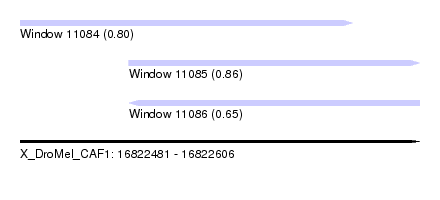

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,822,481 – 16,822,606 |
| Length | 125 |
| Max. P | 0.861834 |

| Location | 16,822,481 – 16,822,585 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 88.74 |
| Mean single sequence MFE | -25.06 |
| Consensus MFE | -19.54 |
| Energy contribution | -19.38 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.804416 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16822481 104 + 22224390 GCAAUGUUGCAAAAUUAAUGCGAUAUCCAAUGCGAUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU--------ACAGGCAAAAAAUGGGAGCUACAACAAGUG ((..(((((((.......)))))))((((.((((.((((....))))).(((((.((((......)))))))))--------....))).....)))).))........... ( -23.70) >DroSec_CAF1 20187 89 + 1 GCAAUGUUGCAAAAUUAAUGCGAUAUCCAAUGCGAUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU--------ACAGGCAGAA---------------AAGUG ...((((((((.......))))))))....((((.((((....))))).(((((.((((......)))))))))--------....)))...---------------..... ( -20.90) >DroSim_CAF1 4935 103 + 1 GCAAUGUUGCAAAAUUAAUGCGAUAUCCAAUGCGAUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU--------AGAGGCAGAA-AUGGGAGCUACAACAAGUG ((..(((((((.......)))))))((((.((((.((((....))))).(((((.((((......)))))))))--------....)))...-.)))).))........... ( -25.50) >DroEre_CAF1 4907 103 + 1 GCAAUGUUGCAAAAUUAAUGCGAUAUCCAAUGCGAUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU--------ACAGGCAGCA-AUGGGAGCUACAACAAGUG ((..(((((((.......)))))))((((.((((.((((....))))).(((((.((((......)))))))))--------.......)))-.)))).))........... ( -27.60) >DroYak_CAF1 5015 111 + 1 GCAAUGUUGCAAAAUUAAUGCGAUAUCCAAUGCGAUUACAGAUGUGACAACAACAUGGGCAAAUAUCCAGUUGUGCUGUUGUGCUGGCUGAA-AUGGGAGCUACAACAAGUG (((((((((((.......))))))))....)))(.((((....))))).(((((.((((......)))))))))(((((((((((..(....-..)..))).))))).))). ( -27.60) >consensus GCAAUGUUGCAAAAUUAAUGCGAUAUCCAAUGCGAUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU________ACAGGCAGAA_AUGGGAGCUACAACAAGUG ((..(((..((....(((((((........))).))))....))..)))(((((.((((......)))))))))............))........................ (-19.54 = -19.38 + -0.16)



| Location | 16,822,515 – 16,822,606 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 87.18 |
| Mean single sequence MFE | -22.48 |
| Consensus MFE | -17.76 |
| Energy contribution | -18.20 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.861834 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16822515 91 + 22224390 AUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU--------ACAGGCAAAAAAUGGGAGCUACAACAAGUGUUACAACCACUUGAAGUUGGU ........((((((((((((.((((......)))))))))--------...(((...........)))........)))))))((((........)))) ( -20.90) >DroSec_CAF1 20221 76 + 1 AUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU--------ACAGGCAGAA---------------AAGUGUUACAACCACUUGAAGUUGGU ........((((((.(((((.((((......)))))))))--------....((....---------------..))))))))((((........)))) ( -18.30) >DroSim_CAF1 4969 90 + 1 AUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU--------AGAGGCAGAA-AUGGGAGCUACAACAAGUGUUACAACCACUUGAAGUUGGU ........(((((((......((((......))))(((((--------((...((...-.))....)))))))...)))))))((((........)))) ( -25.10) >DroEre_CAF1 4941 90 + 1 AUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU--------ACAGGCAGCA-AUGGGAGCUACAACAAGUGUUACAACCACUUGAAGUUGGU ........((((((((((((.((((......)))))))))--------....(.(((.-......))).)......)))))))((((........)))) ( -21.70) >DroYak_CAF1 5049 98 + 1 AUUACAGAUGUGACAACAACAUGGGCAAAUAUCCAGUUGUGCUGUUGUGCUGGCUGAA-AUGGGAGCUACAACAAGUGUUACAACCACUUGAAGUUGGU ........((((((((((((.((((......)))))))))..(((((((((..(....-..)..))).))))))..)))))))((((........)))) ( -26.40) >consensus AUUACAGAUGUGACAACAACAUGGGCAAAUACCCAGUUGU________ACAGGCAGAA_AUGGGAGCUACAACAAGUGUUACAACCACUUGAAGUUGGU ........((((((((((((.((((......)))))))))...........(((...........)))........)))))))((((........)))) (-17.76 = -18.20 + 0.44)

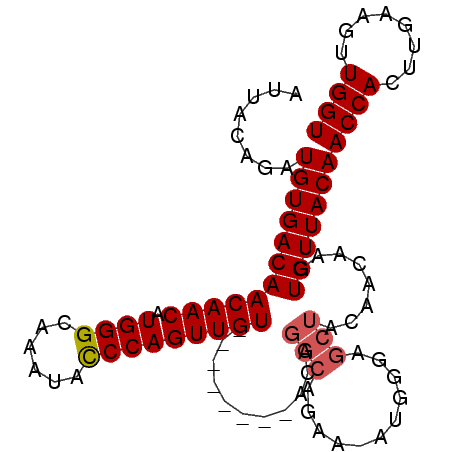

| Location | 16,822,515 – 16,822,606 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 87.18 |
| Mean single sequence MFE | -20.36 |
| Consensus MFE | -15.88 |
| Energy contribution | -16.68 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.652728 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16822515 91 - 22224390 ACCAACUUCAAGUGGUUGUAACACUUGUUGUAGCUCCCAUUUUUUGCCUGU--------ACAACUGGGUAUUUGCCCAUGUUGUUGUCACAUCUGUAAU ....((..((((((.......))))))..))............((((.(((--------((((((((((....))))).)))))....)))...)))). ( -20.80) >DroSec_CAF1 20221 76 - 1 ACCAACUUCAAGUGGUUGUAACACUU---------------UUCUGCCUGU--------ACAACUGGGUAUUUGCCCAUGUUGUUGUCACAUCUGUAAU .........(((((.......)))))---------------.......((.--------((((((((((....))))).)))))...)).......... ( -15.70) >DroSim_CAF1 4969 90 - 1 ACCAACUUCAAGUGGUUGUAACACUUGUUGUAGCUCCCAU-UUCUGCCUCU--------ACAACUGGGUAUUUGCCCAUGUUGUUGUCACAUCUGUAAU ....((..((((((.......))))))..)).........-...((...(.--------((((((((((....))))).))))).)...))........ ( -19.70) >DroEre_CAF1 4941 90 - 1 ACCAACUUCAAGUGGUUGUAACACUUGUUGUAGCUCCCAU-UGCUGCCUGU--------ACAACUGGGUAUUUGCCCAUGUUGUUGUCACAUCUGUAAU ........((((((.......))))))..(((((......-.))))).((.--------((((((((((....))))).)))))...)).......... ( -24.50) >DroYak_CAF1 5049 98 - 1 ACCAACUUCAAGUGGUUGUAACACUUGUUGUAGCUCCCAU-UUCAGCCAGCACAACAGCACAACUGGAUAUUUGCCCAUGUUGUUGUCACAUCUGUAAU ....((..((((((.......))))))..)).........-..(((.....(((((((((....(((........)))))))))))).....))).... ( -21.10) >consensus ACCAACUUCAAGUGGUUGUAACACUUGUUGUAGCUCCCAU_UUCUGCCUGU________ACAACUGGGUAUUUGCCCAUGUUGUUGUCACAUCUGUAAU ....((..((((((.......))))))..))............................((((((((((....))))).)))))............... (-15.88 = -16.68 + 0.80)


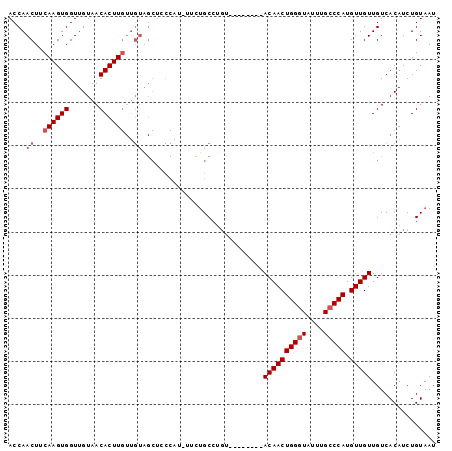
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:22:30 2006