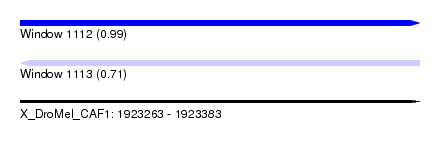

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,923,263 – 1,923,383 |
| Length | 120 |
| Max. P | 0.994263 |

| Location | 1,923,263 – 1,923,383 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.56 |
| Mean single sequence MFE | -39.27 |
| Consensus MFE | -28.18 |
| Energy contribution | -29.77 |
| Covariance contribution | 1.59 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.72 |
| SVM decision value | 2.46 |
| SVM RNA-class probability | 0.994263 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
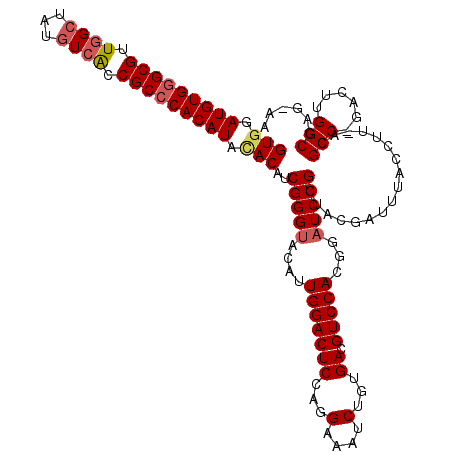
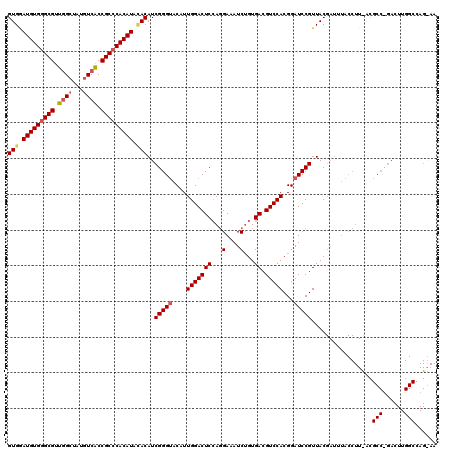
>X_DroMel_CAF1 1923263 120 + 22224390 GUGAAUGUGGGCGUUUGCUAUAUCACCGCCCACAUACACAUCGGGUACAUUGGACUCCAGGAAAUCAGUGACGUCCACGGAUCCGUUACGAUUUACCUUCACGCCCGACUUGGCCAGAAA (((.(((((((((.............))))))))).))).((((((....(((...)))(((((((.((((((((....))..))))))))))).)).....))))))............ ( -42.02) >DroEre_CAF1 7301 120 + 1 GUGGAUGUGAGCGUUGGCUAUGUCGCCGCCCACAUAUACAUCGGGUACAUUGGACUCGAGGAAAUCUGUGACGUCCACGGAUCCGUUACGAUUUACCUUUACGCCUGACUUGGCCAGGAA (((.(((((.(((.((((...)))).))).))))).))).((((((.......))))))(((((((.((((((((....))..))))))))))).)).....(((......)))...... ( -37.80) >DroYak_CAF1 7380 120 + 1 GUGGAUGUGGGCGUUGGCUAUGUCACCGCCCACAUAGACAUCGGGCACGUUGGACUCCAUGAAAUCUGUGACGUCCACGGAUCCGUUACNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN ((..(((((((((.((((...)))).)))))))))..))..((((..(((.((((..(((.......)))..)))))))..))))................................... ( -38.00) >consensus GUGGAUGUGGGCGUUGGCUAUGUCACCGCCCACAUACACAUCGGGUACAUUGGACUCCAGGAAAUCUGUGACGUCCACGGAUCCGUUACGAUUUACCUU_ACGCC_GACUUGGCCAG_AA (((.(((((((((.((((...)))).))))))))).)))..(((((....(((((((...(....)...)).)))))...))))).................(((......)))...... (-28.18 = -29.77 + 1.59)

| Location | 1,923,263 – 1,923,383 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.56 |
| Mean single sequence MFE | -36.17 |
| Consensus MFE | -26.50 |
| Energy contribution | -25.90 |
| Covariance contribution | -0.60 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.714591 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
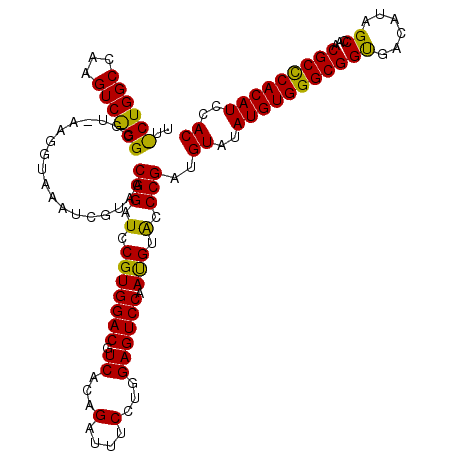
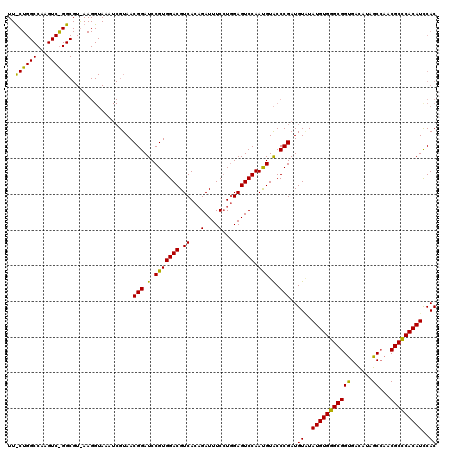
>X_DroMel_CAF1 1923263 120 - 22224390 UUUCUGGCCAAGUCGGGCGUGAAGGUAAAUCGUAACGGAUCCGUGGACGUCACUGAUUUCCUGGAGUCCAAUGUACCCGAUGUGUAUGUGGGCGGUGAUAUAGCAAACGCCCACAUUCAC ...........((((((.((...((....))...))(((((((.(((.(((...))).))))))).)))......))))))(((.(((((((((.............))))))))).))) ( -40.72) >DroEre_CAF1 7301 120 - 1 UUCCUGGCCAAGUCAGGCGUAAAGGUAAAUCGUAACGGAUCCGUGGACGUCACAGAUUUCCUCGAGUCCAAUGUACCCGAUGUAUAUGUGGGCGGCGACAUAGCCAACGCUCACAUCCAC ...(((((...)))))((((...((((...(((...(((..((.(((.(((...))).))).))..))).)))))))..))))..(((((((((((......)))...)))))))).... ( -36.30) >DroYak_CAF1 7380 120 - 1 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNGUAACGGAUCCGUGGACGUCACAGAUUUCAUGGAGUCCAACGUGCCCGAUGUCUAUGUGGGCGGUGACAUAGCCAACGCCCACAUCCAC ....................................((((..(((((((((...(((((....)))))....(....))))))))))(((((((.((.......)).))))))))))).. ( -31.50) >consensus UU_CUGGCCAAGUC_GGCGU_AAGGUAAAUCGUAACGGAUCCGUGGACGUCACAGAUUUCCUGGAGUCCAAUGUACCCGAUGUAUAUGUGGGCGGUGACAUAGCCAACGCCCACAUCCAC ..((((((...))))))..................(((.(.(((((((.((...(....)...)))))).))).).)))..((..(((((((((((......))...)))))))))..)) (-26.50 = -25.90 + -0.60)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:51:19 2006