
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,298,227 – 17,298,333 |
| Length | 106 |
| Max. P | 0.955324 |

| Location | 17,298,227 – 17,298,333 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.25 |
| Mean single sequence MFE | -40.40 |
| Consensus MFE | -28.79 |
| Energy contribution | -28.93 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.847969 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17298227 106 + 22224390 -----GAUGCGGAACC------UGUUG---CGCUUCCGGUGGCGGCCGUGCGCCAUUUGUCAAACGACAAAUGCUGCUUGGCCUUGUUGAAGGUGGAACGCAAUUGCGAAUUGCGUUGAU -----(((((((....------.(((.---(((((.(((..(.(((((.(((.((((((((....)))))))).))).))))))..))).))))).)))((....))...)))))))... ( -42.80) >DroVir_CAF1 12285 114 + 1 UGUUUGCGUUGGCGCU------AGAGGACGAAUUCGUAGAGGCAGCCGUGCGCCAUUUGUCAAACGACAAAUGCUGCUUGGCCUUGUUGAAGGUGGAACGCAAUUGGGAAUUGCGUUGGU ...(..(.(..(((((------.....(((....)))...))).((((.(((.((((((((....)))))))).))).))))...))..)..)..)((((((((.....))))))))... ( -37.90) >DroPse_CAF1 22900 112 + 1 -----GAAGCGGGUCCUGUGCCUGUCC---CUGUGCCGGUGCCGGCCGUGCGCCAUUUGUCAAAGGACAAAUGCUGCUUGGCCUUGUUGAAGGUGGAACGCAAUUGCGAAUUGCGUUGAU -----...(((((..(.......)..)---))))(((...((.(((((.(((.((((((((....)))))))).))).)))))..))....)))..((((((((.....))))))))... ( -41.60) >DroWil_CAF1 7160 106 + 1 -----GAAGACGAUCC------AGAGC---CAGACCCAGUCGCAGCCGUGCGCCAUUUGUCAAAGGACAAAUGCUGCUUGGCCUUGUAAAGGGUUGAACGCAAUUGGGAAUUGCGUUGAU -----.....((((((------.....---..(((...)))(((((((.(((.((((((((....)))))))).))).))))..)))...))))))((((((((.....))))))))... ( -36.90) >DroMoj_CAF1 23088 110 + 1 -GUUCGGAGUGGCGCU------AGAGG---AGCUCGCCGAAGCAGCCGUGCGCCAUUUGUCAAACGACAAAUGCUGCUUGGCCUUGUUGAAGGUGGAACGCAAUUGCGAAUUGCGUUGGU -(((((.(((.(((((------.....---)))(((((..((((((((.(((.((((((((....)))))))).))).))))..))))...)))))...)).))).)))))......... ( -39.90) >DroAna_CAF1 16391 111 + 1 UUGUGGAGCCGGAACC------UGUGG---UGCUUCCGAUGGCGGCCGUGCGCCAUUUGUCGAAGGACAAAUGCUGCUUGGCCUUGUUGAAGGUGGAACGCAAUUGGGAAUUGCGUUGGU .......(((...(((------(....---.(((......)))(((((.(((.((((((((....)))))))).))).))))).......))))..((((((((.....))))))))))) ( -43.30) >consensus _____GAAGCGGAACC______AGAGG___CGCUCCCGGUGGCAGCCGUGCGCCAUUUGUCAAACGACAAAUGCUGCUUGGCCUUGUUGAAGGUGGAACGCAAUUGCGAAUUGCGUUGAU ................................((.((...((((((((.(((.((((((((....)))))))).))).))))..))))...)).))((((((((.....))))))))... (-28.79 = -28.93 + 0.14)
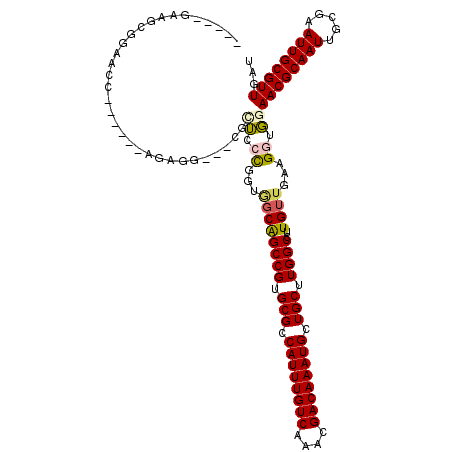


| Location | 17,298,227 – 17,298,333 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.25 |
| Mean single sequence MFE | -32.92 |
| Consensus MFE | -25.92 |
| Energy contribution | -25.67 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.955324 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17298227 106 - 22224390 AUCAACGCAAUUCGCAAUUGCGUUCCACCUUCAACAAGGCCAAGCAGCAUUUGUCGUUUGACAAAUGGCGCACGGCCGCCACCGGAAGCG---CAACA------GGUUCCGCAUC----- ...((((((((.....)))))))).............((((..((..((((((((....))))))))..))..)))).....(((((.(.---.....------).)))))....----- ( -34.60) >DroVir_CAF1 12285 114 - 1 ACCAACGCAAUUCCCAAUUGCGUUCCACCUUCAACAAGGCCAAGCAGCAUUUGUCGUUUGACAAAUGGCGCACGGCUGCCUCUACGAAUUCGUCCUCU------AGCGCCAACGCAAACA ...((((((((.....))))))))...((((....)))).......(((((((((....)))))))(((((..((...))...(((....))).....------.)))))...))..... ( -29.10) >DroPse_CAF1 22900 112 - 1 AUCAACGCAAUUCGCAAUUGCGUUCCACCUUCAACAAGGCCAAGCAGCAUUUGUCCUUUGACAAAUGGCGCACGGCCGGCACCGGCACAG---GGACAGGCACAGGACCCGCUUC----- ....(((((((.....)))))))(((..(((....)))(((..((..((((((((....))))))))..))...((((....))))....---.....)))...)))........----- ( -33.60) >DroWil_CAF1 7160 106 - 1 AUCAACGCAAUUCCCAAUUGCGUUCAACCCUUUACAAGGCCAAGCAGCAUUUGUCCUUUGACAAAUGGCGCACGGCUGCGACUGGGUCUG---GCUCU------GGAUCGUCUUC----- (((((((((((.....))))))))..........((..((((..(..((((((((....)))))))).((((....)))).....)..))---))..)------)))).......----- ( -31.80) >DroMoj_CAF1 23088 110 - 1 ACCAACGCAAUUCGCAAUUGCGUUCCACCUUCAACAAGGCCAAGCAGCAUUUGUCGUUUGACAAAUGGCGCACGGCUGCUUCGGCGAGCU---CCUCU------AGCGCCACUCCGAAC- ...((((((((.....))))))))...((((....)))).(((((.((....)).))))).....((((((..((((((....)).))))---.....------.))))))........- ( -32.80) >DroAna_CAF1 16391 111 - 1 ACCAACGCAAUUCCCAAUUGCGUUCCACCUUCAACAAGGCCAAGCAGCAUUUGUCCUUCGACAAAUGGCGCACGGCCGCCAUCGGAAGCA---CCACA------GGUUCCGGCUCCACAA ...((((((((.....)))))))).............((((..((..((((((((....))))))))..))..))))(((...((((.(.---.....------).)))))))....... ( -35.60) >consensus ACCAACGCAAUUCCCAAUUGCGUUCCACCUUCAACAAGGCCAAGCAGCAUUUGUCCUUUGACAAAUGGCGCACGGCCGCCACCGGGAGCG___CCACA______GGAUCCGCUUC_____ ...((((((((.....)))))))).............((((..((..((((((((....))))))))..))..))))........................................... (-25.92 = -25.67 + -0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:25:49 2006