
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,064,282 – 18,064,442 |
| Length | 160 |
| Max. P | 0.670746 |

| Location | 18,064,282 – 18,064,402 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.43 |
| Mean single sequence MFE | -31.00 |
| Consensus MFE | -27.62 |
| Energy contribution | -27.62 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.670746 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
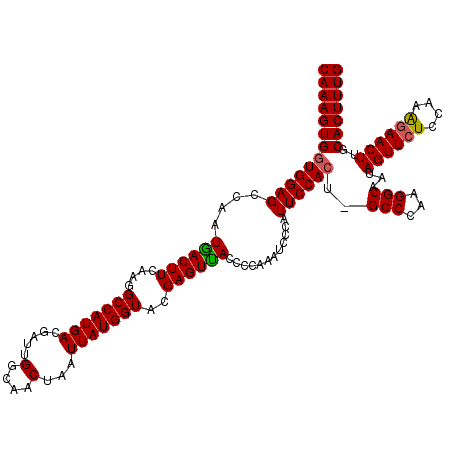
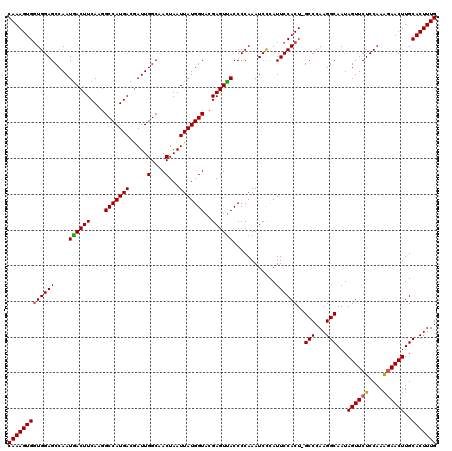
>X_DroMel_CAF1 18064282 120 + 22224390 CAAAGUGUUGGAGCCAAUAACUUCAAGGCCAUGAAAAUUGGCAACUAAUUAUGGUACGAGUUACCCCAAAUCUCAUUCCACUAGCCCAAGGCAAUAGUUCUCCAAAUAACUUGCACUUUG ((((((((((((((((((...((((......)))).))))))(((((....(((...(((...........)))...)))...(((...)))..))))).))))........)))))))) ( -28.70) >DroSec_CAF1 2177 119 + 1 CAAAGUGGUGGAGCCAAUGACUUCAAGGCCAUGACGAUUGGCAACUAAUUAUGGUACGAGUUACCCCAAAUCCCAUUCCACU-GCCCAAGGCAAUAGUUCUCCAAAGAACUUGCACUUUG ((((((((((((((((((..(.(((......))).)))))))..........((((.....))))...........)))))(-(((...))))..((((((....))))))..))))))) ( -31.40) >DroSim_CAF1 5270 119 + 1 CAAAGUGGUGGAGCCAAUGACUUCAAGGCCAUGACGAUUGGCAACUAAUUAUGGUACGAGUCACCCCAAAUCCUAUUCCACU-GCCCAAGGCAAUAGUUCUCCAAGGAACUUGCACUUUG (((((((((((((....((((((....(((((((.....(....)...)))))))..))))))............))))))(-(((...))))..((((((....))))))..))))))) ( -32.89) >consensus CAAAGUGGUGGAGCCAAUGACUUCAAGGCCAUGACGAUUGGCAACUAAUUAUGGUACGAGUUACCCCAAAUCCCAUUCCACU_GCCCAAGGCAAUAGUUCUCCAAAGAACUUGCACUUUG (((((((((((((....((((((....(((((((.....(....)...)))))))..))))))............))))))..(((...)))...((((((....))))))..))))))) (-27.62 = -27.62 + 0.00)

| Location | 18,064,322 – 18,064,442 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.10 |
| Mean single sequence MFE | -27.67 |
| Consensus MFE | -25.94 |
| Energy contribution | -25.83 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.94 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.505520 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18064322 120 + 22224390 GCAACUAAUUAUGGUACGAGUUACCCCAAAUCUCAUUCCACUAGCCCAAGGCAAUAGUUCUCCAAAUAACUUGCACUUUGUGCUACUAAAAAGCGGAUUGCAUCUGCAAUCUUGGCAGCC ((.........(((...(((...........)))...))).((((.(((((....((((........))))....))))).)))).......(((((((((....)))))))..)).)). ( -24.50) >DroSec_CAF1 2217 119 + 1 GCAACUAAUUAUGGUACGAGUUACCCCAAAUCCCAUUCCACU-GCCCAAGGCAAUAGUUCUCCAAAGAACUUGCACUUUGUGCUACUAAAAAGCGGAUUGCAUCUGCAAUCUUGGCAGCC ............((((.....))))...............((-((((((((....((((((....))))))....))))).(((.......)))(((((((....))))))).))))).. ( -28.30) >DroSim_CAF1 5310 119 + 1 GCAACUAAUUAUGGUACGAGUCACCCCAAAUCCUAUUCCACU-GCCCAAGGCAAUAGUUCUCCAAGGAACUUGCACUUUGUGCCACUAAAAAGCGGAUUGCAUCUGCAAUCUUGGCAGCC ((.........((((((((((((..................(-(((...))))..((((((....)))))))).)).)))))))).......(((((((((....)))))))..)).)). ( -30.20) >consensus GCAACUAAUUAUGGUACGAGUUACCCCAAAUCCCAUUCCACU_GCCCAAGGCAAUAGUUCUCCAAAGAACUUGCACUUUGUGCUACUAAAAAGCGGAUUGCAUCUGCAAUCUUGGCAGCC ((.........(((((((((.......................(((...)))...((((((....)))))).....))))))))).......(((((((((....)))))))..)).)). (-25.94 = -25.83 + -0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:33:18 2006