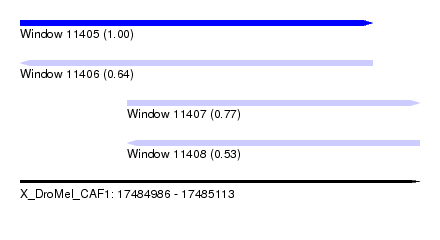

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,484,986 – 17,485,113 |
| Length | 127 |
| Max. P | 0.999991 |

| Location | 17,484,986 – 17,485,098 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 91.03 |
| Mean single sequence MFE | -38.16 |
| Consensus MFE | -36.74 |
| Energy contribution | -36.98 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.08 |
| Mean z-score | -4.13 |
| Structure conservation index | 0.96 |
| SVM decision value | 5.62 |
| SVM RNA-class probability | 0.999991 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17484986 112 + 22224390 ------AGAAACGGAGAAGAAAAACGCGCAAGCGAUUUUCUUAUGCCGGCGCUGCCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAGCGAAUAAUAUACGCGAAUCGCGAUUUCACAA ------.(((((((..(((((((.(((....))).)))))))...))).(((((.((((((((.(((.....))))))))))).))))).........((((...)))).)))).... ( -42.60) >DroSec_CAF1 2918 109 + 1 ------AGAAACGGAGAAGAAAAACGCGGAAGCGAUUUUCUUAUGCCGGCGCUGCCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAGCGAA---UAUCCGCGAAACGCGAUUUCACAA ------.(((((((..(((((((.(((....))).)))))))...))).(((((.((((((((.(((.....))))))))))).)))))..---....((((...)))).)))).... ( -42.40) >DroSim_CAF1 2679 109 + 1 ------AGAAAUGGAGAAGAAAAACGCGCAAGCGAUUUUCUUAUGCCGGCGCUGCCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAGCGAA---UAUCCGCGAAACGCGAUUUCACAA ------.(((((((..(((((((.(((....))).)))))))...))..(((((.((((((((.(((.....))))))))))).)))))..---....((((...))))))))).... ( -41.30) >DroEre_CAF1 2958 109 + 1 ------AGAAACGGAGAAGAAAAACGCGAAAGCGAUUUUUUUAUGCCGGCGCUACCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAACGAA---CUUCCGCGAAUAGUGAUUUCACAA ------.(((((((..(((((((.(((....))).)))))))...))).(((((.((((((((.(((.....)))))))))))....((..---......))..))))).)))).... ( -30.10) >DroYak_CAF1 2965 115 + 1 ACACACAGAAACGGAGAAGAAAAACGCGAAAGCGAUUUUCUUAUGCCGGCGCUACCCGAAAAAUCAUUCCAUGUGAUUUUUGGCCAACGAA---UUUCCGCGAAUUGUGAUUUCACAA ...(((((...((((((((((((.(((....))).)))))))..(((((......)).(((((((((.....)))))))))))).......---.)))))....)))))......... ( -34.40) >consensus ______AGAAACGGAGAAGAAAAACGCGAAAGCGAUUUUCUUAUGCCGGCGCUGCCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAGCGAA___UAUCCGCGAAUCGCGAUUUCACAA .......(((((((..(((((((.(((....))).)))))))...))).(((((.(((((((.((((.....))))))))))).))))).........(((.....))).)))).... (-36.74 = -36.98 + 0.24)

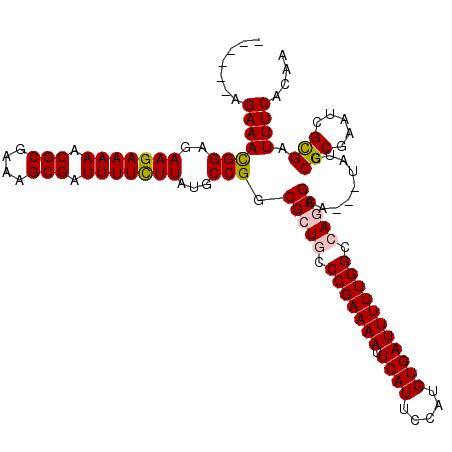
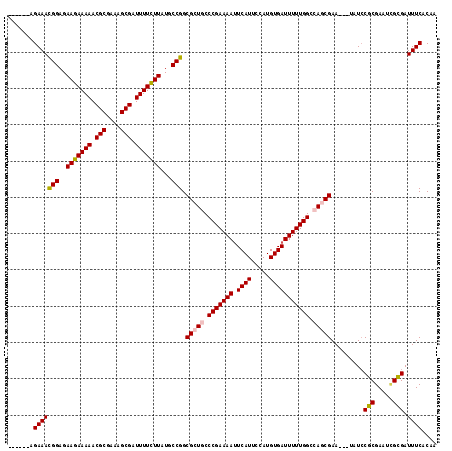
| Location | 17,484,986 – 17,485,098 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 91.03 |
| Mean single sequence MFE | -30.48 |
| Consensus MFE | -26.24 |
| Energy contribution | -26.16 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.637354 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17484986 112 - 22224390 UUGUGAAAUCGCGAUUCGCGUAUAUUAUUCGCUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGCAGCGCCGGCAUAAGAAAAUCGCUUGCGCGUUUUUCUUCUCCGUUUCU------ ..((((.((((((...)))).)).)))).(((((.((((((.(((.......))).)))).)).))))).(((...(((((((.(((....))).)))))))..))).....------ ( -30.20) >DroSec_CAF1 2918 109 - 1 UUGUGAAAUCGCGUUUCGCGGAUA---UUCGCUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGCAGCGCCGGCAUAAGAAAAUCGCUUCCGCGUUUUUCUUCUCCGUUUCU------ (((((((((...)))))))))...---..(((((.((((((.(((.......))).)))).)).))))).(((...(((((((.(((....))).)))))))..))).....------ ( -31.70) >DroSim_CAF1 2679 109 - 1 UUGUGAAAUCGCGUUUCGCGGAUA---UUCGCUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGCAGCGCCGGCAUAAGAAAAUCGCUUGCGCGUUUUUCUUCUCCAUUUCU------ ..(((((((...)))))))((...---..(((((.((((((.(((.......))).)))).)).)))))))((...(((((((.(((....))).)))))))..))......------ ( -30.70) >DroEre_CAF1 2958 109 - 1 UUGUGAAAUCACUAUUCGCGGAAG---UUCGUUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGUAGCGCCGGCAUAAAAAAAUCGCUUUCGCGUUUUUCUUCUCCGUUUCU------ ..(((....)))....(.((((((---(((((...(((.........))).))))))))))).)......(((.....(((((.(((....)))))))).....))).....------ ( -22.60) >DroYak_CAF1 2965 115 - 1 UUGUGAAAUCACAAUUCGCGGAAA---UUCGUUGGCCAAAAAUCACAUGGAAUGAUUUUUCGGGUAGCGCCGGCAUAAGAAAAUCGCUUUCGCGUUUUUCUUCUCCGUUUCUGUGUGU (((((....)))))..((((((((---..(((((.((((((((((.......)))))))).)).))))).(((...(((((((.(((....))).)))))))..)))))))))))... ( -37.20) >consensus UUGUGAAAUCGCGAUUCGCGGAUA___UUCGCUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGCAGCGCCGGCAUAAGAAAAUCGCUUGCGCGUUUUUCUUCUCCGUUUCU______ .((((((.......)))))).........(((((.((((((.(((.......))).)))).)).))))).(((...(((((((.(((....))).)))))))..)))........... (-26.24 = -26.16 + -0.08)

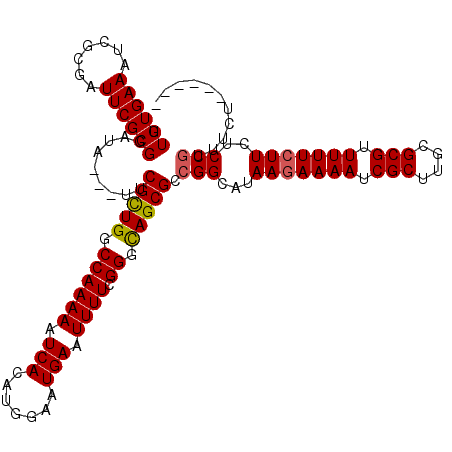
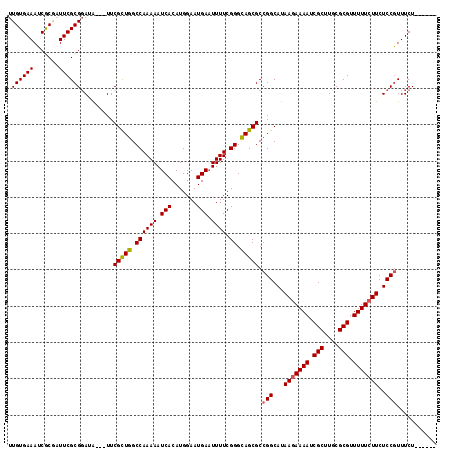
| Location | 17,485,020 – 17,485,113 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 91.38 |
| Mean single sequence MFE | -23.70 |
| Consensus MFE | -20.60 |
| Energy contribution | -21.00 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.767273 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17485020 93 + 22224390 UUAUGCCGGCGCUGCCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAGCGAAUAAUAUACGCGAAUCGCGAUUUCACAAAAAAUGCACAGGGC-U ....(((..(((((.((((((((.(((.....))))))))))).))))).........((((...))))....................)))-. ( -27.10) >DroSec_CAF1 2952 90 + 1 UUAUGCCGGCGCUGCCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAGCGAA---UAUCCGCGAAACGCGAUUUCACAACAAACGCACAGGGC-U ....(((.((((((.((((((((.(((.....))))))))))).))))(((---....((((...))))..))).........))....)))-. ( -27.60) >DroSim_CAF1 2713 90 + 1 UUAUGCCGGCGCUGCCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAGCGAA---UAUCCGCGAAACGCGAUUUCACAACAAACGCACAGGGC-U ....(((.((((((.((((((((.(((.....))))))))))).))))(((---....((((...))))..))).........))....)))-. ( -27.60) >DroEre_CAF1 2992 91 + 1 UUAUGCCGGCGCUACCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAACGAA---CUUCCGCGAAUAGUGAUUUCACAACAAAUGCACAGGGUUU .....((.((((...((((((((.(((.....))))))))))).....(..---....))).....(((....))).......))...)).... ( -15.70) >DroYak_CAF1 3005 90 + 1 UUAUGCCGGCGCUACCCGAAAAAUCAUUCCAUGUGAUUUUUGGCCAACGAA---UUUCCGCGAAUUGUGAUUUCACAACAAAUGCACAGGGC-U ....(((((((.....).(((((((((.....))))))))).)))......---.....(((..(((((....)))))....)))....)))-. ( -20.50) >consensus UUAUGCCGGCGCUGCCCGAAAAUUCAUUCCAUGUGAUUUUUGGCCAGCGAA___UAUCCGCGAAUCGCGAUUUCACAACAAAUGCACAGGGC_U ....(((..(((((.(((((((.((((.....))))))))))).))))).........(((.....)))....................))).. (-20.60 = -21.00 + 0.40)

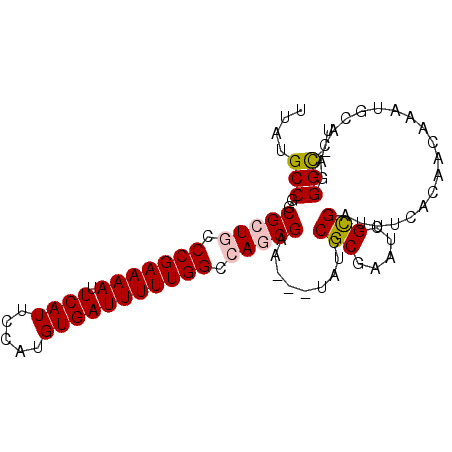
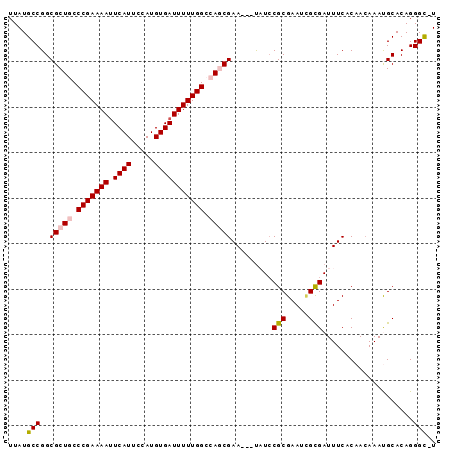
| Location | 17,485,020 – 17,485,113 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 91.38 |
| Mean single sequence MFE | -26.12 |
| Consensus MFE | -22.26 |
| Energy contribution | -22.62 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.85 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.527792 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17485020 93 - 22224390 A-GCCCUGUGCAUUUUUUGUGAAAUCGCGAUUCGCGUAUAUUAUUCGCUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGCAGCGCCGGCAUAA .-(((.(((((.....(((((....))))).....))))).....(((((.((((((.(((.......))).)))).)).)))))..))).... ( -25.70) >DroSec_CAF1 2952 90 - 1 A-GCCCUGUGCGUUUGUUGUGAAAUCGCGUUUCGCGGAUA---UUCGCUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGCAGCGCCGGCAUAA .-(((....(((.....((((....))))...)))((...---..(((((.((((((.(((.......))).)))).)).)))))))))).... ( -26.50) >DroSim_CAF1 2713 90 - 1 A-GCCCUGUGCGUUUGUUGUGAAAUCGCGUUUCGCGGAUA---UUCGCUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGCAGCGCCGGCAUAA .-(((....(((.....((((....))))...)))((...---..(((((.((((((.(((.......))).)))).)).)))))))))).... ( -26.50) >DroEre_CAF1 2992 91 - 1 AAACCCUGUGCAUUUGUUGUGAAAUCACUAUUCGCGGAAG---UUCGUUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGUAGCGCCGGCAUAA ....((.((((.......(((....)))....(.((((((---(((((...(((.........))).))))))))))).)..)))).))..... ( -23.80) >DroYak_CAF1 3005 90 - 1 A-GCCCUGUGCAUUUGUUGUGAAAUCACAAUUCGCGGAAA---UUCGUUGGCCAAAAAUCACAUGGAAUGAUUUUUCGGGUAGCGCCGGCAUAA .-((((((((.....((((((....)))))).)))))...---..(((((.((((((((((.......)))))))).)).)))))..))).... ( -28.10) >consensus A_GCCCUGUGCAUUUGUUGUGAAAUCGCGAUUCGCGGAUA___UUCGCUGGCCAAAAAUCACAUGGAAUGAAUUUUCGGGCAGCGCCGGCAUAA ..((((((((.....((((((....)))))).)))))........(((((.((((((.(((.......))).)))).)).)))))..))).... (-22.26 = -22.62 + 0.36)


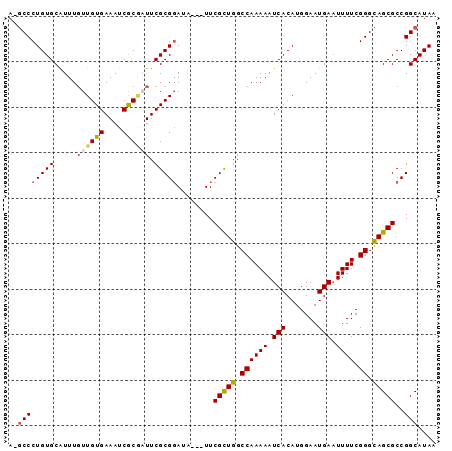
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:27:20 2006