
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,954,214 – 16,954,318 |
| Length | 104 |
| Max. P | 0.998796 |

| Location | 16,954,214 – 16,954,318 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.71 |
| Mean single sequence MFE | -45.02 |
| Consensus MFE | -36.20 |
| Energy contribution | -38.08 |
| Covariance contribution | 1.87 |
| Combinations/Pair | 1.19 |
| Mean z-score | -3.44 |
| Structure conservation index | 0.80 |
| SVM decision value | 3.23 |
| SVM RNA-class probability | 0.998796 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16954214 104 + 22224390 G--UGGGAAAAAUGGGGGCGUGGUGCCAAAAGUGGGUUGCAUUUUGAUAUGCACAAGGCA------------CCACUUUUCACCACUCCCCCCCUGUGUCCCAGCA--AAUUUAUUUAUG .--(((((.....(((((.((((((..(((((((((((((((......)))))....)).------------)))))))))))))).)))))......)))))...--............ ( -44.60) >DroSec_CAF1 31397 114 + 1 G--UGGGAA-AAUGGGGGCGUGGUACCAAAAGUGGGUUGCAUUUUGAUAUGCACAAGGCAUGGGCGUGCGACCCCCUUUUCACCACUCCCCGC---UGUCCCAGCAUAAAUUUAUUUACG .--(((((.-..((((((.(((((...(((((.(((((((((.....(((((.....)))))...))))))))).))))).))))))))))).---..)))))................. ( -48.60) >DroEre_CAF1 18701 114 + 1 GGGGGGAGA-AAUGGGGGCGUGGUGGCAAAAGUGGGUUGCAUUUUGAUAUGCACAAGGCAUGGGCGUGCGACCCCCUUUUCAGCACUUCCCAC---UGUGUUUGCU--AAUUUAUUUAUG .((.(((.(-..((((((.(((.(((..((((.(((((((((.....(((((.....)))))...))))))))).))))))).))))))))).---..).))).))--............ ( -39.00) >DroYak_CAF1 18172 114 + 1 GUGGGGGGA-AACGGGGGCGUGGUGCCAAAAGUGGGUUGCAUUUUGAUAUGCACAAGACAUGGGCGUGCGACCCCCUUUUCACCACUCCCCGC---UGUCUCUGCU--AAUUUAUUUAUG ..(.(((((-..((((((.((((((..(((((.(((((((((.....(((((....).))))...))))))))).))))))))))))))))).---..))))).).--............ ( -47.90) >consensus G__GGGGAA_AAUGGGGGCGUGGUGCCAAAAGUGGGUUGCAUUUUGAUAUGCACAAGGCAUGGGCGUGCGACCCCCUUUUCACCACUCCCCGC___UGUCCCAGCA__AAUUUAUUUAUG ...(((((....((((((.((((((..(((((.(((((((((.....(((((.....)))))...))))))))).)))))))))))))))))......)))))................. (-36.20 = -38.08 + 1.87)


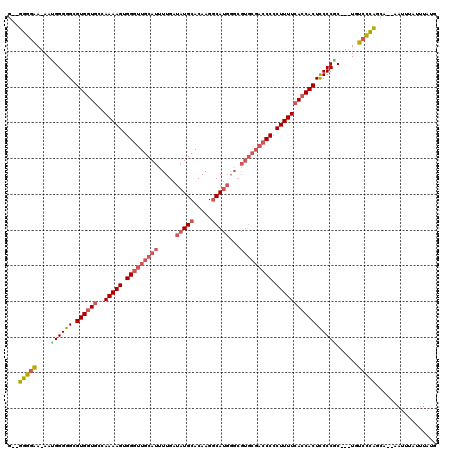
| Location | 16,954,214 – 16,954,318 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.71 |
| Mean single sequence MFE | -37.60 |
| Consensus MFE | -25.88 |
| Energy contribution | -27.50 |
| Covariance contribution | 1.62 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.94 |
| Structure conservation index | 0.69 |
| SVM decision value | 2.23 |
| SVM RNA-class probability | 0.990710 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16954214 104 - 22224390 CAUAAAUAAAUU--UGCUGGGACACAGGGGGGGAGUGGUGAAAAGUGG------------UGCCUUGUGCAUAUCAAAAUGCAACCCACUUUUGGCACCACGCCCCCAUUUUUCCCA--C ............--...(((((......(((((.((((((.(((((((------------.(.....(((((......))))).))))))))...)))))).))))).....)))))--. ( -42.60) >DroSec_CAF1 31397 114 - 1 CGUAAAUAAAUUUAUGCUGGGACA---GCGGGGAGUGGUGAAAAGGGGGUCGCACGCCCAUGCCUUGUGCAUAUCAAAAUGCAACCCACUUUUGGUACCACGCCCCCAUU-UUCCCA--C ((((((....)))))).(((((.(---(.((((.((((((.(((((((((.(((.....((((.....)))).......))).)))).)))))..))))))..)))).))-.)))))--. ( -42.10) >DroEre_CAF1 18701 114 - 1 CAUAAAUAAAUU--AGCAAACACA---GUGGGAAGUGCUGAAAAGGGGGUCGCACGCCCAUGCCUUGUGCAUAUCAAAAUGCAACCCACUUUUGCCACCACGCCCCCAUU-UCUCCCCCC ............--.........(---(((((..(((.((.(((((((((.(((.....((((.....)))).......))).)))).)))))..)).)))...))))))-......... ( -27.20) >DroYak_CAF1 18172 114 - 1 CAUAAAUAAAUU--AGCAGAGACA---GCGGGGAGUGGUGAAAAGGGGGUCGCACGCCCAUGUCUUGUGCAUAUCAAAAUGCAACCCACUUUUGGCACCACGCCCCCGUU-UCCCCCCAC ............--....(((((.---(.((((.((((((.(((((((((.(((.....((((.....)))).......))).)))).)))))..)))))).))))))))-))....... ( -38.50) >consensus CAUAAAUAAAUU__AGCAGAGACA___GCGGGGAGUGGUGAAAAGGGGGUCGCACGCCCAUGCCUUGUGCAUAUCAAAAUGCAACCCACUUUUGGCACCACGCCCCCAUU_UCCCCA__C ...........................(.((((.(((((((((((.((((.(((.....((((.....)))).......))).)))).)))))..)))))).)))))............. (-25.88 = -27.50 + 1.62)

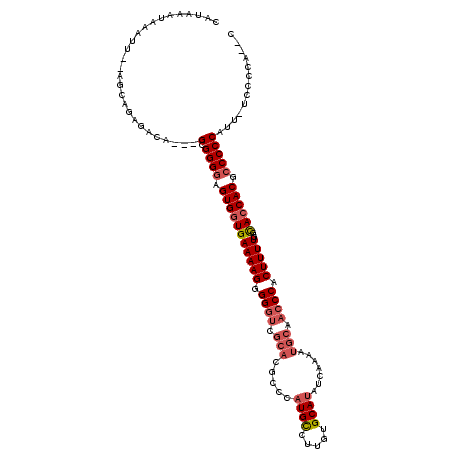

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:23:32 2006