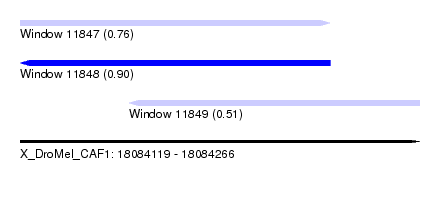

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,084,119 – 18,084,266 |
| Length | 147 |
| Max. P | 0.902757 |

| Location | 18,084,119 – 18,084,233 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 96.49 |
| Mean single sequence MFE | -32.90 |
| Consensus MFE | -28.40 |
| Energy contribution | -28.84 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.759824 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18084119 114 + 22224390 GUACGCUCAAAGUCAAAGUCAAUUGGAUAUAGCAAAAACCUUGAGCUAACCCACUUUUCUGGGCUUUUGGCCAAAUGCAUGCUGCUUUCAACCAAAUGAGAGUGUACUUAUUUG ((((((((....(((.......(((((..(((((......(((.(((((((((......))))...)))))))).....)))))..))))).....)))))))))))....... ( -28.80) >DroSec_CAF1 6570 114 + 1 GUACGCUCAAAGUCAAAGUCAAUUGGACAUGGCAAAAACCUUGGGCUAACCCACUUUUCUGGGCUUUUGGCCAAAUGCAUGCUGCUUUCAGCCAAAUGAGAGUGUACUUACUUG ((((((((.........(((.....))).((((..........((((((((((......))))...))))))....((.....)).....)))).....))))))))....... ( -34.60) >DroSim_CAF1 6345 114 + 1 GUACGCUCAAAGUCAAAGUCAAUUGGACAUGGCAAAAACCUUGGGCUAACCCACUUUUCUGGGCUUUUUGCCAAAUGCAUGCUGCUUUCAGCCAAAUGAGAGUGUACUUACUUG ((((((((.........(((.....))).((((((((((((((((....)))).......))).))))))))).......((((....)))).......))))))))....... ( -35.31) >consensus GUACGCUCAAAGUCAAAGUCAAUUGGACAUGGCAAAAACCUUGGGCUAACCCACUUUUCUGGGCUUUUGGCCAAAUGCAUGCUGCUUUCAGCCAAAUGAGAGUGUACUUACUUG ((((((((..........(((.((((....((((.........((((((((((......))))...))))))..........)))).....)))).)))))))))))....... (-28.40 = -28.84 + 0.45)

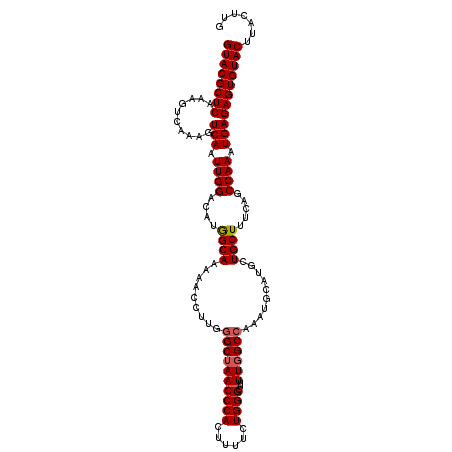

| Location | 18,084,119 – 18,084,233 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 96.49 |
| Mean single sequence MFE | -33.60 |
| Consensus MFE | -31.42 |
| Energy contribution | -31.53 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.902757 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18084119 114 - 22224390 CAAAUAAGUACACUCUCAUUUGGUUGAAAGCAGCAUGCAUUUGGCCAAAAGCCCAGAAAAGUGGGUUAGCUCAAGGUUUUUGCUAUAUCCAAUUGACUUUGACUUUGAGCGUAC .......((((.......((((((..((.((.....)).))..))))))((((((......)))))).((((((((((...(.((........)).)...)))))))))))))) ( -35.70) >DroSec_CAF1 6570 114 - 1 CAAGUAAGUACACUCUCAUUUGGCUGAAAGCAGCAUGCAUUUGGCCAAAAGCCCAGAAAAGUGGGUUAGCCCAAGGUUUUUGCCAUGUCCAAUUGACUUUGACUUUGAGCGUAC .......((((.......((((((..((.((.....)).))..))))))((((((......)))))).((.(((((((...(.((........)).)...))))))).)))))) ( -34.20) >DroSim_CAF1 6345 114 - 1 CAAGUAAGUACACUCUCAUUUGGCUGAAAGCAGCAUGCAUUUGGCAAAAAGCCCAGAAAAGUGGGUUAGCCCAAGGUUUUUGCCAUGUCCAAUUGACUUUGACUUUGAGCGUAC .......((((.((((((.(((((((....))))..(((..((((((((((((((......)))))).(((...)))))))))))))).))).)))..........))).)))) ( -30.90) >consensus CAAGUAAGUACACUCUCAUUUGGCUGAAAGCAGCAUGCAUUUGGCCAAAAGCCCAGAAAAGUGGGUUAGCCCAAGGUUUUUGCCAUGUCCAAUUGACUUUGACUUUGAGCGUAC .......((((.(((...((((((..((.((.....)).))..))))))((((((......))))))....(((((((.(((.......)))..))))))).....))).)))) (-31.42 = -31.53 + 0.11)


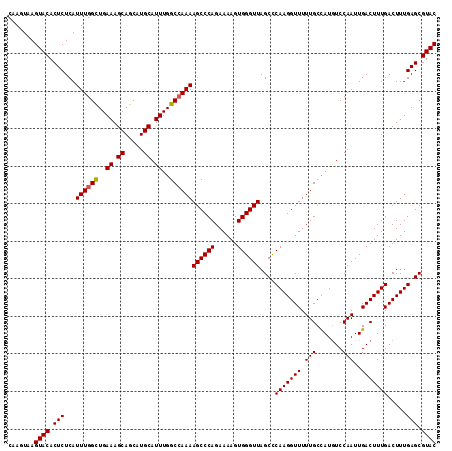
| Location | 18,084,159 – 18,084,266 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 91.20 |
| Mean single sequence MFE | -25.40 |
| Consensus MFE | -22.14 |
| Energy contribution | -22.03 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.87 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.508555 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18084159 107 - 22224390 UAUGAUAG-------ACCAUAGACCUAACAUCUGACAAUACAAAUAAGUACACUCUCAUUUGGUUGAAAGCAGCAUGCAUUUGGCCAAAAGCCCAGAAAAGUGGGUUAGCUCAA ((((....-------..))))((((((...((((....(((......)))........((((((..((.((.....)).))..))))))....))))....))))))....... ( -24.20) >DroSec_CAF1 6610 113 - 1 -AUGAUAGACCAUAGACCAUAGACCUACCAUCUGACAAUCCAAGUAAGUACACUCUCAUUUGGCUGAAAGCAGCAUGCAUUUGGCCAAAAGCCCAGAAAAGUGGGUUAGCCCAA -....................(((((((..((((.........(..((....))..).((((((..((.((.....)).))..))))))....))))...)))))))....... ( -26.70) >DroSim_CAF1 6385 113 - 1 -AUGAUAGACCAUAGACCAUAGACCUACUAUCUGACAAUCCAAGUAAGUACACUCUCAUUUGGCUGAAAGCAGCAUGCAUUUGGCAAAAAGCCCAGAAAAGUGGGUUAGCCCAA -....................((((((((.((((......(((((.((......)).)))))((..((.((.....)).))..))........))))..))))))))....... ( -25.30) >consensus _AUGAUAGACCAUAGACCAUAGACCUACCAUCUGACAAUCCAAGUAAGUACACUCUCAUUUGGCUGAAAGCAGCAUGCAUUUGGCCAAAAGCCCAGAAAAGUGGGUUAGCCCAA .....................(((((((..((((......(((((.((......)).)))))((..((.((.....)).))..))........))))...)))))))....... (-22.14 = -22.03 + -0.11)

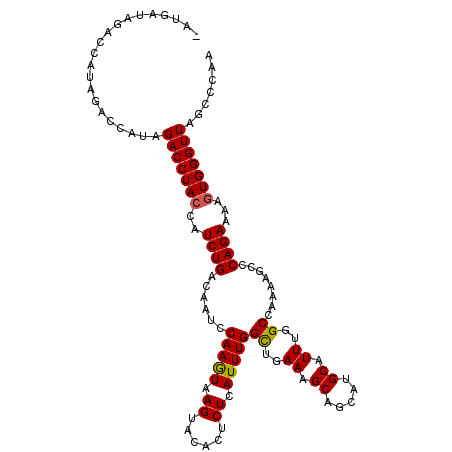

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:33:53 2006