
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,778,872 – 16,778,979 |
| Length | 107 |
| Max. P | 0.981516 |

| Location | 16,778,872 – 16,778,979 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.44 |
| Mean single sequence MFE | -21.80 |
| Consensus MFE | -14.21 |
| Energy contribution | -14.10 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.742049 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16778872 107 + 22224390 A-G-GUUUGUCAAAACAAUAUGGUAAAUGACCAUAAAUCUGUAACCCUUCAGUAGGUACACCUAAAGAUUCUUAUUUAAACGUAUU-----------UCGUUGGCAAAAAGAGUUACAGA .-.-((((....))))..((((((.....))))))..((((((((...((..((((....))))..)).((((.((..((((....-----------.))))...)).)))))))))))) ( -25.30) >DroSec_CAF1 15784 108 + 1 AUG-GCUUGUCAAAACAAUAUGGUAAAUGACCAUAAAUCUGUAACCCUUCAGUAGAUACACCUAAAGAUUCAUAUUUAAACGUAGU-----------UUGUGGGCAAAAAGACUUACAGA ..(-(..((((.......((((((.....))))))...(((........)))..))))..)).......................(-----------(((((((........)))))))) ( -16.40) >DroYak_CAF1 15609 119 + 1 UUGUAUUUGUCAAAAUAAUAUGGUAAAUGACCAUAAAUCUGUAACCCUUUAAUAGGUGCACCUAAAGAUUCUC-CCUAUACACAGUGACAGAUAUAUGUAUGUAUAUACACAGUUACAGA .(((((((((((......((((((.....))))))...((((..........((((....)))).((......-.))....)))))))))))))))(((((....))))).......... ( -23.70) >consensus AUG_GUUUGUCAAAACAAUAUGGUAAAUGACCAUAAAUCUGUAACCCUUCAGUAGGUACACCUAAAGAUUCUUAUUUAAACGUAGU___________UUGUGGGCAAAAAGAGUUACAGA ..................((((((.....))))))..((((((((...((..((((....))))..)).............................(((....))).....)))))))) (-14.21 = -14.10 + -0.11)

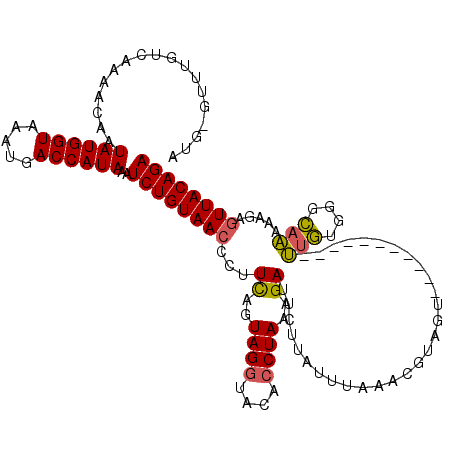
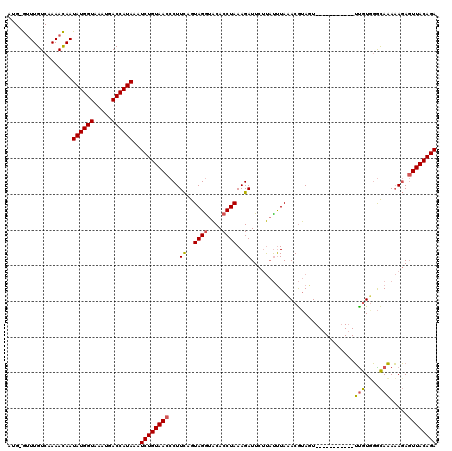
| Location | 16,778,872 – 16,778,979 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.44 |
| Mean single sequence MFE | -24.69 |
| Consensus MFE | -18.99 |
| Energy contribution | -18.00 |
| Covariance contribution | -0.99 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.89 |
| SVM RNA-class probability | 0.981516 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16778872 107 - 22224390 UCUGUAACUCUUUUUGCCAACGA-----------AAUACGUUUAAAUAAGAAUCUUUAGGUGUACCUACUGAAGGGUUACAGAUUUAUGGUCAUUUACCAUAUUGUUUUGACAAAC-C-U (((((((((((((.....((((.-----------....))))..............((((....))))..)))))))))))))..((((((.....))))))((((....))))..-.-. ( -25.90) >DroSec_CAF1 15784 108 - 1 UCUGUAAGUCUUUUUGCCCACAA-----------ACUACGUUUAAAUAUGAAUCUUUAGGUGUAUCUACUGAAGGGUUACAGAUUUAUGGUCAUUUACCAUAUUGUUUUGACAAGC-CAU (((((((.((((((((....)))-----------..((((..((((........))))..))))......))))).)))))))..((((((.....))))))((((....))))..-... ( -20.90) >DroYak_CAF1 15609 119 - 1 UCUGUAACUGUGUAUAUACAUACAUAUAUCUGUCACUGUGUAUAGG-GAGAAUCUUUAGGUGCACCUAUUAAAGGGUUACAGAUUUAUGGUCAUUUACCAUAUUAUUUUGACAAAUACAA ..((((..(((((....)))))((((.((((((.(((.(..(((((-....(((....)))...)))))...).))).)))))).))))((((...............))))...)))). ( -27.26) >consensus UCUGUAACUCUUUUUGCCCACAA___________ACUACGUUUAAAUAAGAAUCUUUAGGUGUACCUACUGAAGGGUUACAGAUUUAUGGUCAUUUACCAUAUUGUUUUGACAAAC_CAU (((((((((((((.((....))..............((((..((((........))))..))))......)))))))))))))..((((((.....)))))).................. (-18.99 = -18.00 + -0.99)

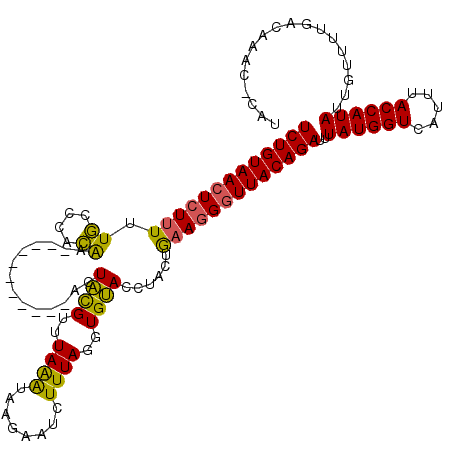
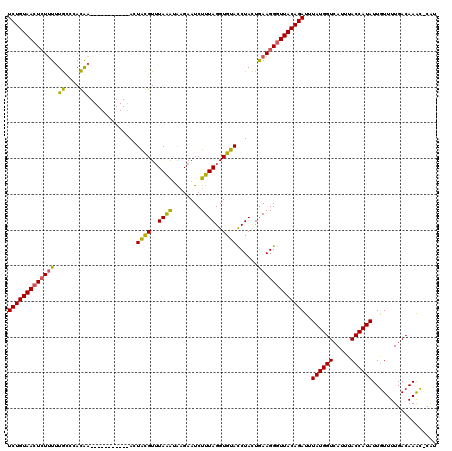
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:22:11 2006