
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,734,128 – 16,734,368 |
| Length | 240 |
| Max. P | 0.977019 |

| Location | 16,734,128 – 16,734,248 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.17 |
| Mean single sequence MFE | -29.71 |
| Consensus MFE | -28.74 |
| Energy contribution | -28.78 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.78 |
| SVM RNA-class probability | 0.977019 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
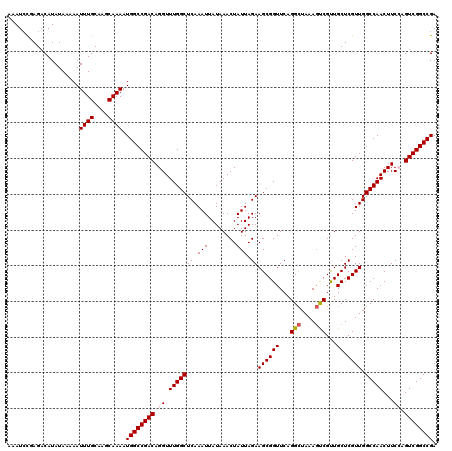
>X_DroMel_CAF1 16734128 120 - 22224390 UCGGCCAUUUUGCUUGCAAAUUUUUAUAUGUCUCAGAUUUUGUUUAUGUUUUCGAUUUUGGAGUUCUUCGAACUCGUCGGGGUUUUUGAGAUGUCAUGAAAAAUUGCCCAACCCGCUGAC (((((....(((...((((.(((((((((((((((((............(..((((....((((((...))))))))))..)..)))))))))).))))))).)))).)))...))))). ( -31.94) >DroSec_CAF1 21859 120 - 1 UCGGCCAUUUUGCUUGCAAAUUUUUAUAUGUCUCGGAUUUUGUUUAUGUUUUCGAUUUCGGAGUUCUUCGAACUCGUCGGGGUUUUUGAGAUCUCAUGAAAAAUUGCCCAACCCGCUGAC (((((....(((...((((.(((((((..((((((((............(..((((....((((((...))))))))))..)..))))))))...))))))).)))).)))...))))). ( -28.84) >DroSim_CAF1 19240 120 - 1 UCGGCCAUUUUGCUUGCAAAUUUUUAUAUGUCUCGGAUUUUGUUUAUGUUUUCGAUUUCGGAGUUCUUCGAACUCGUCGGGGUUUUUGAGAUCUCAUGAAAAAUUGCCCAACCCGCUGAC (((((....(((...((((.(((((((..((((((((............(..((((....((((((...))))))))))..)..))))))))...))))))).)))).)))...))))). ( -28.84) >DroEre_CAF1 13252 120 - 1 UCGGCCAUUUUGCUUGCAAAUUUUUAUAUGUCUCGGAUUUUGUUUAUGUUUUCGAUUUCGGAGUUCUUCGAACUCGUCGGGGUUUUUGAGAUCUCAUGAAAAAUUGCUCAACCCGCUGAC (((((....(((...((((.(((((((..((((((((............(..((((....((((((...))))))))))..)..))))))))...))))))).)))).)))...))))). ( -29.54) >DroYak_CAF1 13011 120 - 1 UCGGCCAUUUUGCUUGCAAAUUUUUAUAUGUCUCGCAUUUUGUUUAUGUUUUCGAUUUUGGAGUUCUUCGAACUCGUCGGGGUUUUUGAGAUCUCAUGAAAAAUUGCCCAACCCGCUGAC (((((....(((...((((.(((((((..(((((((((.......))))(..((((....((((((...))))))))))..).....)))))...))))))).)))).)))...))))). ( -29.40) >consensus UCGGCCAUUUUGCUUGCAAAUUUUUAUAUGUCUCGGAUUUUGUUUAUGUUUUCGAUUUCGGAGUUCUUCGAACUCGUCGGGGUUUUUGAGAUCUCAUGAAAAAUUGCCCAACCCGCUGAC (((((....(((...((((.(((((((..((((((((............(..((((....((((((...))))))))))..)..))))))))...))))))).)))).)))...))))). (-28.74 = -28.78 + 0.04)

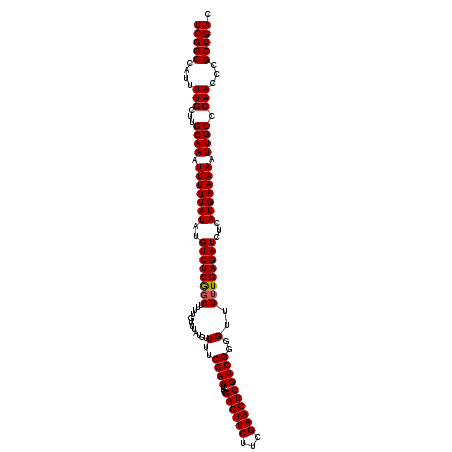

| Location | 16,734,208 – 16,734,328 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.42 |
| Mean single sequence MFE | -32.01 |
| Consensus MFE | -30.14 |
| Energy contribution | -30.58 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963221 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
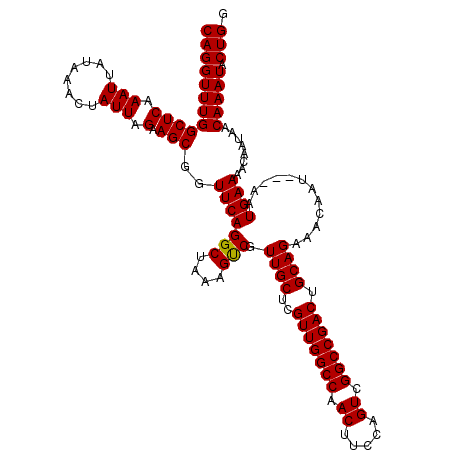
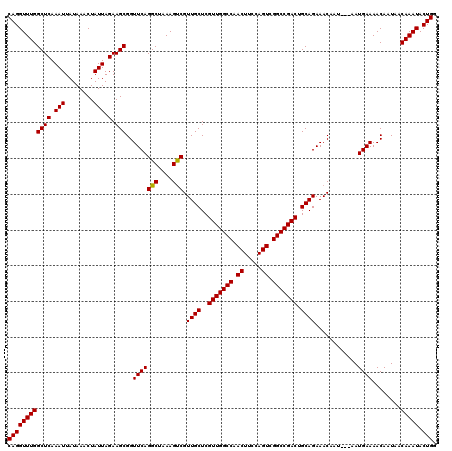
>X_DroMel_CAF1 16734208 120 + 22224390 AAAUCUGAGACAUAUAAAAAUUUGCAAGCAAAAUGGCCGACAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGCUAAAGCCGUUGCUCGUUGGCCAACUUCCAGUCGGCCGA ....................((((....)))).((((((((.((.(((((((.(((........))).))((((((...(((....)))...)).)))).)))))...)).)))))))). ( -34.90) >DroSec_CAF1 21939 120 + 1 AAAUCCGAGACAUAUAAAAAUUUGCAAGCAAAAUGGCCGACAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGCUAAAGUCGUUGCUCGUUGGCCAACUUCCAGUCGGCCGA ....................((((....)))).((((((((.((.(((((((.(((........))).))((((((...(((....)))...)).)))).)))))...)).)))))))). ( -32.20) >DroSim_CAF1 19320 120 + 1 AAAUCCGAGACAUAUAAAAAUUUGCAAGCAAAAUGGCCGACAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGCUAAAGUCGUUGCUCGUUGGCCAACUUCCAGUCGGCCGA ....................((((....)))).((((((((.((.(((((((.(((........))).))((((((...(((....)))...)).)))).)))))...)).)))))))). ( -32.20) >DroEre_CAF1 13332 120 + 1 AAAUCCGAGACAUAUAAAAAUUUGCAAGCAAAAUGGCCGACAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGUUAAAUUCGCUGCUCGUUGGCCAACUUCGAGUCGGCCGA ....................((((....)))).((((((((.((.(((((.(((..............((.(((((...((......))))))))).))))))))..))..)))))))). ( -29.36) >DroYak_CAF1 13091 120 + 1 AAAUGCGAGACAUAUAAAAAUUUGCAAGCAAAAUGGCCGACAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGCUAAAGUCGUUGCUCGUUGGCCAACUUCAAGUCGGCCGA ...((((((...........)))))).......(((((((((((((.(((((.(((........))).))((((((...(((....)))...)).)))).))))))))...)))))))). ( -31.40) >consensus AAAUCCGAGACAUAUAAAAAUUUGCAAGCAAAAUGGCCGACAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGCUAAAGUCGUUGCUCGUUGGCCAACUUCCAGUCGGCCGA ....................((((....)))).((((((((.((.(((((((.(((........))).))((((((...(((....)))...)).)))).)))))...)).)))))))). (-30.14 = -30.58 + 0.44)


| Location | 16,734,248 – 16,734,368 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.64 |
| Mean single sequence MFE | -30.00 |
| Consensus MFE | -29.62 |
| Energy contribution | -29.40 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820554 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16734248 120 + 22224390 CAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGCUAAAGCCGUUGCUCGUUGGCCAACUUCCAGUCGGCCGACUGCAGAAACAAUAAUAAUGAAAACAAUAACAAAUACUGG ((((((((((((.(((........))).).)))..(((((((....))).((((..(((((((.((.....)).))))))).))))............))))........))))).))). ( -31.20) >DroSec_CAF1 21979 117 + 1 CAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGCUAAAGUCGUUGCUCGUUGGCCAACUUCCAGUCGGCCGACUGCAGAAACAAU---AAUGAAAACAAUAACAAAUACUGG ((((((((((((.(((........))).).)))..(((((((....))).((((..(((((((.((.....)).))))))).)))).......---..))))........))))).))). ( -28.50) >DroYak_CAF1 13131 117 + 1 CAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGCUAAAGUCGUUGCUCGUUGGCCAACUUCAAGUCGGCCGACUGCAGAAACCAU---AAUGAAAACAAUAACAAAUACUGG ((((((((((((.(((........))).).)))(((((.(((....)))..(((..(((((((.((.....)).))))))).)))).))))..---..............))))).))). ( -30.30) >consensus CAGGUUUGGCUCAAAUUAUAAACUAUUAGAAGCGGUUCAGGCUAAAGUCGUUGCUCGUUGGCCAACUUCCAGUCGGCCGACUGCAGAAACAAU___AAUGAAAACAAUAACAAAUACUGG ((((((((((((.(((........))).).)))..(((((((....))).((((..(((((((.((.....)).))))))).))))............))))........))))).))). (-29.62 = -29.40 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:20:27 2006