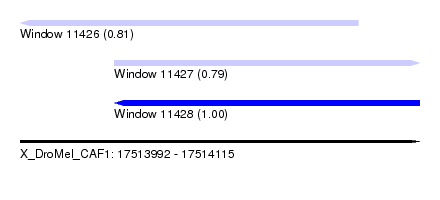
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,513,992 – 17,514,115 |
| Length | 123 |
| Max. P | 0.998194 |

| Location | 17,513,992 – 17,514,096 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 78.98 |
| Mean single sequence MFE | -22.44 |
| Consensus MFE | -13.92 |
| Energy contribution | -14.12 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.813869 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17513992 104 - 22224390 CCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAAAGUCAUCCAUUGAUUCCUAAAAACUC---AGAAU----AUUGGCAUUUCAAGAACCUAUAAAC----UA--AA-- ....(.((((((.(((.(((((....))))))))...(((....(((((.....))))))))........---.....----..))).))).)..............----..--..-- ( -18.50) >DroSec_CAF1 4860 97 - 1 CCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAUAGUCAUCCAUGGAUUCCUAAAAAC-C---A---U----ACUAUGAACUCAGAA---UAUAGAC----UA--AA-- ...((.((((((.(((.(((((....))))))))....(((....))).....)))))).))......-.---.---.----.(((((.........---)))))..----..--..-- ( -21.80) >DroSim_CAF1 3211 97 - 1 CCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAUAGUCAUCCAUGGAUUCCUAAAAAC-C---A---U----ACUAUGAACUCAGAA---UAUAGAC----UA--AA-- ...((.((((((.(((.(((((....))))))))....(((....))).....)))))).))......-.---.---.----.(((((.........---)))))..----..--..-- ( -21.80) >DroEre_CAF1 3133 109 - 1 CCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAAAGUCGUCCAUUGUUUCCUGAAAAC-C---A---UAUGCAUUAUGAACUGGCGA---UAUUGGUAUUUUAAGAACC .............(((.(((((....))))))))((((.(((((.((((.(((..((((.....))))-(---(---((.....))))...))))))---).))))).))))....... ( -25.80) >DroYak_CAF1 3123 111 - 1 CCAAGCGAUCCACUCCACCGUGUACACA-GGGGAAAAAGGCUAAAGUCAUCCAUUGUUUCCUUACAAC-CUUAA---UGUAUAUGAUGAACUGGAAA---UAUUGGCAUUUUAAUAACU ((((....((((...((.((((((((..-((((.....(((....))).....((((......)))))-)))..---)))))))).))...))))..---..))))............. ( -24.30) >consensus CCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAAAGUCAUCCAUUGAUUCCUAAAAAC_C___A___U____AUUAUGAACUCAAAA___UAUAGAC____UA__AA__ ...((.((..(..(((.(((((....))))))))....(((....))).......)..))))......................................................... (-13.92 = -14.12 + 0.20)


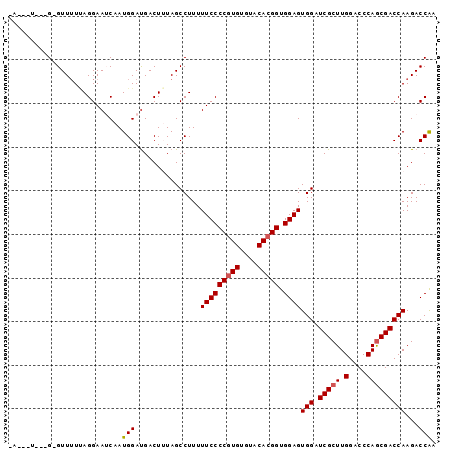
| Location | 17,514,021 – 17,514,115 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 91.44 |
| Mean single sequence MFE | -29.48 |
| Consensus MFE | -23.20 |
| Energy contribution | -23.44 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.790189 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17514021 94 + 22224390 -AUUCU---GAGUUUUUAGGAAUCAAUGGAUGACUUUAGCCUUUUUCCCCGUGUGUACACGGUGGAGUGGAUCGCUUGGACCCAGCGACCAAGACCAA -.((((---((....)))))).....(((...............(((((((((....))))).))))(((.(((((.(....)))))))))...))). ( -27.60) >DroSec_CAF1 4886 90 + 1 -A---U---G-GUUUUUAGGAAUCCAUGGAUGACUAUAGCCUUUUUCCCCGUGUGUACACGGUGGAGUGGAUCGCUUGGACCCAGCGACCAAGACCAA -.---(---(-((((((((..(((....)))..)))........(((((((((....))))).)))).((.(((((.(....))))))))))))))). ( -33.00) >DroSim_CAF1 3237 90 + 1 -A---U---G-GUUUUUAGGAAUCCAUGGAUGACUAUAGCCUUUUUCCCCGUGUGUACACGGUGGAGUGGAUCGCUUGGACCCAGCGACCAAGACCAA -.---(---(-((((((((..(((....)))..)))........(((((((((....))))).)))).((.(((((.(....))))))))))))))). ( -33.00) >DroEre_CAF1 3170 91 + 1 UA---U---G-GUUUUCAGGAAACAAUGGACGACUUUAGCCUUUUUCCCCGUGUGUACACGGUGGAGUGGAUCGCUUGGACCCAGCGACCAAGACCAA ..---(---(-((((...(....)..(((..........((...(((((((((....))))).)))).)).(((((.(....))))))))))))))). ( -31.00) >DroYak_CAF1 3160 93 + 1 CA---UUAAG-GUUGUAAGGAAACAAUGGAUGACUUUAGCCUUUUUCCCC-UGUGUACACGGUGGAGUGGAUCGCUUGGACCCAACGACCAAGACCGA ..---....(-(((....(....)..(((..........((...((((.(-(((....)))).)))).)).(((.(((....))))))))).)))).. ( -22.80) >consensus _A___U___G_GUUUUUAGGAAUCAAUGGAUGACUUUAGCCUUUUUCCCCGUGUGUACACGGUGGAGUGGAUCGCUUGGACCCAGCGACCAAGACCAA ..........................(((...............(((((((((....))))).))))(((.(((((.(....)))))))))...))). (-23.20 = -23.44 + 0.24)


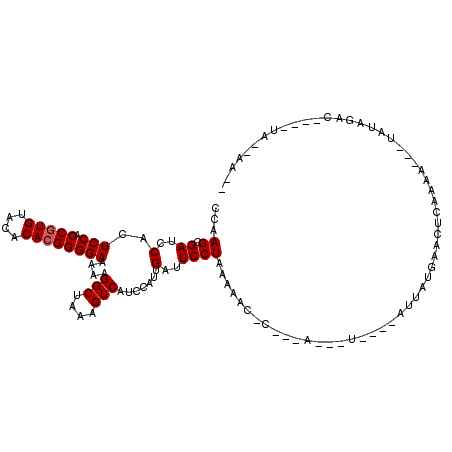
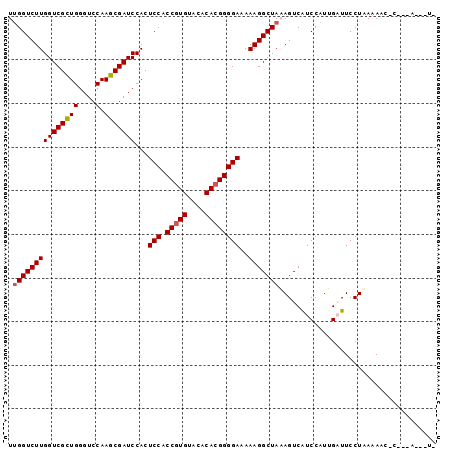
| Location | 17,514,021 – 17,514,115 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 91.44 |
| Mean single sequence MFE | -28.80 |
| Consensus MFE | -25.88 |
| Energy contribution | -26.12 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.90 |
| SVM decision value | 3.03 |
| SVM RNA-class probability | 0.998194 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17514021 94 - 22224390 UUGGUCUUGGUCGCUGGGUCCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAAAGUCAUCCAUUGAUUCCUAAAAACUC---AGAAU- ((((((((((((((((....).)))))))...(((.(((((....))))))))..))))))))(((((.....)))))...........---.....- ( -30.60) >DroSec_CAF1 4886 90 - 1 UUGGUCUUGGUCGCUGGGUCCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAUAGUCAUCCAUGGAUUCCUAAAAAC-C---A---U- .(((((((((((((((....).)))))))...(((.(((((....))))))))..)))))))(((..(((....)))..))).....-.---.---.- ( -31.70) >DroSim_CAF1 3237 90 - 1 UUGGUCUUGGUCGCUGGGUCCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAUAGUCAUCCAUGGAUUCCUAAAAAC-C---A---U- .(((((((((((((((....).)))))))...(((.(((((....))))))))..)))))))(((..(((....)))..))).....-.---.---.- ( -31.70) >DroEre_CAF1 3170 91 - 1 UUGGUCUUGGUCGCUGGGUCCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAAAGUCGUCCAUUGUUUCCUGAAAAC-C---A---UA ((((((((((((((((....).)))))))...(((.(((((....))))))))..))))))))........................-.---.---.. ( -28.70) >DroYak_CAF1 3160 93 - 1 UCGGUCUUGGUCGUUGGGUCCAAGCGAUCCACUCCACCGUGUACACA-GGGGAAAAAGGCUAAAGUCAUCCAUUGUUUCCUUACAAC-CUUAA---UG ..((..((((((..((((((.....)))))).(((.((.((....))-)))))....))))))..)).....((((......)))).-.....---.. ( -21.30) >consensus UUGGUCUUGGUCGCUGGGUCCAAGCGAUCCACUCCACCGUGUACACACGGGGAAAAAGGCUAAAGUCAUCCAUUGAUUCCUAAAAAC_C___A___U_ .(((((((((((((((....).)))))))...(((.(((((....))))))))..))))))).................................... (-25.88 = -26.12 + 0.24)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:27:37 2006