
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,365,023 – 16,365,156 |
| Length | 133 |
| Max. P | 0.996593 |

| Location | 16,365,023 – 16,365,123 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 84.74 |
| Mean single sequence MFE | -26.10 |
| Consensus MFE | -20.81 |
| Energy contribution | -21.38 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.34 |
| Structure conservation index | 0.80 |
| SVM decision value | 2.72 |
| SVM RNA-class probability | 0.996593 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16365023 100 - 22224390 AAAGCGACGAUGUGAGCGAGAUUGGCGUAAAAGGGACAAGGGUGUCUCAAAUUGCCUUGGCUUAGCUCACAAAAAAUUGAACCAAAAA--AAAAAAAAAAUG ..........((((((((((...((((.....((((((....))))))....))))....))).))))))).................--............ ( -25.20) >DroSec_CAF1 37830 99 - 1 AAAGCGACUAUGUGAGCGAGAUUGGCGUAAAAGGGACAAGGGUGUCUCAAAUUGCCUUGGUUUAGCUCACAAAAAAUUGAAUCUAAAA--AA-AAAAAAAUU ..........(((((((.((((..(.((((..((((((....))))))...)))).)..)))).))))))).................--..-......... ( -25.30) >DroSim_CAF1 26186 99 - 1 AAAGCGACUGUGUGAGCGAGAUUGGCGAAAAAGGGACAAGGGUGUCUCAAAUUGCCUUGGCUUAGCUCACACAAAAUUGAAUCUAAAA--AA-AAAAAAAUG ........((((((((((((...(((((....((((((....))))))...)))))....))).)))))))))...............--..-......... ( -30.10) >DroEre_CAF1 23501 90 - 1 -AAGCGGCGAUGAGAGCGCGAUUGGCGUAAGAGGGACAAGGGUUUCUCAAAUUGCCUUGGCUUAGCUCACAAAAAACU----------GAAC-AUCAAAAUG -.......((((.((((((.(..((((...(((..((....))..)))....)))).).))...)))).((......)----------)..)-)))...... ( -23.80) >consensus AAAGCGACGAUGUGAGCGAGAUUGGCGUAAAAGGGACAAGGGUGUCUCAAAUUGCCUUGGCUUAGCUCACAAAAAAUUGAAUCUAAAA__AA_AAAAAAAUG ..........((((((((((...((((.....((((((....))))))....))))....))).)))))))............................... (-20.81 = -21.38 + 0.56)


| Location | 16,365,048 – 16,365,156 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.46 |
| Mean single sequence MFE | -30.46 |
| Consensus MFE | -20.34 |
| Energy contribution | -21.18 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.605364 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16365048 108 - 22224390 UCCGAA------GAGAGCA-GGAUAUGCAAGAGAGCGCAGA-----AAGCGACGAUGUGAGCGAGAUUGGCGUAAAAGGGACAAGGGUGUCUCAAAUUGCCUUGGCUUAGCUCACAAAAA ..((..------....(((-.....))).......(((...-----..))).)).((((((((((...((((.....((((((....))))))....))))....))).))))))).... ( -31.90) >DroSec_CAF1 37854 108 - 1 UCCAAA------GAGAGCA-GGAUAUUCAAGAGAGCGCUGA-----AAGCGACUAUGUGAGCGAGAUUGGCGUAAAAGGGACAAGGGUGUCUCAAAUUGCCUUGGUUUAGCUCACAAAAA (((...------.......-)))..........(((((...-----..))).)).(((((((.((((..(.((((..((((((....))))))...)))).)..)))).))))))).... ( -30.30) >DroSim_CAF1 26210 108 - 1 UCCAAA------GAGAGCA-GGAUAUUCAAGAGAGCGCUGA-----AAGCGACUGUGUGAGCGAGAUUGGCGAAAAAGGGACAAGGGUGUCUCAAAUUGCCUUGGCUUAGCUCACACAAA (((...------.......-)))..........(((((...-----..))).))(((((((((((...(((((....((((((....))))))...)))))....))).))))))))... ( -34.30) >DroEre_CAF1 23517 105 - 1 UCCACG------GAGAGCA-GCGAAUUGCAGAGAGCGA--------AAGCGGCGAUGAGAGCGCGAUUGGCGUAAGAGGGACAAGGGUUUCUCAAAUUGCCUUGGCUUAGCUCACAAAAA ......------..(((((-((.((.........((..--------..))((((((....((((.....))))..(((..((....))..)))..)))))))).)))..))))....... ( -35.00) >DroYak_CAF1 22544 120 - 1 UCCAAAAAGAGAGAGAGCAAAUAAAUUCAAGAGAGCGACGAAAUUCAAGCGACGAUGAGAGCGAGAUUGGCGUAAAAGGGACAAGGGUGUCUCAAAUUGCCUUGGCUUAGCGCACAAAAA ................((.......((((.....((............)).....)))).(((((...((((.....((((((....))))))....))))....))).))))....... ( -20.80) >consensus UCCAAA______GAGAGCA_GGAUAUUCAAGAGAGCGCUGA_____AAGCGACGAUGUGAGCGAGAUUGGCGUAAAAGGGACAAGGGUGUCUCAAAUUGCCUUGGCUUAGCUCACAAAAA ..................................((............)).....((((((((((...((((.....((((((....))))))....))))....))).))))))).... (-20.34 = -21.18 + 0.84)

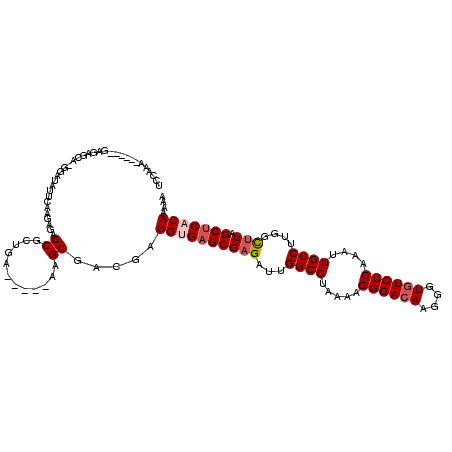

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:16:14 2006