
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,679,841 – 17,679,952 |
| Length | 111 |
| Max. P | 0.917954 |

| Location | 17,679,841 – 17,679,952 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 76.25 |
| Mean single sequence MFE | -34.45 |
| Consensus MFE | -19.50 |
| Energy contribution | -20.32 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.57 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.917954 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17679841 111 + 22224390 UGCCCCCUCCGAUUUAUCUAC-----UCCAAUCGCGCGAUUAGCGAGCCGAAAUUUCACAGUGUGUUUUCAGAGUCAGA--GAUACGGCAAAUCGGAGUGGGCACGGCCUUGCUGAUA (((((.(((((((((......-----.....((((.......))))((((..(((((..(.(.((....)).).)..))--))).))))))))))))).)))))((((...))))... ( -40.30) >DroSec_CAF1 400 117 + 1 UGCC-CCUCCGAUUUAUCUAUCUAUAUCCAAUCGCGCGAUUAGCGAGCCGAAAUUUCACAGUGUGUUUCCAGCGUCAGAGAGAUACGGCAAAUCGGAGUGGGCACGUCCAUGCUGAUA ((((-((((((((((................((((.......))))(((((....))...((((.((((........)))).)))))))))))))))).))))).............. ( -39.80) >DroEre_CAF1 590 108 + 1 UGCC-CCUCCGAUUUGUCUAC-----UCCAAUCGCGCGAUUAGGGAGAAGCAU--UUGCAAUGUAUUCCGAACGUCAGA--GAUAUGGUAAAUCGGAGUGGGCACGUCCAUGCUGAUA ((((-((((((((((..(..(-----(((.(((....)))...))))..((..--..))...((((..(........).--.)))))..))))))))).))))).............. ( -35.80) >DroYak_CAF1 398 86 + 1 UGCC-CCUCCGAUUUAUCUAC-----UCCAAUCG------------------------CAAUGUAUUUCAAACGUCAGA--GAUAUGGCAAAUCGGAAUGGGCACGACCAUGCUGAUA ((((-(.((((((((......-----.......(------------------------(...(((((((........))--))))).))))))))))..))))).............. ( -21.91) >consensus UGCC_CCUCCGAUUUAUCUAC_____UCCAAUCGCGCGAUUAGCGAGCCGAAA__UCACAAUGUAUUUCCAACGUCAGA__GAUACGGCAAAUCGGAGUGGGCACGUCCAUGCUGAUA ((((.((((((((((..(.............((((.......))))................(((((..............))))))..))))))))).))))).............. (-19.50 = -20.32 + 0.81)

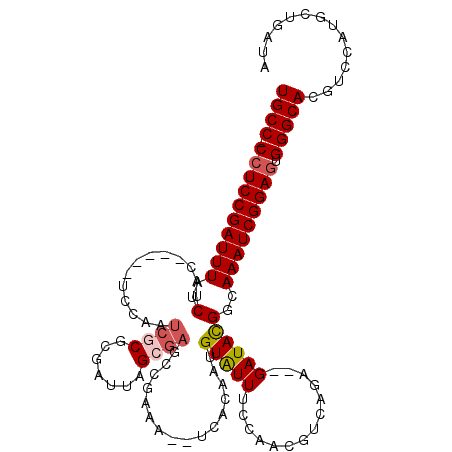
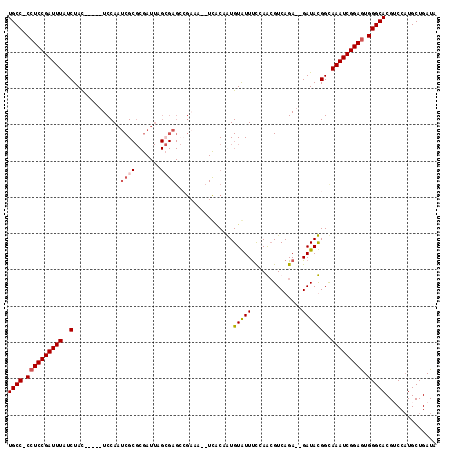
| Location | 17,679,841 – 17,679,952 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 76.25 |
| Mean single sequence MFE | -34.68 |
| Consensus MFE | -19.42 |
| Energy contribution | -19.67 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820872 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17679841 111 - 22224390 UAUCAGCAAGGCCGUGCCCACUCCGAUUUGCCGUAUC--UCUGACUCUGAAAACACACUGUGAAAUUUCGGCUCGCUAAUCGCGCGAUUGGA-----GUAGAUAAAUCGGAGGGGGCA ..............(((((.((((((((((.(....(--((..(..((((((.(((...)))...)))))).((((.......)))))..))-----)..).)))))))))).))))) ( -36.60) >DroSec_CAF1 400 117 - 1 UAUCAGCAUGGACGUGCCCACUCCGAUUUGCCGUAUCUCUCUGACGCUGGAAACACACUGUGAAAUUUCGGCUCGCUAAUCGCGCGAUUGGAUAUAGAUAGAUAAAUCGGAGG-GGCA ..............(((((.((((((((((.(.(((((.((..(.(((((((.(((...)))...)))))))((((.......)))))..))...)))))).)))))))))))-)))) ( -40.60) >DroEre_CAF1 590 108 - 1 UAUCAGCAUGGACGUGCCCACUCCGAUUUACCAUAUC--UCUGACGUUCGGAAUACAUUGCAA--AUGCUUCUCCCUAAUCGCGCGAUUGGA-----GUAGACAAAUCGGAGG-GGCA ..............(((((.(((((((((........--((((.....))))...........--...((.((((..((((....)))))))-----).))..))))))))))-)))) ( -33.10) >DroYak_CAF1 398 86 - 1 UAUCAGCAUGGUCGUGCCCAUUCCGAUUUGCCAUAUC--UCUGACGUUUGAAAUACAUUG------------------------CGAUUGGA-----GUAGAUAAAUCGGAGG-GGCA ..............(((((.((((((((((...((.(--((..((((.((.....))..)------------------------)).)..))-----)))..)))))))))))-)))) ( -28.40) >consensus UAUCAGCAUGGACGUGCCCACUCCGAUUUGCCAUAUC__UCUGACGCUGGAAACACACUGUGA__UUUCGGCUCGCUAAUCGCGCGAUUGGA_____GUAGAUAAAUCGGAGG_GGCA ..............(((((.((((((((((.(.(((...((..((((....................................))).)..)).....)))).))))))))))).)))) (-19.42 = -19.67 + 0.25)

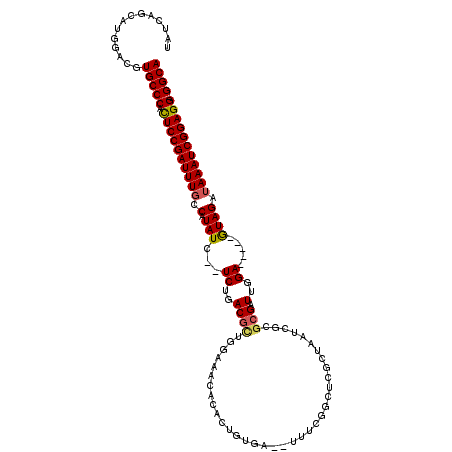

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:29:09 2006