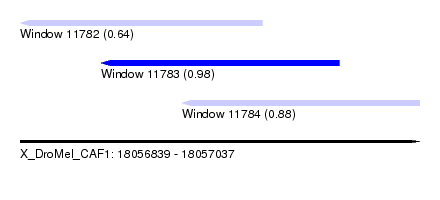
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,056,839 – 18,057,037 |
| Length | 198 |
| Max. P | 0.982586 |

| Location | 18,056,839 – 18,056,959 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -26.67 |
| Consensus MFE | -24.80 |
| Energy contribution | -24.80 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.644204 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18056839 120 - 22224390 ACUGUCCUUGGAUUCUUGCCUUCUCCAGGAUUCUGGGAAUCUACUUAAACUUUGUCUUGAGUUGAUUUCCUUUUACAGUUUGCUCAUUCUUGUUGCUGGAAAAACUUAAUUUGCUGUCUG ...(((((.(((...........))))))))...(((((((.((((((........)))))).)))))))((((.((((..((........)).)))).))))................. ( -24.60) >DroSec_CAF1 1504 120 - 1 ACUGUCCUUGGAUCCUCUCCUUCUCCAGGAUACUGGGAAUCUACUUAAACUUUGUCUUGAGUUGAUUUCCUUUUACAGUUUGCUCAUUCUUGUUGCUGGAAAAACUUAAUUUGCUGUCUG ..((((((.(((...........)))))))))..(((((((.((((((........)))))).)))))))((((.((((..((........)).)))).))))................. ( -25.90) >DroSim_CAF1 1522 120 - 1 ACUGUCCUGGGAUUCUUGCCAUCUCCAGGAUUCUGGGAAUCUACUUAAACUUUGUCUUGAGUUGAUUUCCUUUUACAGUUUGCUCAUUCUUGUUGCUGGAAAAACUUAAUUUGCUGUCUG ...((((((((((.......))).)))))))...(((((((.((((((........)))))).)))))))((((.((((..((........)).)))).))))................. ( -29.50) >consensus ACUGUCCUUGGAUUCUUGCCUUCUCCAGGAUUCUGGGAAUCUACUUAAACUUUGUCUUGAGUUGAUUUCCUUUUACAGUUUGCUCAUUCUUGUUGCUGGAAAAACUUAAUUUGCUGUCUG ...(((((.(((...........))))))))...(((((((.((((((........)))))).)))))))((((.((((..((........)).)))).))))................. (-24.80 = -24.80 + 0.00)

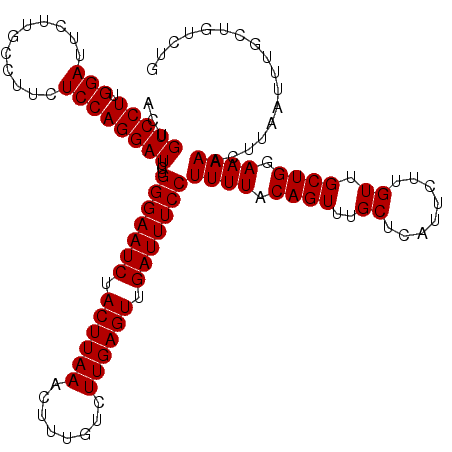

| Location | 18,056,879 – 18,056,997 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.02 |
| Mean single sequence MFE | -23.73 |
| Consensus MFE | -21.60 |
| Energy contribution | -21.60 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.66 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.92 |
| SVM RNA-class probability | 0.982586 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18056879 118 - 22224390 ACACAUA--UAUUUUAAACACACUUUACCCAUUUAUAUCCACUGUCCUUGGAUUCUUGCCUUCUCCAGGAUUCUGGGAAUCUACUUAAACUUUGUCUUGAGUUGAUUUCCUUUUACAGUU .......--...............................(((((....(((.(((((.......))))).)))(((((((.((((((........)))))).)))))))....))))). ( -22.40) >DroSec_CAF1 1544 118 - 1 ACACAUA--UAUUUUAAAUACAAUUUUCCCAUUUAUACCCACUGUCCUUGGAUCCUCUCCUUCUCCAGGAUACUGGGAAUCUACUUAAACUUUGUCUUGAGUUGAUUUCCUUUUACAGUU .......--...............................(((((.((((((...........)))))).....(((((((.((((((........)))))).)))))))....))))). ( -22.30) >DroSim_CAF1 1562 120 - 1 ACACAUAUAUAUUUUAAAUACAAUUUCCCCAUUUAUACCCACUGUCCUGGGAUUCUUGCCAUCUCCAGGAUUCUGGGAAUCUACUUAAACUUUGUCUUGAGUUGAUUUCCUUUUACAGUU ........................................(((((((((((((.......))).))))))....(((((((.((((((........)))))).))))))).....)))). ( -26.50) >consensus ACACAUA__UAUUUUAAAUACAAUUUACCCAUUUAUACCCACUGUCCUUGGAUUCUUGCCUUCUCCAGGAUUCUGGGAAUCUACUUAAACUUUGUCUUGAGUUGAUUUCCUUUUACAGUU ........................................((((((((.(((...........)))))))....(((((((.((((((........)))))).))))))).....)))). (-21.60 = -21.60 + -0.00)


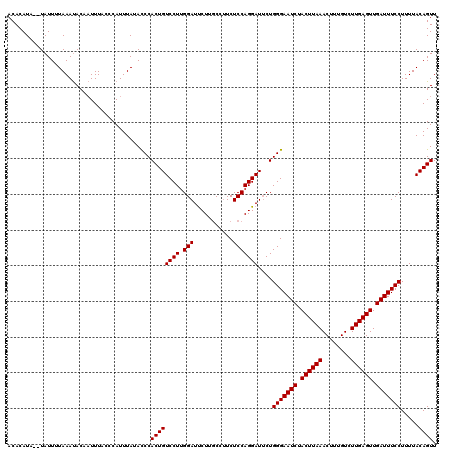
| Location | 18,056,919 – 18,057,037 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.58 |
| Mean single sequence MFE | -18.99 |
| Consensus MFE | -16.05 |
| Energy contribution | -16.17 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.882292 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18056919 118 - 22224390 UAUGUACCCUACUACUCCAUCUUCUUGUUAUACACGAAACACACAUA--UAUUUUAAACACACUUUACCCAUUUAUAUCCACUGUCCUUGGAUUCUUGCCUUCUCCAGGAUUCUGGGAAU .....................(((.((.....)).))).........--............................((((..(((((.(((...........))))))))..))))... ( -13.80) >DroSec_CAF1 1584 115 - 1 UAUGUGCCAUA---CUCUAUCUUCUUGUUAUACACGAAACACACAUA--UAUUUUAAAUACAAUUUUCCCAUUUAUACCCACUGUCCUUGGAUCCUCUCCUUCUCCAGGAUACUGGGAAU ((((((.....---...........((((........))))))))))--............................((((.((((((.(((...........))))))))).))))... ( -19.59) >DroSim_CAF1 1602 117 - 1 UAUGUGCCCUA---CUCUAUCUUCUUGUUAUACACGAAACACACAUAUAUAUUUUAAAUACAAUUUCCCCAUUUAUACCCACUGUCCUGGGAUUCUUGCCAUCUCCAGGAUUCUGGGAAU ((((((.....---...........((((........))))))))))..............................((((..((((((((((.......))).)))))))..))))... ( -23.59) >consensus UAUGUGCCCUA___CUCUAUCUUCUUGUUAUACACGAAACACACAUA__UAUUUUAAAUACAAUUUACCCAUUUAUACCCACUGUCCUUGGAUUCUUGCCUUCUCCAGGAUUCUGGGAAU ((((((...................((((........))))))))))..............................((((..(((((.(((...........))))))))..))))... (-16.05 = -16.17 + 0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:32:56 2006