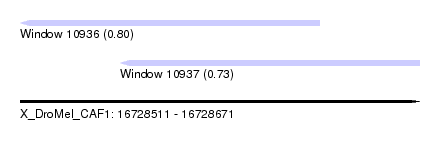
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,728,511 – 16,728,671 |
| Length | 160 |
| Max. P | 0.801804 |

| Location | 16,728,511 – 16,728,631 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.06 |
| Mean single sequence MFE | -33.23 |
| Consensus MFE | -22.54 |
| Energy contribution | -24.35 |
| Covariance contribution | 1.81 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.801804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16728511 120 - 22224390 AUCGCCACUGAGGCAUGAUCGCAUCCAGUGAUCUAAAUGAUCAGCGAUCGGAUUGCAGCAUCGACUUCCCUCACAGCCGCUGUAUCGUUUACACAAUGAUAGCCAUAUAGCCCACAGGCA ...(((.....(((..((((((......(((((.....)))))))))))(((...(......)...)))......)))(((((((.(((...........))).))))))).....))). ( -34.40) >DroSec_CAF1 9308 100 - 1 AUCGCC--------AUGAUCGCGUCCAGUGAUCUAAAUGAUCAGCGAUCGGAUUGCAGCAUCU---------GUAGCCGCUGUAUCGUUUACACAAUGAUAGCCAUA---GCCGUGGGCA ...(((--------..((((((.....))))))(((((((((((((..(((((......))))---------)....))))).))))))))(((.(((.....))).---...)))))). ( -31.70) >DroEre_CAF1 7921 110 - 1 AUCUCCACUGAGGCAUGAUCGCAUCUAAUGACCUAAAUGAUCAGCGAUCGGAUGGCAGCUUCGGCUUCCC-CACAGCCGCUGUAUCGUUUACACAAUGAUACUCAU---------AGGCA ....((..((((((..(((((((((.............)))..))))))((..((.(((....))).)))-)...)))...((((((((.....))))))))))).---------.)).. ( -32.32) >DroYak_CAF1 8374 110 - 1 AUCUCCACUGGGGCAUGAUCGCAUCUAAUGAUCCAAAUGAUCAGCGAUCGGAUUGCAGCUUCGGCUUCCC-CACAGCCGCUGUAUCGUUUACACAAUGAUAGCCGU---------AGGCA ........((((((((((((((......(((((.....))))))))))))...)))(((....)))..))-))..(((((.((((((((.....)))))).)).))---------.))). ( -34.50) >consensus AUCGCCACUGAGGCAUGAUCGCAUCCAAUGAUCUAAAUGAUCAGCGAUCGGAUUGCAGCAUCGGCUUCCC_CACAGCCGCUGUAUCGUUUACACAAUGAUAGCCAU_________AGGCA ...(((..........((((((......(((((.....)))))))))))((...(((((...((((........)))))))))((((((.....))))))..))............))). (-22.54 = -24.35 + 1.81)


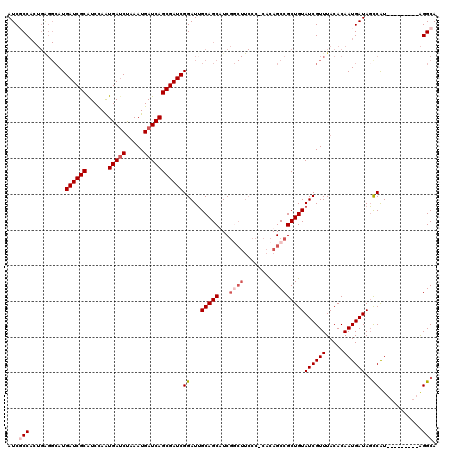
| Location | 16,728,551 – 16,728,671 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.17 |
| Mean single sequence MFE | -36.08 |
| Consensus MFE | -24.32 |
| Energy contribution | -25.32 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.729738 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16728551 120 - 22224390 UGAGGGAUCCCUGCACUUCCUAAUUACUCAAUCAACUGGGAUCGCCACUGAGGCAUGAUCGCAUCCAGUGAUCUAAAUGAUCAGCGAUCGGAUUGCAGCAUCGACUUCCCUCACAGCCGC (((((((((.(((((..(((......((((......))))...(((.....)))..((((((......(((((.....)))))))))))))).)))))....))..)))))))....... ( -39.90) >DroSec_CAF1 9345 103 - 1 UGAGGGAUCCCUGCACUUCCUAAUUACUCAAUCAACUGGGAUCGCC--------AUGAUCGCGUCCAGUGAUCUAAAUGAUCAGCGAUCGGAUUGCAGCAUCU---------GUAGCCGC ...((((((((..........................)))))).))--------..((((((......(((((.....)))))))))))((.((((((...))---------)))))).. ( -31.17) >DroEre_CAF1 7952 119 - 1 UGAGGGAUCCCUGCACUUCCUAAUUACUCAAUCAACUGGGAUCUCCACUGAGGCAUGAUCGCAUCUAAUGACCUAAAUGAUCAGCGAUCGGAUGGCAGCUUCGGCUUCCC-CACAGCCGC ((.((((((((..........................))))))))))....(((..(((((((((.............)))..))))))((..((.(((....))).)))-)...))).. ( -36.29) >DroYak_CAF1 8405 119 - 1 UGAGGGAUCCCUGCACUUCCUAAUUACUCAAUCAACUGGGAUCUCCACUGGGGCAUGAUCGCAUCUAAUGAUCCAAAUGAUCAGCGAUCGGAUUGCAGCUUCGGCUUCCC-CACAGCCGC ((.((((((((..........................)))))))))).((((((((((((((......(((((.....))))))))))))...)))(((....)))..))-))....... ( -36.97) >consensus UGAGGGAUCCCUGCACUUCCUAAUUACUCAAUCAACUGGGAUCGCCACUGAGGCAUGAUCGCAUCCAAUGAUCUAAAUGAUCAGCGAUCGGAUUGCAGCAUCGGCUUCCC_CACAGCCGC ...((((((((..........................)))))).)).....(((.(((((((......(((((.....))))))))))))((........)).............))).. (-24.32 = -25.32 + 1.00)


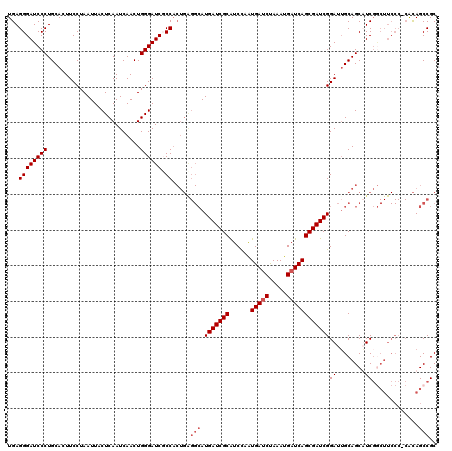
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:20:19 2006