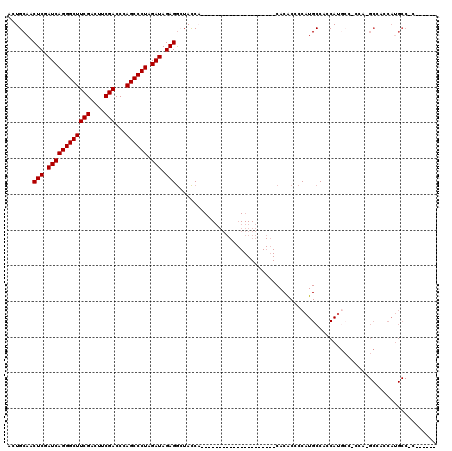
| Location | 18,085,635 – 18,085,748 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.16 |
| Mean single sequence MFE | -34.64 |
| Consensus MFE | -24.62 |
| Energy contribution | -24.92 |
| Covariance contribution | 0.30 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.17 |
| Structure conservation index | 0.71 |
| SVM decision value | 2.63 |
| SVM RNA-class probability | 0.995887 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18085635 113 + 22224390 CCCAUGCCUCCAUGCCCCCAUGCCGCCAUGCCAC------ACACGCC-CCAUGCCACUCCAGUUUGAGGCAGCCUGAUCAACUUGUUGUGGCUUUGGUGGCAUGCAAUGACCACCAUCAA ..((((....))))....((((((((((.(((((------....(((-.((.((.......)).)).)))(((..(.....)..))))))))..))))))))))................ ( -34.20) >DroSec_CAF1 8066 103 + 1 CCCAUGCCCCCAU--------GCCACCAUGCC--------ACAUGCC-CCAUGCCACUCCAGUUUGAGGCAGCCUGAUCAACUUGUUGUGGCUUUGGUGGCAUGCAAUGACCACCAUCAA ..(((.....(((--------(((((((.(((--------(((((((-.((.((.......)).)).))))((..(.....)..))))))))..))))))))))..)))........... ( -34.30) >DroSim_CAF1 7848 87 + 1 CCCAUGCCACCAU-------------------------------GCC-UCAUGCCACUCCAGUUUGAGG-AGCCUGAUCAACUUGUUGUGGCUUUGGUGGCAUGCAAUGACCACCAUCAA ..((((((((((.-------------------------------.((-(((.((.......)).)))))-((((....(((....))).)))).))))))))))................ ( -31.00) >DroEre_CAF1 5654 120 + 1 CCCAUGCCACCAUGCCACCAUGCAACCAUGCCUCCAUGCCCCAUGCCACCAUGCCACUCCAGUUUGAGGCAGCCUGAUCAACUUGUUGUGGCUUUGGUGGCAUGCAAUGACCACCAUCAA ..((((....((((....))))....))))....(((....((((((((((.(((((..(((.((((..(.....).)))).)))..)))))..))))))))))..)))........... ( -38.10) >DroYak_CAF1 6658 119 + 1 UCCAUGCCCCCAAGCCCCCAAGCUACCAUGCCACCAUGCCACAUGCC-CCAUGCCACUCCAGUUUGCGGCAGCCUGAUCAACUUGUUGUGGCUUUGGUGGCAUGCAAUGACCACCAUCAA ..(((.......(((......)))..((((((((((.((((((((((-.((.((.......)).)).))))((..(.....)..))))))))..))))))))))..)))........... ( -35.60) >consensus CCCAUGCCACCAUGCC_CCA_GCCACCAUGCC_C______ACAUGCC_CCAUGCCACUCCAGUUUGAGGCAGCCUGAUCAACUUGUUGUGGCUUUGGUGGCAUGCAAUGACCACCAUCAA ..((((((.(((.........((......)).....................(((((..(((.((((..........)))).)))..)))))..))).))))))................ (-24.62 = -24.92 + 0.30)



| Location | 18,085,635 – 18,085,748 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.16 |
| Mean single sequence MFE | -42.62 |
| Consensus MFE | -27.93 |
| Energy contribution | -28.36 |
| Covariance contribution | 0.43 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939212 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18085635 113 - 22224390 UUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGUUGAUCAGGCUGCCUCAAACUGGAGUGGCAUGG-GGCGUGU------GUGGCAUGGCGGCAUGGGGGCAUGGAGGCAUGGG .((((....))))((.((((((.(((..((((((((.((((.....))))((((((..((.....))..)))-))))).)------))))).))).)))))).)).((((....)))).. ( -40.20) >DroSec_CAF1 8066 103 - 1 UUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGUUGAUCAGGCUGCCUCAAACUGGAGUGGCAUGG-GGCAUGU--------GGCAUGGUGGC--------AUGGGGGCAUGGG ....(.(((.((....((((((((((..((((((...((((.....))))((((((..((.....))..)))-))).)))--------))).)))))))--------))).)).))).). ( -42.60) >DroSim_CAF1 7848 87 - 1 UUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGUUGAUCAGGCU-CCUCAAACUGGAGUGGCAUGA-GGC-------------------------------AUGGUGGCAUGGG .((((....))))...((((((((((..(((.(((....))).(((.((.-.(((......)))..)).)))-)))-------------------------------.)))))))))).. ( -34.90) >DroEre_CAF1 5654 120 - 1 UUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGUUGAUCAGGCUGCCUCAAACUGGAGUGGCAUGGUGGCAUGGGGCAUGGAGGCAUGGUUGCAUGGUGGCAUGGUGGCAUGGG .((((....))))...((((((((((..(((((.........((((.((..(..(.....)..)..)).))))..((((.((((((....))).))).))))))))).)))))))))).. ( -47.10) >DroYak_CAF1 6658 119 - 1 UUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGUUGAUCAGGCUGCCGCAAACUGGAGUGGCAUGG-GGCAUGUGGCAUGGUGGCAUGGUAGCUUGGGGGCUUGGGGGCAUGGA ....(((((((........)))))))..(((....((((((..(((((((((((...((((..((.(((((.-..))))).)).))))....))))))))))).))))))..)))..... ( -48.30) >consensus UUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGUUGAUCAGGCUGCCUCAAACUGGAGUGGCAUGG_GGCAUGU______G_GGCAUGGUGGC_UGG_GGCAUGGGGGCAUGGG .((((....))))...((((((.(((..(((((..((..((((...((...))))))..))..))))).......((((...........))))..............))).)))))).. (-27.93 = -28.36 + 0.43)

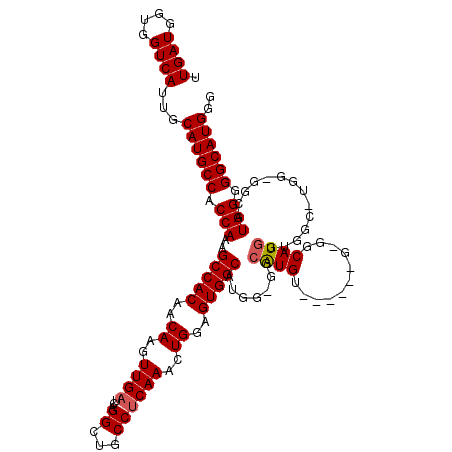

| Location | 18,085,708 – 18,085,824 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.36 |
| Mean single sequence MFE | -27.34 |
| Consensus MFE | -23.40 |
| Energy contribution | -23.80 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.584505 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18085708 116 - 22224390 ACAAAACACAGGCAGCGCAUGCAACACACACAACUUGGAC----AACAGAACACAGCGACGACGUGAUCUCAGUGCGGCUUUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGU ...........(((.....)))..........(((((...----...(((.(((.........))).)))...((.(((((...(((((((........)))))))))))).)).))))) ( -28.10) >DroSec_CAF1 8129 107 - 1 ACAAAACACAGGCAGCACAUGCAACACAUAC-------------AACAGAACACAGCGACGACGUGAUCUCAGUGCGGCAUUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGU ..........(((..((.((((..(((....-------------...(((.(((.........))).)))..)))..)))))).(((((((........)))))))..)))......... ( -25.70) >DroSim_CAF1 7895 116 - 1 ACAAAACACAGGCAGCACAUGCAACACACACAACUUGGAC----AACAGAACACAGCGACGACGUGAUCUCAGUGCGGCUUUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGU ...........(((.....)))..........(((((...----...(((.(((.........))).)))...((.(((((...(((((((........)))))))))))).)).))))) ( -28.10) >DroEre_CAF1 5734 114 - 1 ACAAAACACAGGCAGCACAUGCAACACA--CAACUUGGAC----AACAGAACACAGCGACGACGUGAUCUCAGUGCGGCUUUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGU ...........(((.....)))......--..(((((...----...(((.(((.........))).)))...((.(((((...(((((((........)))))))))))).)).))))) ( -28.10) >DroYak_CAF1 6737 118 - 1 ACAAAACACAGGCAGCACAUGCAACACA--CAACUUGGACACACAACAAAACACAGCGACGACGUGAUCUCAGUGCGGCUUUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGU ...........(((.....)))......--..(((((..................(((..((.......))..)))(((((...(((((((........))))))))))))....))))) ( -26.70) >consensus ACAAAACACAGGCAGCACAUGCAACACA_ACAACUUGGAC____AACAGAACACAGCGACGACGUGAUCUCAGUGCGGCUUUGAUGGUGGUCAUUGCAUGCCACCAAAGCCACAACAAGU ......(((..(((.....))).........................(((.(((.........))).)))..))).(((((...(((((((........))))))))))))......... (-23.40 = -23.80 + 0.40)

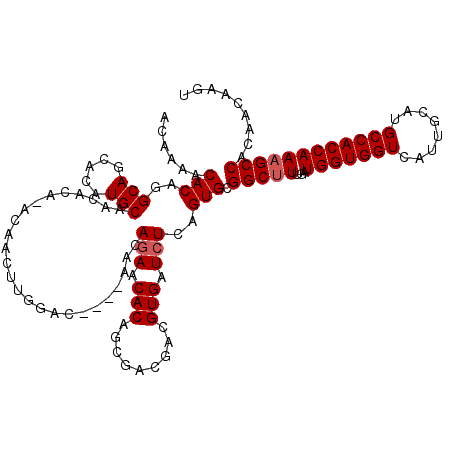

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:34:02 2006