
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,648,706 – 16,648,808 |
| Length | 102 |
| Max. P | 0.993293 |

| Location | 16,648,706 – 16,648,808 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 75.46 |
| Mean single sequence MFE | -25.97 |
| Consensus MFE | -14.21 |
| Energy contribution | -14.90 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.55 |
| SVM decision value | 2.39 |
| SVM RNA-class probability | 0.993293 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16648706 102 + 22224390 ---UUUGUCGUCUGCUGUUGCCUUUGAUUUGCCUCUGGAUCCAGCCACUAUAUAGUUUAUAG--------------UUUAUAGUAUAUAGUAUAUUGUAUACUGUUCACUAUGGCGACU ---...((((((.((((...((...((......)).))...))))......(((((..((((--------------(.(((((((((...))))))))).)))))..))))))))))). ( -29.20) >DroSec_CAF1 7942 88 + 1 ---UUUGUCGUCUGCUGCUGCCUUUGAUUUGUCUCUGGAUCCAGCCACUAUAUAGUUUAUAG---------------------CAU-------AUAGUAUACUGUUCAGUAUGGCGACU ---...((((((.((((...((...((....))...))...)))).(((((((.((.....)---------------------)))-------)))))(((((....))))))))))). ( -24.10) >DroSim_CAF1 8402 95 + 1 ---UUUGUCGUCUGCUGCUGCCUUUGAUUUGCCUCUGGAUCCAGCCACUAUAUAGUUUAUAA---------------------CAUAUAGUAUAUAGUAUACUGUUCAGUAUGGCGACU ---...((((((.((((...((...((......)).))...)))).((((((((.(.(((..---------------------.))).).))))))))(((((....))))))))))). ( -23.90) >DroYak_CAF1 8015 116 + 1 CCUUUUGUCGUCUGC---UGCCUUUGAUUUGCCUUUGGAUCCAGCUACUAUAUACCAUAUAUCAUAUAGCAUCUAGCAUAUAGCAUUUAGAAUAUAGUAUAGAGUAUAGUAUGGCGACU ......((((((.((---((..((..(.......)..))..))))((((((((....(((((.((((....((((((.....))...)))))))).)))))..)))))))).)))))). ( -26.70) >consensus ___UUUGUCGUCUGCUGCUGCCUUUGAUUUGCCUCUGGAUCCAGCCACUAUAUAGUUUAUAG_____________________CAUAUAGUAUAUAGUAUACUGUUCAGUAUGGCGACU ......((((((.((((...((...((......)).))...)))).(((..(((((.(((....................................))).)))))..)))..)))))). (-14.21 = -14.90 + 0.69)


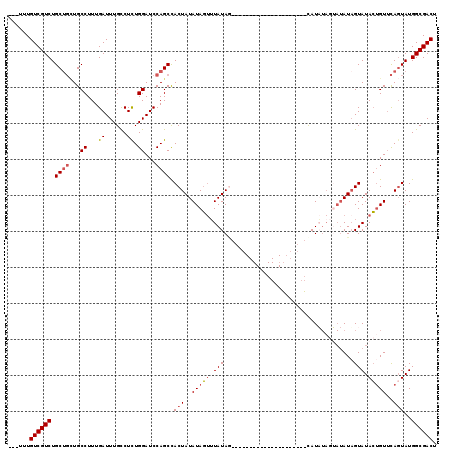
| Location | 16,648,706 – 16,648,808 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 75.46 |
| Mean single sequence MFE | -22.48 |
| Consensus MFE | -11.67 |
| Energy contribution | -12.74 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873365 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16648706 102 - 22224390 AGUCGCCAUAGUGAACAGUAUACAAUAUACUAUAUACUAUAAA--------------CUAUAAACUAUAUAGUGGCUGGAUCCAGAGGCAAAUCAAAGGCAACAGCAGACGACAAA--- .((((((((((((.(.((((((...)))))).).))))))..(--------------(((((.....))))))..(((....))).)))........(....).......)))...--- ( -23.30) >DroSec_CAF1 7942 88 - 1 AGUCGCCAUACUGAACAGUAUACUAU-------AUG---------------------CUAUAAACUAUAUAGUGGCUGGAUCCAGAGACAAAUCAAAGGCAGCAGCAGACGACAAA--- .((((.(...(((..((((.((((((-------(((---------------------........)))))))))))))....))).............((....)).).))))...--- ( -19.80) >DroSim_CAF1 8402 95 - 1 AGUCGCCAUACUGAACAGUAUACUAUAUACUAUAUG---------------------UUAUAAACUAUAUAGUGGCUGGAUCCAGAGGCAAAUCAAAGGCAGCAGCAGACGACAAA--- .(((((((((((....)))))......(((((((((---------------------........))))))))).(((....))).))).........((....))....)))...--- ( -22.50) >DroYak_CAF1 8015 116 - 1 AGUCGCCAUACUAUACUCUAUACUAUAUUCUAAAUGCUAUAUGCUAGAUGCUAUAUGAUAUAUGGUAUAUAGUAGCUGGAUCCAAAGGCAAAUCAAAGGCA---GCAGACGACAAAAGG .((((.(.............((((((((.(((.(((.(((((((.....)).))))).))).))).))))))))((((...((..............))))---)).).))))...... ( -24.34) >consensus AGUCGCCAUACUGAACAGUAUACUAUAUACUAUAUG_____________________CUAUAAACUAUAUAGUGGCUGGAUCCAGAGGCAAAUCAAAGGCAGCAGCAGACGACAAA___ .((((((.............((((((((......................................)))))))).(((....))).))).........((....))....)))...... (-11.67 = -12.74 + 1.06)

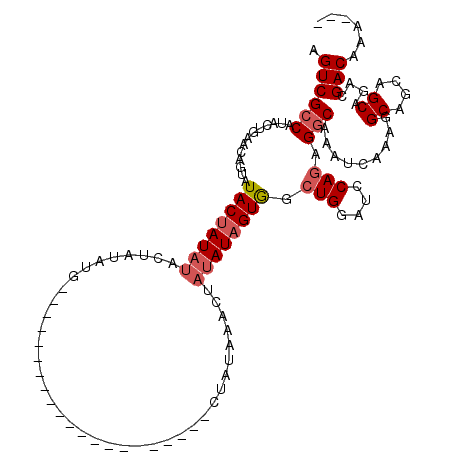

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:19:35 2006