
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,070,791 – 18,071,031 |
| Length | 240 |
| Max. P | 0.989414 |

| Location | 18,070,791 – 18,070,911 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.88 |
| Mean single sequence MFE | -23.30 |
| Consensus MFE | -22.70 |
| Energy contribution | -22.70 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.636955 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18070791 120 + 22224390 AACACAAAAAAAUGGAUAAAAAACAAGGGAGAAAAUAGAAAAAUAGGACAAAAAUAUCAUGUUGGAAACCAGAGCUGCUCGCUCCGGAGUGACCGGUGCAGGUACGAGUGAGUAUCUGCG ............(((......((((.((............................)).)))).....)))..((((.(((((....))))).))))((((((((......)))))))). ( -24.09) >DroSec_CAF1 8460 117 + 1 AACUCAAAAAAAUUGAUAAAAAAU---GAAGAAAAUAGAAAAAUAGGACAAAAAUAUCAUGUUGGAAACCAGAGCUGCUCGCUCCGGAGUGACCGGUGCAGGUACGAGUGAGUAUCUGCG ..(((...................---...................((((.........))))(....)..)))(((.(((((....))))).))).((((((((......)))))))). ( -23.10) >DroSim_CAF1 13057 109 + 1 --------AAAAUUGAUAAAAAAU---GAAGAAAAUAGAAAAAUAGGACAAAAAUAUCAUGUUGGAAACCAGAGCUGCUCGCUCCGGAGUGACCGGUGCAGGUACGAGUGAGUAUCUGCG --------................---..................................((((...)))).((((.(((((....))))).))))((((((((......)))))))). ( -22.70) >consensus AAC_CAAAAAAAUUGAUAAAAAAU___GAAGAAAAUAGAAAAAUAGGACAAAAAUAUCAUGUUGGAAACCAGAGCUGCUCGCUCCGGAGUGACCGGUGCAGGUACGAGUGAGUAUCUGCG .............................................................((((...)))).((((.(((((....))))).))))((((((((......)))))))). (-22.70 = -22.70 + -0.00)
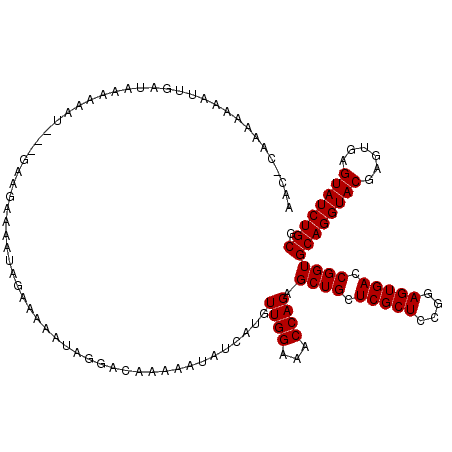


| Location | 18,070,791 – 18,070,911 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.88 |
| Mean single sequence MFE | -24.37 |
| Consensus MFE | -23.80 |
| Energy contribution | -23.80 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.98 |
| SVM decision value | 2.16 |
| SVM RNA-class probability | 0.989414 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18070791 120 - 22224390 CGCAGAUACUCACUCGUACCUGCACCGGUCACUCCGGAGCGAGCAGCUCUGGUUUCCAACAUGAUAUUUUUGUCCUAUUUUUCUAUUUUCUCCCUUGUUUUUUAUCCAUUUUUUUGUGUU .((((.(((......))).))))...((.....(((((((.....)))))))...)).....((((....)))).............................................. ( -23.80) >DroSec_CAF1 8460 117 - 1 CGCAGAUACUCACUCGUACCUGCACCGGUCACUCCGGAGCGAGCAGCUCUGGUUUCCAACAUGAUAUUUUUGUCCUAUUUUUCUAUUUUCUUC---AUUUUUUAUCAAUUUUUUUGAGUU .((((.(((......))).))))...((.....(((((((.....)))))))...))(((..((((....))))...................---........((((.....))))))) ( -25.50) >DroSim_CAF1 13057 109 - 1 CGCAGAUACUCACUCGUACCUGCACCGGUCACUCCGGAGCGAGCAGCUCUGGUUUCCAACAUGAUAUUUUUGUCCUAUUUUUCUAUUUUCUUC---AUUUUUUAUCAAUUUU-------- .((((.(((......))).))))...((.....(((((((.....)))))))...)).....((((....))))...................---................-------- ( -23.80) >consensus CGCAGAUACUCACUCGUACCUGCACCGGUCACUCCGGAGCGAGCAGCUCUGGUUUCCAACAUGAUAUUUUUGUCCUAUUUUUCUAUUUUCUUC___AUUUUUUAUCAAUUUUUUUG_GUU .((((.(((......))).))))...((.....(((((((.....)))))))...)).....((((....)))).............................................. (-23.80 = -23.80 + 0.00)



| Location | 18,070,831 – 18,070,951 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 100.00 |
| Mean single sequence MFE | -32.60 |
| Consensus MFE | -32.60 |
| Energy contribution | -32.60 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.92 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.881664 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18070831 120 - 22224390 AAUGAAUAUUUUAUAGAAAAAUGUGCAAUGUUCGCAAGGCCGCAGAUACUCACUCGUACCUGCACCGGUCACUCCGGAGCGAGCAGCUCUGGUUUCCAACAUGAUAUUUUUGUCCUAUUU ...(((((((.....((((...(.((..(((((((......((((.(((......))).)))).((((.....)))).))))))))).)...))))......)))))))........... ( -32.60) >DroSec_CAF1 8497 120 - 1 AAUGAAUAUUUUAUAGAAAAAUGUGCAAUGUUCGCAAGGCCGCAGAUACUCACUCGUACCUGCACCGGUCACUCCGGAGCGAGCAGCUCUGGUUUCCAACAUGAUAUUUUUGUCCUAUUU ...(((((((.....((((...(.((..(((((((......((((.(((......))).)))).((((.....)))).))))))))).)...))))......)))))))........... ( -32.60) >DroSim_CAF1 13086 120 - 1 AAUGAAUAUUUUAUAGAAAAAUGUGCAAUGUUCGCAAGGCCGCAGAUACUCACUCGUACCUGCACCGGUCACUCCGGAGCGAGCAGCUCUGGUUUCCAACAUGAUAUUUUUGUCCUAUUU ...(((((((.....((((...(.((..(((((((......((((.(((......))).)))).((((.....)))).))))))))).)...))))......)))))))........... ( -32.60) >consensus AAUGAAUAUUUUAUAGAAAAAUGUGCAAUGUUCGCAAGGCCGCAGAUACUCACUCGUACCUGCACCGGUCACUCCGGAGCGAGCAGCUCUGGUUUCCAACAUGAUAUUUUUGUCCUAUUU ...(((((((.....((((...(.((..(((((((......((((.(((......))).)))).((((.....)))).))))))))).)...))))......)))))))........... (-32.60 = -32.60 + 0.00)



| Location | 18,070,911 – 18,071,031 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 100.00 |
| Mean single sequence MFE | -32.90 |
| Consensus MFE | -32.90 |
| Energy contribution | -32.90 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.34 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.543349 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18070911 120 + 22224390 GCCUUGCGAACAUUGCACAUUUUUCUAUAAAAUAUUCAUUAACCCGAUAUUCAUUAAAAUGACGCCGCCUCGUUAGGGUUAAUGGCGCCAGUGGCCCCCAGUCGGUCGCCAAGCGCUCCG ....(((((...)))))...................((((((((((.....).......(((((......))))))))))))))((((...((((..((....))..)))).)))).... ( -32.90) >DroSec_CAF1 8577 120 + 1 GCCUUGCGAACAUUGCACAUUUUUCUAUAAAAUAUUCAUUAACCCGAUAUUCAUUAAAAUGACGCCGCCUCGUUAGGGUUAAUGGCGCCAGUGGCCCCCAGUCGGUCGCCAAGCGCUCCG ....(((((...)))))...................((((((((((.....).......(((((......))))))))))))))((((...((((..((....))..)))).)))).... ( -32.90) >DroSim_CAF1 13166 120 + 1 GCCUUGCGAACAUUGCACAUUUUUCUAUAAAAUAUUCAUUAACCCGAUAUUCAUUAAAAUGACGCCGCCUCGUUAGGGUUAAUGGCGCCAGUGGCCCCCAGUCGGUCGCCAAGCGCUCCG ....(((((...)))))...................((((((((((.....).......(((((......))))))))))))))((((...((((..((....))..)))).)))).... ( -32.90) >consensus GCCUUGCGAACAUUGCACAUUUUUCUAUAAAAUAUUCAUUAACCCGAUAUUCAUUAAAAUGACGCCGCCUCGUUAGGGUUAAUGGCGCCAGUGGCCCCCAGUCGGUCGCCAAGCGCUCCG ....(((((...)))))...................((((((((((.....).......(((((......))))))))))))))((((...((((..((....))..)))).)))).... (-32.90 = -32.90 + -0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:33:40 2006