
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,855,269 – 16,855,372 |
| Length | 103 |
| Max. P | 0.762331 |

| Location | 16,855,269 – 16,855,372 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 84.14 |
| Mean single sequence MFE | -21.32 |
| Consensus MFE | -13.47 |
| Energy contribution | -14.66 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.63 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.505531 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16855269 103 + 22224390 AAGAGCGGUACUCAU---CAUGCCCCGCCCAUCGUCCUCCGGUGAUGCAUAUAAGUCGUAACUAUCCCCACUCCCACCUGAGGCACCUGGAUCACCUGCACCAUCA ..((((((..(....---...)..))))(((..((((((.((((.........(((....)))...........)))).)))).)).))).))............. ( -20.35) >DroSec_CAF1 53950 103 + 1 AUGAGCGGGACUCAU---CAUGCCCCGCCCAUCGUCCUUCUGUGAUGCAUAUAAGUCGUAACUAUCCCCACUCCCACCUGCGGCACCUGCAUCACCUGCACCAUCA (((.(((((.(....---...).))))).))).........((((((((.....((((((..................))))))...))))))))........... ( -26.07) >DroEre_CAF1 39235 94 + 1 AUGAGCGGGACUCAU---CUUGCUCCGCCCAUCGUUCUCCGGUGAUGCAUACAAGCCCUAACUAUCCCCACUCCCACC---------UGCAUCACCUGCACCAUCA ..((((((((....)---)))))))...............(((((((((.............................---------))))))))).......... ( -22.45) >DroYak_CAF1 38692 97 + 1 AUGAGCGGAACUCAUCAUCAUGACCCGCCCAUCGUUCUCCGAUGAUGCAUAUAAGCCGUAACUUUGCCCACUUCCACC---------UGCAUCACCUGCACCAUCA (((.((((...((((....)))).)))).)))........((((.((((...(((......)))(((...........---------.))).....)))).)))). ( -16.40) >consensus AUGAGCGGGACUCAU___CAUGCCCCGCCCAUCGUCCUCCGGUGAUGCAUAUAAGCCGUAACUAUCCCCACUCCCACC_________UGCAUCACCUGCACCAUCA ..(.(((((.(..........).))))).)..........(((((((((......................................))))))))).......... (-13.47 = -14.66 + 1.19)

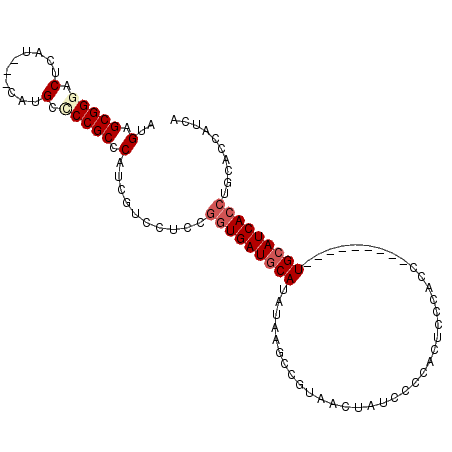

| Location | 16,855,269 – 16,855,372 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 84.14 |
| Mean single sequence MFE | -33.55 |
| Consensus MFE | -26.42 |
| Energy contribution | -27.23 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.762331 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16855269 103 - 22224390 UGAUGGUGCAGGUGAUCCAGGUGCCUCAGGUGGGAGUGGGGAUAGUUACGACUUAUAUGCAUCACCGGAGGACGAUGGGCGGGGCAUG---AUGAGUACCGCUCUU (((.((..(.((....))..)..)))))...(((((((((........).((((((((((..(.(((........)))..)..)))).---)))))).)))))))) ( -31.00) >DroSec_CAF1 53950 103 - 1 UGAUGGUGCAGGUGAUGCAGGUGCCGCAGGUGGGAGUGGGGAUAGUUACGACUUAUAUGCAUCACAGAAGGACGAUGGGCGGGGCAUG---AUGAGUCCCGCUCAU .....((.(..((((((((....(((....)))((((.(.........).))))...))))))))....).)).(((((((((((...---....))))))))))) ( -38.40) >DroEre_CAF1 39235 94 - 1 UGAUGGUGCAGGUGAUGCA---------GGUGGGAGUGGGGAUAGUUAGGGCUUGUAUGCAUCACCGGAGAACGAUGGGCGGAGCAAG---AUGAGUCCCGCUCAU .....((.(.(((((((((---------.(..((..(((......)))...))..).))))))))).)...)).((((((((..(...---....)..)))))))) ( -31.80) >DroYak_CAF1 38692 97 - 1 UGAUGGUGCAGGUGAUGCA---------GGUGGAAGUGGGCAAAGUUACGGCUUAUAUGCAUCAUCGGAGAACGAUGGGCGGGUCAUGAUGAUGAGUUCCGCUCAU .(((((((((.((((.((.---------.((((...(....)...)))).)))))).)))))))))........(((((((((((((....))))..))))))))) ( -33.00) >consensus UGAUGGUGCAGGUGAUGCA_________GGUGGGAGUGGGGAUAGUUACGACUUAUAUGCAUCACCGGAGAACGAUGGGCGGGGCAUG___AUGAGUCCCGCUCAU ..((.((.(.(((((((((..............((((.(.........).))))...)))))))))...).)).))(((((((((..........))))))))).. (-26.42 = -27.23 + 0.81)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:22:45 2006