
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,985,945 – 17,986,055 |
| Length | 110 |
| Max. P | 0.980173 |

| Location | 17,985,945 – 17,986,055 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.51 |
| Mean single sequence MFE | -35.55 |
| Consensus MFE | -18.81 |
| Energy contribution | -18.70 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.53 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.523671 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
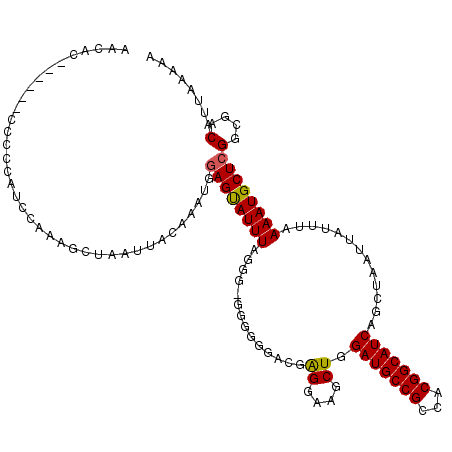
>X_DroMel_CAF1 17985945 110 + 22224390 CAAAG------UCCCCGUCCAAAACUAAUUACAAAUGUAGCAUUUAGA----AGGGACUGGGAAGCUGGAUGCCGCCACGGCAUCAGCUAAUUAUUUAAAAUGCUCGGCGACAUUAAAAA ....(------(((((((((....((((((((....))))...)))).----..)))).)))..(((((((((((...)))))))(((..............))))))))))........ ( -33.14) >DroEre_CAF1 17654 114 + 1 AACAC------CCCCCAUCCACAGCUAAUUACAAAUGGAGUAUUUUGGGCGGGGGUACGAGGAAGCUGGAUGCCGCCACGGCAUCAGCUAAUUAUUUAAAAUGCUCGGCGACAUUAAAAA .....------(((((..(((..(((............)))....)))..)))))..((((..(((((.((((((...))))))))))).(((......))).))))............. ( -32.80) >DroYak_CAF1 16209 120 + 1 AUCCCCCCCUCCCCCAAGCCAAAGCUAAUUACGAGUGGAGUAUUUAGGGAGGGGGGAUGAGGGAACUGGAUGCCGCCACGGCAUCAGCUAAUUAUUUAAAAUGCUCGGCGACAUUAAAAA ...((((((((((...(((....)))...(((.......)))....))))))))))...((....)).((((.((((..(((((....(((....)))..))))).)))).))))..... ( -40.70) >consensus AACAC______CCCCCAUCCAAAGCUAAUUACAAAUGGAGUAUUUAGGG_GGGGGGACGAGGAAGCUGGAUGCCGCCACGGCAUCAGCUAAUUAUUUAAAAUGCUCGGCGACAUUAAAAA .....................................((((((((..............((....)).(((((((...))))))).............))))))))(....)........ (-18.81 = -18.70 + -0.11)


| Location | 17,985,945 – 17,986,055 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.51 |
| Mean single sequence MFE | -37.92 |
| Consensus MFE | -25.77 |
| Energy contribution | -25.44 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.68 |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.980173 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
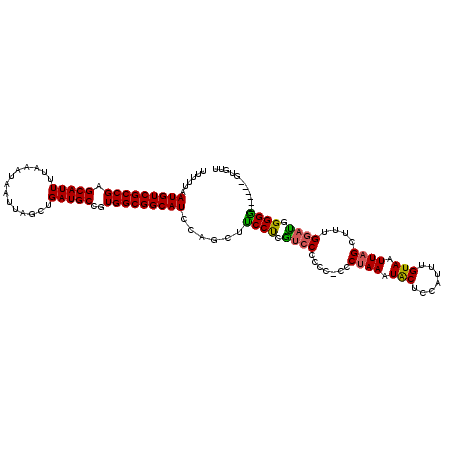
>X_DroMel_CAF1 17985945 110 - 22224390 UUUUUAAUGUCGCCGAGCAUUUUAAAUAAUUAGCUGAUGCCGUGGCGGCAUCCAGCUUCCCAGUCCCU----UCUAAAUGCUACAUUUGUAAUUAGUUUUGGACGGGGA------CUUUG ......(((((((((.(((((..............)))))..))))))))).(((..((((.((((..----.((((.(((.......))).))))....)))).))))------..))) ( -34.54) >DroEre_CAF1 17654 114 - 1 UUUUUAAUGUCGCCGAGCAUUUUAAAUAAUUAGCUGAUGCCGUGGCGGCAUCCAGCUUCCUCGUACCCCCGCCCAAAAUACUCCAUUUGUAAUUAGCUGUGGAUGGGGG------GUGUU ..........(((((.((.............(((((((((((...)))))).))))).............)).)......(((((((..((......))..))))))))------))).. ( -33.27) >DroYak_CAF1 16209 120 - 1 UUUUUAAUGUCGCCGAGCAUUUUAAAUAAUUAGCUGAUGCCGUGGCGGCAUCCAGUUCCCUCAUCCCCCCUCCCUAAAUACUCCACUCGUAAUUAGCUUUGGCUUGGGGGAGGGGGGGAU ......(((((((((.(((((..............)))))..)))))))))............((((((((((((...(((.......)))...(((....)))..)))))))))))).. ( -45.94) >consensus UUUUUAAUGUCGCCGAGCAUUUUAAAUAAUUAGCUGAUGCCGUGGCGGCAUCCAGCUUCCUCGUCCCCCC_CCCUAAAUACUCCAUUUGUAAUUAGCUUUGGAUGGGGG______GUGUU ......(((((((((.(((((..............)))))..)))))))))......((((.((((.......((((.(((.......))).))))....)))).))))........... (-25.77 = -25.44 + -0.33)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:32:03 2006