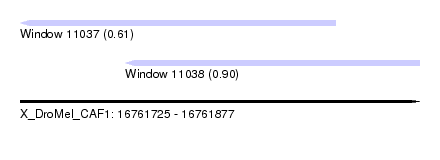

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,761,725 – 16,761,877 |
| Length | 152 |
| Max. P | 0.899168 |

| Location | 16,761,725 – 16,761,845 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.08 |
| Mean single sequence MFE | -17.73 |
| Consensus MFE | -10.49 |
| Energy contribution | -12.80 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.607792 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16761725 120 - 22224390 CCAGGCAUUGGCAUAGAAAUUCUCAAAGGAAAUAAGCACAGCAAAUUAACGAUUGCUAUCAUUAAUGCUAACAAUAAAUAAUAAAAAAACUCCAGCAUAAAAAUGCUAAAAACUUUAUGA ...(((((((((.((....((((....)))).)).))..(((((........))))).....)))))))........................(((((....)))))............. ( -16.50) >DroSec_CAF1 48475 119 - 1 CCAGACAUUGGCAUAGAAAUUCUCAAAGGAAAUAAGCACAGCAAAUUAACGAUUGCUAUCGUUAAUGCUAACAAUAAAUAAUAAA-AAACUCCAGCAUGAAAAUGCUAAAAACUUUAUGA ((((...))))((((((..........(((.....((...))..(((((((((....)))))))))...................-....)))(((((....)))))......)))))). ( -21.20) >DroSim_CAF1 46417 119 - 1 CCAGACAUUGGCAUAGAAAUUCUCAAAGGAAAUAAGCACAGCAAAUUAACGAUUGCUAUCGUUAAUGCUAACAAUAAAUAAUAAA-AAACUUCAGCAUAAAAAUGCUAAAAACUUUAUGA ((((...))))((((((..((((....))))........((((..((((((((....))))))))))))................-.......(((((....)))))......)))))). ( -18.70) >DroYak_CAF1 40734 98 - 1 CCAGACAUUGGCAUAGAAAUUCUCAAAGGAGAUACGCUCAACAAAUAAUCAAGCG--------------------AAAUAAUCA--AAACUCCAGCAUAAAAAUGCUAAAAACUUUAUGA ((((...))))((((((..........((((....(((.............)))(--------------------(.....)).--...))))(((((....)))))......)))))). ( -14.52) >consensus CCAGACAUUGGCAUAGAAAUUCUCAAAGGAAAUAAGCACAGCAAAUUAACGAUUGCUAUCGUUAAUGCUAACAAUAAAUAAUAAA_AAACUCCAGCAUAAAAAUGCUAAAAACUUUAUGA ((((...))))((((((..........(((.....(.....)..(((((((((....)))))))))........................)))(((((....)))))......)))))). (-10.49 = -12.80 + 2.31)



| Location | 16,761,765 – 16,761,877 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.90 |
| Mean single sequence MFE | -18.95 |
| Consensus MFE | -12.25 |
| Energy contribution | -15.50 |
| Covariance contribution | 3.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.899168 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16761765 112 - 22224390 AUAAAA--------AAAAAAAUUGGGUAAACUUUUUAUGGCCAGGCAUUGGCAUAGAAAUUCUCAAAGGAAAUAAGCACAGCAAAUUAACGAUUGCUAUCAUUAAUGCUAACAAUAAAUA ......--------.......(((((.....((((((((.((((...))))))))))))..))))).............((((..((((.(((....))).))))))))........... ( -16.10) >DroSec_CAF1 48514 112 - 1 AUAAAA--------AAAAAAAUUGGGUAAACUUUUUAUGGCCAGACAUUGGCAUAGAAAUUCUCAAAGGAAAUAAGCACAGCAAAUUAACGAUUGCUAUCGUUAAUGCUAACAAUAAAUA ......--------.......(((((.....((((((((.((((...))))))))))))..))))).............((((..((((((((....))))))))))))........... ( -21.80) >DroSim_CAF1 46456 111 - 1 AUAAAA--------A-AAAAAUUGGGUAAACUUUUUAUGGCCAGACAUUGGCAUAGAAAUUCUCAAAGGAAAUAAGCACAGCAAAUUAACGAUUGCUAUCGUUAAUGCUAACAAUAAAUA ......--------.-.....(((((.....((((((((.((((...))))))))))))..))))).............((((..((((((((....))))))))))))........... ( -21.80) >DroYak_CAF1 40772 100 - 1 AUAAAAACAAAAAAAAAAAAAUUGGGUAAACUUUUUAUGGCCAGACAUUGGCAUAGAAAUUCUCAAAGGAGAUACGCUCAACAAAUAAUCAAGCG--------------------AAAUA .....................((((((....((((((((.((((...)))))))))))).((((....))))...))))))..............--------------------..... ( -16.10) >consensus AUAAAA________AAAAAAAUUGGGUAAACUUUUUAUGGCCAGACAUUGGCAUAGAAAUUCUCAAAGGAAAUAAGCACAGCAAAUUAACGAUUGCUAUCGUUAAUGCUAACAAUAAAUA .....................(((((.....((((((((.((((...))))))))))))..))))).............((((..((((((((....))))))))))))........... (-12.25 = -15.50 + 3.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:21:49 2006