
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,469,589 – 16,469,749 |
| Length | 160 |
| Max. P | 0.979030 |

| Location | 16,469,589 – 16,469,709 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -39.47 |
| Consensus MFE | -36.77 |
| Energy contribution | -37.10 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.748461 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16469589 120 + 22224390 UUUGGCCCAAAACGAUGCAAACGAUGCAGCCGAUGCAACGCGACGCCUGCAACUGACAAACGGCGGCAUCAGCCACUGUGCGUUCGUUUUAGUUCGCUUUUCGUUUUCGUUUUGGUUUUU ......(((((((((...((((((((((.....))))..((((...(((.(((.(((..((((.(((....))).))))..))).))).))).))))...)))))))))))))))..... ( -40.60) >DroSec_CAF1 7611 113 + 1 UUUGGCCCAAAACGAUGCAAACGAUGCAGCCGAUGCAACGCGACGCCUGCAACUGACAAACGGCGGCAUCAGCCACUGUGCGUUCGUUUCAGUUCGCUUUUCGUUUUGGUUUU------- ......(((((((((.((.(((..(((((.((.(((...))).)).)))))((.(((..((((.(((....))).))))..))).))....))).))...)))))))))....------- ( -38.90) >DroSim_CAF1 4898 120 + 1 UUUGGCCCAAAACGAUGCAAACGAUGCAGCCGAUGCAACGCGACGCCUGCAACUGACAAACGGCGGCAUCAGCCACUGUGCGUUCGUUUCAGUUCGCUUUUCGUUUUGGUUUUGGUUUUU ......(((((((((.((.(((..(((((.((.(((...))).)).)))))((.(((..((((.(((....))).))))..))).))....))).))...)))))))))........... ( -38.90) >consensus UUUGGCCCAAAACGAUGCAAACGAUGCAGCCGAUGCAACGCGACGCCUGCAACUGACAAACGGCGGCAUCAGCCACUGUGCGUUCGUUUCAGUUCGCUUUUCGUUUUGGUUUUGGUUUUU ......(((((((((.((.(((..(((((.((.(((...))).)).)))))((.(((..((((.(((....))).))))..))).))....))).))...)))))))))........... (-36.77 = -37.10 + 0.33)

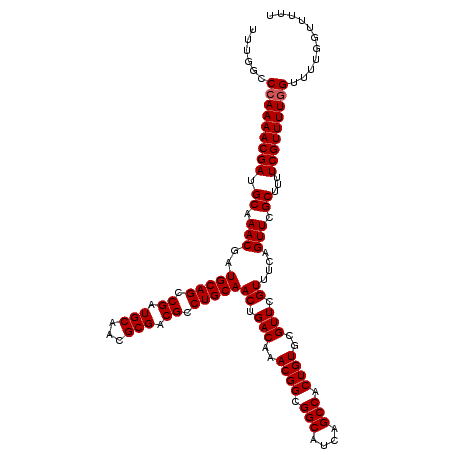
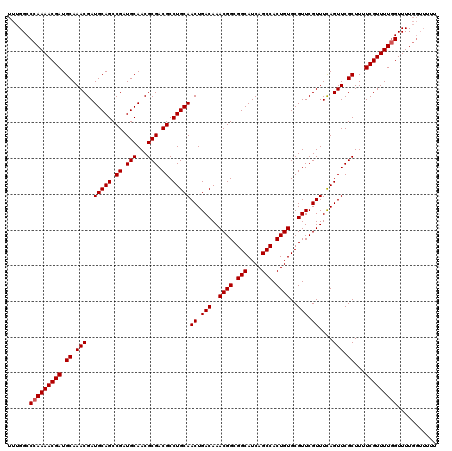
| Location | 16,469,589 – 16,469,709 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -47.63 |
| Consensus MFE | -44.37 |
| Energy contribution | -45.03 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -4.27 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.979030 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16469589 120 - 22224390 AAAAACCAAAACGAAAACGAAAAGCGAACUAAAACGAACGCACAGUGGCUGAUGCCGCCGUUUGUCAGUUGCAGGCGUCGCGUUGCAUCGGCUGCAUCGUUUGCAUCGUUUUGGGCCAAA .....(((((((((((((((...(((..(......)..)))...(..((((((((((((((((((.....)))))))..)))..))))))))..).))))))...)))))))))...... ( -47.10) >DroSec_CAF1 7611 113 - 1 -------AAAACCAAAACGAAAAGCGAACUGAAACGAACGCACAGUGGCUGAUGCCGCCGUUUGUCAGUUGCAGGCGUCGCGUUGCAUCGGCUGCAUCGUUUGCAUCGUUUUGGGCCAAA -------....(((((((((.(((((((((((..((((((....(((((....))))))))))))))))(((((.((..((...))..)).)))))))))))...)))))))))...... ( -47.90) >DroSim_CAF1 4898 120 - 1 AAAAACCAAAACCAAAACGAAAAGCGAACUGAAACGAACGCACAGUGGCUGAUGCCGCCGUUUGUCAGUUGCAGGCGUCGCGUUGCAUCGGCUGCAUCGUUUGCAUCGUUUUGGGCCAAA ...........(((((((((.(((((((((((..((((((....(((((....))))))))))))))))(((((.((..((...))..)).)))))))))))...)))))))))...... ( -47.90) >consensus AAAAACCAAAACCAAAACGAAAAGCGAACUGAAACGAACGCACAGUGGCUGAUGCCGCCGUUUGUCAGUUGCAGGCGUCGCGUUGCAUCGGCUGCAUCGUUUGCAUCGUUUUGGGCCAAA ...........(((((((((.((((((((((..(((((((....(((((....))))))))))))))))(((((.((..((...))..)).)))))))))))...)))))))))...... (-44.37 = -45.03 + 0.67)

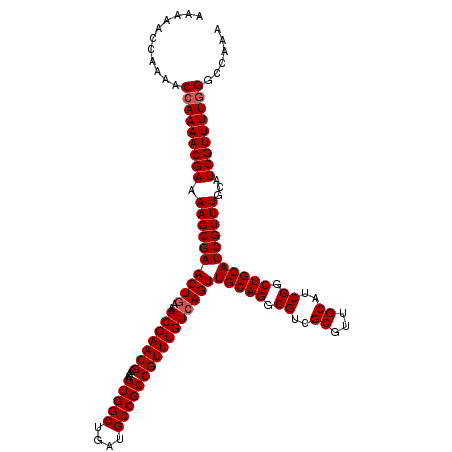
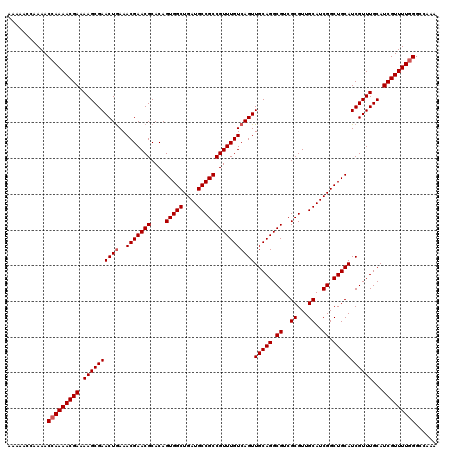
| Location | 16,469,629 – 16,469,749 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.83 |
| Mean single sequence MFE | -24.19 |
| Consensus MFE | -23.40 |
| Energy contribution | -23.73 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.628271 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16469629 120 - 22224390 CCAUUUUCGCAUUAAAAUUUGUAACCAAACCAAAAAGAAAAAAAACCAAAACGAAAACGAAAAGCGAACUAAAACGAACGCACAGUGGCUGAUGCCGCCGUUUGUCAGUUGCAGGCGUCG .....(((((.......((((....))))......................((....))....)))))......(((.(((...(..(((((((........)))))))..)..)))))) ( -22.70) >DroSec_CAF1 7651 104 - 1 CCAUUUUCGCAUUAAAAUUUGUAACCAAACC----------------AAAACCAAAACGAAAAGCGAACUGAAACGAACGCACAGUGGCUGAUGCCGCCGUUUGUCAGUUGCAGGCGUCG ((.((((((........((((.........)----------------))).......))))))((.((((((..((((((....(((((....))))))))))))))))))).))..... ( -25.06) >DroSim_CAF1 4938 114 - 1 CCAUUUUCGCAUUAAAAUUUGUAACCAAACCAAA------AAAAACCAAAACCAAAACGAAAAGCGAACUGAAACGAACGCACAGUGGCUGAUGCCGCCGUUUGUCAGUUGCAGGCGUCG .......(((.......((((....)))).....------.......................((.((((((..((((((....(((((....)))))))))))))))))))..)))... ( -24.80) >consensus CCAUUUUCGCAUUAAAAUUUGUAACCAAACCAAA______AAAAACCAAAACCAAAACGAAAAGCGAACUGAAACGAACGCACAGUGGCUGAUGCCGCCGUUUGUCAGUUGCAGGCGUCG .......(((.......((((....))))..................................((.(((((..(((((((....(((((....)))))))))))))))))))..)))... (-23.40 = -23.73 + 0.33)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:18:00 2006