
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,047,453 – 18,047,613 |
| Length | 160 |
| Max. P | 0.792107 |

| Location | 18,047,453 – 18,047,573 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.78 |
| Mean single sequence MFE | -40.40 |
| Consensus MFE | -38.22 |
| Energy contribution | -38.80 |
| Covariance contribution | 0.58 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.792107 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18047453 120 + 22224390 UUCGCGGAUCGUCGUCCUUCAAAUAGCCGUGUCCAUGUGCAUGUCCAUGUCCAUGACCAUGUCCGUGCCCAUGUCCGUGACCAUGUCCCUGGGCAUGCGGACGACUGAGCACCAUCUUGC .....((.(((((((((........((.(((((((.(..((((..((((..((((..(((....)))..))))..))))..))))..).)))))))))))))))).))...))....... ( -43.60) >DroSec_CAF1 35315 108 + 1 UUCGCGGAUCGUCGUCCUUCAAAUAGCCGUGUCCAUGUGCAUGUCCAUGUCCAUGACCAUGU------------CCGUGACCAUGUCCCUGGGCAUGCGAACGACUGAGCACCAUCUUGC (((((((((....))))...........(((((((.(..((((..((((..(((....))).------------.))))..))))..).)))))))))))).......(((......))) ( -34.00) >DroSim_CAF1 36419 120 + 1 UUCGCGGAUCGUCGUCCUUCAAAUAGCCGUGUCCAUGUGCAUGUCCAUGUCCAUGACCAUGUCCGUGCCCAUGUCCGUGACCAUGUCCCUGGGCAUGCGGACGACUGAGCACCAUCUUGC .....((.(((((((((........((.(((((((.(..((((..((((..((((..(((....)))..))))..))))..))))..).)))))))))))))))).))...))....... ( -43.60) >consensus UUCGCGGAUCGUCGUCCUUCAAAUAGCCGUGUCCAUGUGCAUGUCCAUGUCCAUGACCAUGUCCGUGCCCAUGUCCGUGACCAUGUCCCUGGGCAUGCGGACGACUGAGCACCAUCUUGC .....((.(((((((((..........(((....))).(((((((((.(..((((..((((..(((....)))..))))..))))..).)))))))))))))))).))...))....... (-38.22 = -38.80 + 0.58)

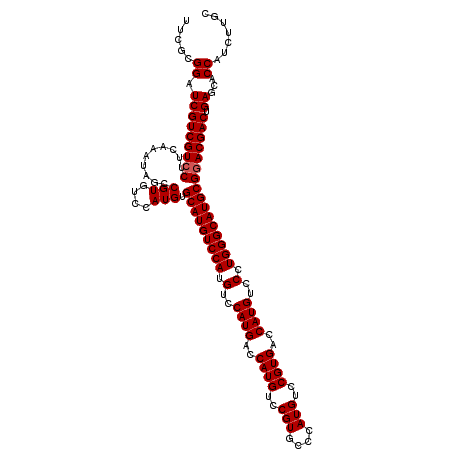

| Location | 18,047,493 – 18,047,613 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.78 |
| Mean single sequence MFE | -49.17 |
| Consensus MFE | -44.87 |
| Energy contribution | -44.73 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.669216 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18047493 120 - 22224390 CGCAUCAUGAUGGUCUACAGGCGGACCUGCGAGCGCCGCCGCAAGAUGGUGCUCAGUCGUCCGCAUGCCCAGGGACAUGGUCACGGACAUGGGCACGGACAUGGUCAUGGACAUGGACAU ....(((((((.((((....((((((..((((((((((.(....).)))))))).)).)))))).((((((.(..(........)..).)))))).))))...))))))).......... ( -51.90) >DroSec_CAF1 35355 108 - 1 CGCAUCAUGAUGGUCUACAGGCGGACCUGCGAGCGCCGCCGCAAGAUGGUGCUCAGUCGUUCGCAUGCCCAGGGACAUGGUCACGG------------ACAUGGUCAUGGACAUGGACAU .((...((((((((((......)))))...((((((((.(....).)))))))).)))))..))(((.(((.(..((((..((...------------...))..))))..).))).))) ( -43.70) >DroSim_CAF1 36459 120 - 1 CGCAUCAUGAUGGUCUACAGGCGGACCUGCGAGCGCCGCCGCAAGAUGGUGCUCAGUCGUCCGCAUGCCCAGGGACAUGGUCACGGACAUGGGCACGGACAUGGUCAUGGACAUGGACAU ....(((((((.((((....((((((..((((((((((.(....).)))))))).)).)))))).((((((.(..(........)..).)))))).))))...))))))).......... ( -51.90) >consensus CGCAUCAUGAUGGUCUACAGGCGGACCUGCGAGCGCCGCCGCAAGAUGGUGCUCAGUCGUCCGCAUGCCCAGGGACAUGGUCACGGACAUGGGCACGGACAUGGUCAUGGACAUGGACAU .((((.....((.....)).((((((..((((((((((.(....).)))))))).)).))))))))))(((.(..((((..((.(..(........)..).))..))))..).))).... (-44.87 = -44.73 + -0.14)


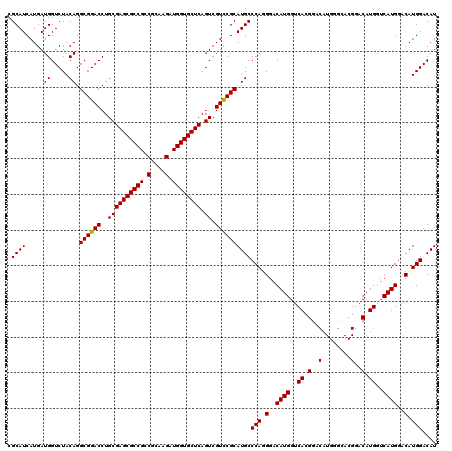
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:32:48 2006