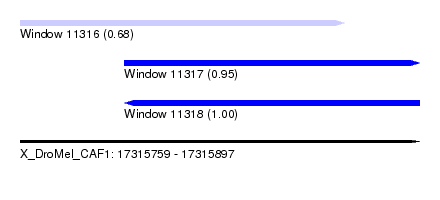
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,315,759 – 17,315,897 |
| Length | 138 |
| Max. P | 0.997820 |

| Location | 17,315,759 – 17,315,871 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.57 |
| Mean single sequence MFE | -34.24 |
| Consensus MFE | -21.34 |
| Energy contribution | -22.22 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.682990 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17315759 112 + 22224390 UUUGCAA-GCAC---ACUCAGAUACAUAUAUCACUUUCCCAAAAGUUCCUAUUAGCUAGGUUCCACUGC----AGCUACCCAGUUGUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAA ((..(..-.(((---.....((((....))))(((((....)))))......(((((((((.((((.((----((((....)))))).)))))))))))))......)))....)..)). ( -30.60) >DroSec_CAF1 32930 108 + 1 UUUGCAA-GCAC---ACUCAGAU----AUAUCACUUUUCCAAGAGUUCCAGUUAGCUAGGUUCCACUGC----AGCUACCCAGUUGUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAA .......-....---........----...((((((((((..(.....).((.((((((((.((((.((----((((....)))))).)))))))))))))).....).))))))))).. ( -32.30) >DroSim_CAF1 22381 108 + 1 UUUGCAA-GCAC---ACUCAGAU----AUAUCACUUUUCCAAGAGUUCCUGUUAGCUAGGUUCCACUGC----AGCUACCCAGUUGUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAA .......-....---........----...(((((((((.....((...(((.((((((((.((((.((----((((....)))))).)))))))))))))))))....))))))))).. ( -33.00) >DroEre_CAF1 33957 112 + 1 UAUGCAUCAUAUGUUACUCAGAU----GUAACACUUUUCUACGAGUUCUUGCUAGCCAGGCUCCACUGC----AGCUACGUUGGUUUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAA ..((((((............)))----))).((((((((.((..((...(((.((((((((.((((.(.----((((.....)))).))))))))))))))))))..))))))))))... ( -41.50) >DroYak_CAF1 23697 112 + 1 UUUGCAG-ACAA---GCUCAGAU----AUAUCACUUCUCUGGGAGUUCUCAUUAGCUAGGUUCCACUGCCGCCAGCUACCCAGUUUUCGUGGGCUUGGCUGCAACAAGUGAAAAGAGGAA .((((((-.(((---(((((((.----...))......(((((.(....)..(((((.(((......)))...))))))))))......)))))))).))))))................ ( -33.80) >consensus UUUGCAA_GCAC___ACUCAGAU____AUAUCACUUUUCCAAGAGUUCCUGUUAGCUAGGUUCCACUGC____AGCUACCCAGUUGUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAA ...............................((((((((.....(((.....(((((((((.((((.((....(((......))))).))))))))))))).)))....))))))))... (-21.34 = -22.22 + 0.88)
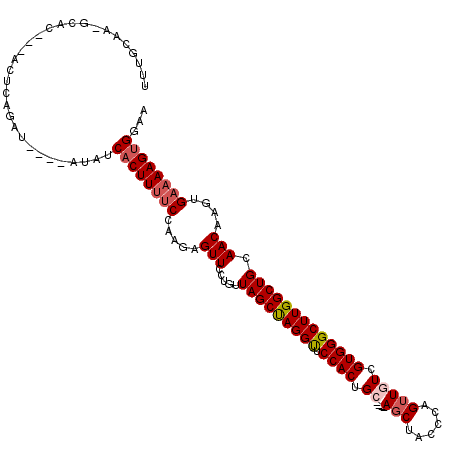



| Location | 17,315,795 – 17,315,897 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 93.63 |
| Mean single sequence MFE | -31.82 |
| Consensus MFE | -27.19 |
| Energy contribution | -27.88 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.39 |
| SVM RNA-class probability | 0.949034 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17315795 102 + 22224390 AAAAGUUCCUAUUAGCUAGGUUCCACUGCAGCUACCCAGUUGUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAAGAUAGCGAAAGAGAAAGGGGAAAGGG .....(((((....((((..((((((((((((((((((.......))))..)))))))..(....)...)))))))..))))...........))))).... ( -32.96) >DroSec_CAF1 32962 102 + 1 AAGAGUUCCAGUUAGCUAGGUUCCACUGCAGCUACCCAGUUGUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAAGAUAGCGAAAGAGAAAGGGGAAAGGG .....((((..((.((((..((((((((((((((((((.......))))..)))))))..(....)...)))))))..)))).......))..))))..... ( -32.10) >DroSim_CAF1 22413 102 + 1 AAGAGUUCCUGUUAGCUAGGUUCCACUGCAGCUACCCAGUUGUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAAGAUAGCGAAAGAGAAAGGGGAAAGGG .....(((((.((.((((..((((((((((((((((((.......))))..)))))))..(....)...)))))))..))))........)).))))).... ( -33.70) >DroEre_CAF1 33993 102 + 1 ACGAGUUCUUGCUAGCCAGGCUCCACUGCAGCUACGUUGGUUUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAAGAUAGCGAAAGAGAAAGGGGAAAGGG .((...((((((.((((((((.((((.(.((((.....)))).)))))))))))))))..((..(....).)).))))...))................... ( -28.50) >consensus AAGAGUUCCUGUUAGCUAGGUUCCACUGCAGCUACCCAGUUGUCGUGGGCUUGGCUGCAACAAGUGAAAAGUGGAAGAUAGCGAAAGAGAAAGGGGAAAGGG .....(((((.((.((((..((((((((((((((((((.......))))..)))))))..(....)...)))))))..)))).....))...)))))..... (-27.19 = -27.88 + 0.69)



| Location | 17,315,795 – 17,315,897 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 93.63 |
| Mean single sequence MFE | -26.97 |
| Consensus MFE | -26.30 |
| Energy contribution | -26.30 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.15 |
| Mean z-score | -3.38 |
| Structure conservation index | 0.98 |
| SVM decision value | 2.94 |
| SVM RNA-class probability | 0.997820 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17315795 102 - 22224390 CCCUUUCCCCUUUCUCUUUCGCUAUCUUCCACUUUUCACUUGUUGCAGCCAAGCCCACGACAACUGGGUAGCUGCAGUGGAACCUAGCUAAUAGGAACUUUU ...........((((..((.((((..(((((((...........(((((...(((((.......))))).))))))))))))..)))).))..))))..... ( -29.40) >DroSec_CAF1 32962 102 - 1 CCCUUUCCCCUUUCUCUUUCGCUAUCUUCCACUUUUCACUUGUUGCAGCCAAGCCCACGACAACUGGGUAGCUGCAGUGGAACCUAGCUAACUGGAACUCUU ...........((((..((.((((..(((((((...........(((((...(((((.......))))).))))))))))))..)))).))..))))..... ( -28.40) >DroSim_CAF1 22413 102 - 1 CCCUUUCCCCUUUCUCUUUCGCUAUCUUCCACUUUUCACUUGUUGCAGCCAAGCCCACGACAACUGGGUAGCUGCAGUGGAACCUAGCUAACAGGAACUCUU ...........((((..((.((((..(((((((...........(((((...(((((.......))))).))))))))))))..)))).))..))))..... ( -28.60) >DroEre_CAF1 33993 102 - 1 CCCUUUCCCCUUUCUCUUUCGCUAUCUUCCACUUUUCACUUGUUGCAGCCAAGCCCACGAAACCAACGUAGCUGCAGUGGAGCCUGGCUAGCAAGAACUCGU .......................................((.(((((((((.((((((..................)))).)).))))).)))).))..... ( -21.47) >consensus CCCUUUCCCCUUUCUCUUUCGCUAUCUUCCACUUUUCACUUGUUGCAGCCAAGCCCACGACAACUGGGUAGCUGCAGUGGAACCUAGCUAACAGGAACUCUU ...........((((..((.((((..(((((((...........(((((...(((((.......))))).))))))))))))..)))).))..))))..... (-26.30 = -26.30 + 0.00)


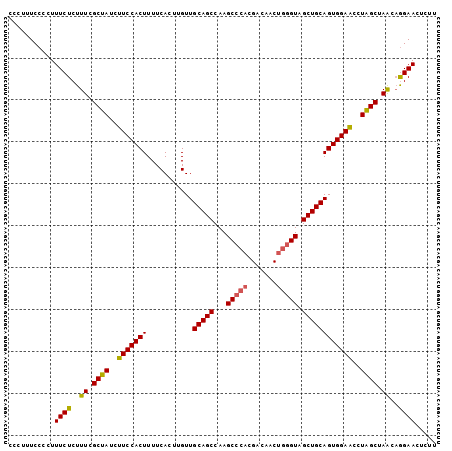
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:25:58 2006