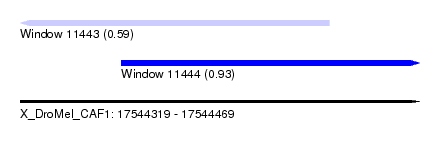

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,544,319 – 17,544,469 |
| Length | 150 |
| Max. P | 0.933291 |

| Location | 17,544,319 – 17,544,435 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.48 |
| Mean single sequence MFE | -24.27 |
| Consensus MFE | -20.56 |
| Energy contribution | -21.62 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.592959 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17544319 116 - 22224390 UUGUUAACACCAGAAACCAGUAAAGAAACUUUAUUUUGCACCUUACAUACGCG--UGUGUGUGUGCGUGCGCGUAUAUUUAAUUCUGGUGUUUUGUAUUAUUUUUAAAUCACGAACUC-- .....((((((((((....((((((........)))))).......(((((((--..(((....)))..)))))))......))))))))))..........................-- ( -31.80) >DroSec_CAF1 36435 105 - 1 UUGUUAUCACCCGAAACCAGUAAAGAAACUUUAUUUUGCACCUUACAU------------GUGUGCGUGCGCGUAUAUUUAAUUCUGGUGUUUUGUAUUAUUUUUAAAUCACGA-CUC-- .......................(((((...(((...(((((......------------(((((((....)))))))........)))))...)))...))))).........-...-- ( -15.44) >DroSim_CAF1 25952 107 - 1 UUGUUAUCACCCGAAACCAGUAAAGAAACUUUAUUUUGCACCUUACAU------------GUGUGCGUGCGCGUAUAUUUAAUUCUGGGGUUUUGUAUUAUUUUUAAAUCUUAA-AUCGU ...........(((((((.((((((........)))))).((......------------(((((((....)))))))........)))))))))...................-..... ( -18.24) >DroYak_CAF1 33245 117 - 1 UUGUUAUCACCAGAAACCAGUAAAGAAACUUUAUUUUGCACCUUACAUACGCGCGCGUGUGUGUGUGUGUGCGUAUAUUUUAUUCUGGUGUUUUGUAUUAUUUUUAAAUCACGA-CUC-- .......((((((((...(((((((...)))))))..((((..((((((((((....)))))))))).))))..........))))))))........................-...-- ( -31.60) >consensus UUGUUAUCACCAGAAACCAGUAAAGAAACUUUAUUUUGCACCUUACAU____________GUGUGCGUGCGCGUAUAUUUAAUUCUGGUGUUUUGUAUUAUUUUUAAAUCACGA_CUC__ .......((((((((...(((((((...)))))))..((((..(((((((((....))))))))).))))............)))))))).............................. (-20.56 = -21.62 + 1.06)


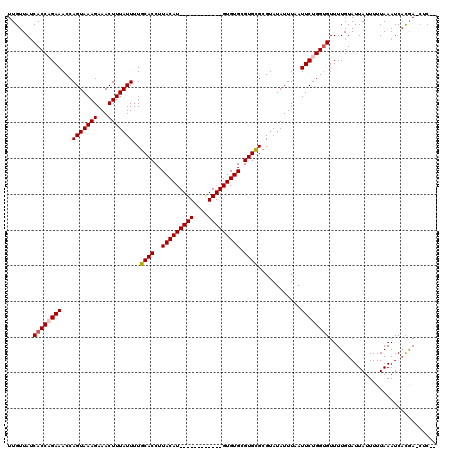
| Location | 17,544,357 – 17,544,469 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 87.32 |
| Mean single sequence MFE | -21.01 |
| Consensus MFE | -16.84 |
| Energy contribution | -17.40 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.933291 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
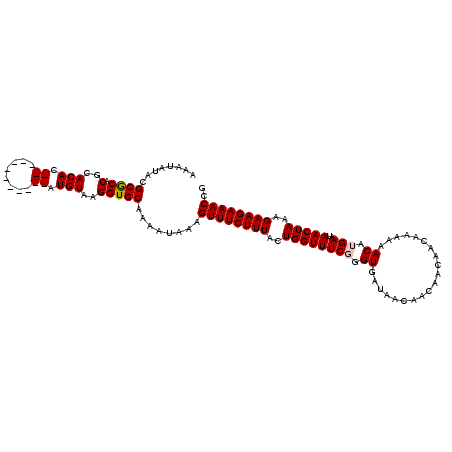
>X_DroMel_CAF1 17544357 112 + 22224390 AAAUAUACGCGCACGCACACACACA--CGCGUAUGUAAGGUGCAAAAUAAAGUUUCUUUACUGGUUUCUGGUGUUAACAAC---AACAAAAAACAUGAAUAACUAAAGAAGAAACCG ..(((((((((..............--)))))))))...............((((((((..(((((..(.(((((......---.......))))).)..)))))..)))))))).. ( -24.16) >DroSec_CAF1 36472 105 + 1 AAAUAUACGCGCACGCACAC------------AUGUAAGGUGCAAAAUAAAGUUUCUUUACUGGUUUCGGGUGAUAACAACAACAACAAAAAACGUGAAUAACUAAAGAAGAAACCG ........((((..(((...------------.)))...))))........((((((((..((((((((.((....................)).)))..)))))..)))))))).. ( -19.15) >DroSim_CAF1 25991 105 + 1 AAAUAUACGCGCACGCACAC------------AUGUAAGGUGCAAAAUAAAGUUUCUUUACUGGUUUCGGGUGAUAACAACAACAACAAAAAACGUGAAUAACUAAAGAAGAAACCG ........((((..(((...------------.)))...))))........((((((((..((((((((.((....................)).)))..)))))..)))))))).. ( -19.15) >DroYak_CAF1 33282 115 + 1 AAAUAUACGCACACACACACACACGCGCGCGUAUGUAAGGUGCAAAAUAAAGUUUCUUUACUGGUUUCUGGUGAUAACAACAAA--AAAAAAACAUGAAUAACUAAAGAAGAAACCA ........((((......(((.(((....))).)))...))))........((((((((..(((((..(.(((...........--.......))).)..)))))..)))))))).. ( -21.57) >consensus AAAUAUACGCGCACGCACAC____________AUGUAAGGUGCAAAAUAAAGUUUCUUUACUGGUUUCGGGUGAUAACAACAACAACAAAAAACAUGAAUAACUAAAGAAGAAACCG ........((((.(..(((.((........)).)))..)))))........((((((((..((((((((.((....................)).)))..)))))..)))))))).. (-16.84 = -17.40 + 0.56)

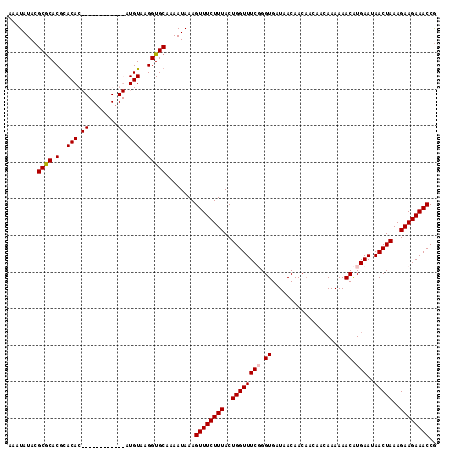
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:27:52 2006