
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,631,299 – 17,631,403 |
| Length | 104 |
| Max. P | 0.968463 |

| Location | 17,631,299 – 17,631,403 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 75.65 |
| Mean single sequence MFE | -23.92 |
| Consensus MFE | -14.99 |
| Energy contribution | -17.30 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873130 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17631299 104 + 22224390 AAAACAAGACUGGAAGCCCGAAGAGCCGAAAAGCGAACAACGCAACCCAUCAACAUCAGCGACUAAUGGGACUCUUGUUCGCAUCGGAGAAAAGGGGGCAAUUG ...............((((......((((...((((((((.....(((((...............)))))....)))))))).))))........))))..... ( -27.60) >DroPse_CAF1 45814 83 + 1 AAAAUCAAACCG-AAGCAAACCGAACCGUACAACAGCCAGGG---CCCAUCAACAUCAACGACUAAUGGG-CUCUUCUCC---UCUGAGGA------------- ............-.........................((((---(((((...............)))))-))))((((.---...)))).------------- ( -13.76) >DroEre_CAF1 27207 102 + 1 AAAACAAGACUGGAAGCCUGAAGAGCCGAAAAGCGAACAACGCAACCCAUCAACAUCAGCGACUAAUGGAACUCUUGUUCGCAUCGGAAA--AAAGGGCAAUUG ...............((((......((((...((((((((......((((...............)))).....)))))))).))))...--...))))..... ( -24.46) >DroYak_CAF1 25618 102 + 1 AAAACAAGACUGGAAGCCCAAAGAGCCGAAAAGCGAACAACGCAACCCAUCAACAUCAGCGACUAAUGGGACUCUUGUUCGCAUCGGAAA--ACGGGGCAAUUG ...............((((......((((...((((((((.....(((((...............)))))....)))))))).))))...--...))))..... ( -29.86) >consensus AAAACAAGACUGGAAGCCCAAAGAGCCGAAAAGCGAACAACGCAACCCAUCAACAUCAGCGACUAAUGGGACUCUUGUUCGCAUCGGAAA__A_GGGGCAAUUG ...............((((......((((...((((((((.....(((((...............)))))....)))))))).))))........))))..... (-14.99 = -17.30 + 2.31)


| Location | 17,631,299 – 17,631,403 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 75.65 |
| Mean single sequence MFE | -30.17 |
| Consensus MFE | -19.61 |
| Energy contribution | -21.86 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.63 |
| SVM RNA-class probability | 0.968463 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17631299 104 - 22224390 CAAUUGCCCCCUUUUCUCCGAUGCGAACAAGAGUCCCAUUAGUCGCUGAUGUUGAUGGGUUGCGUUGUUCGCUUUUCGGCUCUUCGGGCUUCCAGUCUUGUUUU .....((((........((((.(((((((((...((((((((.((....))))))))))...).))))))))...))))......))))............... ( -33.14) >DroPse_CAF1 45814 83 - 1 -------------UCCUCAGA---GGAGAAGAG-CCCAUUAGUCGUUGAUGUUGAUGGG---CCCUGGCUGUUGUACGGUUCGGUUUGCUU-CGGUUUGAUUUU -------------...(((((---...((((.(-((((((((.((....))))))))))---)((..((((.....))))..))....)))-)..))))).... ( -24.90) >DroEre_CAF1 27207 102 - 1 CAAUUGCCCUUU--UUUCCGAUGCGAACAAGAGUUCCAUUAGUCGCUGAUGUUGAUGGGUUGCGUUGUUCGCUUUUCGGCUCUUCAGGCUUCCAGUCUUGUUUU .....(((....--...((((.(((((((((...((((((((.((....))))))))))...).))))))))...)))).......)))............... ( -28.34) >DroYak_CAF1 25618 102 - 1 CAAUUGCCCCGU--UUUCCGAUGCGAACAAGAGUCCCAUUAGUCGCUGAUGUUGAUGGGUUGCGUUGUUCGCUUUUCGGCUCUUUGGGCUUCCAGUCUUGUUUU .....((((...--...((((.(((((((((...((((((((.((....))))))))))...).))))))))...))))......))))............... ( -34.30) >consensus CAAUUGCCCC_U__UCUCCGAUGCGAACAAGAGUCCCAUUAGUCGCUGAUGUUGAUGGGUUGCGUUGUUCGCUUUUCGGCUCUUCGGGCUUCCAGUCUUGUUUU .....((((........((((.(((((((((...((((((((.((....))))))))))...).))))))))...))))......))))............... (-19.61 = -21.86 + 2.25)
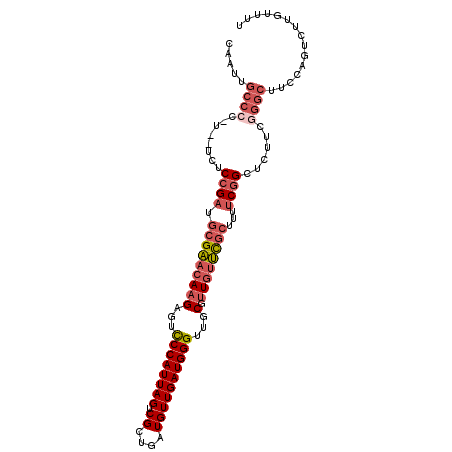



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:28:55 2006