
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,308,399 – 16,308,519 |
| Length | 120 |
| Max. P | 0.896625 |

| Location | 16,308,399 – 16,308,519 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.39 |
| Mean single sequence MFE | -30.13 |
| Consensus MFE | -30.13 |
| Energy contribution | -30.13 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.85 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873258 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
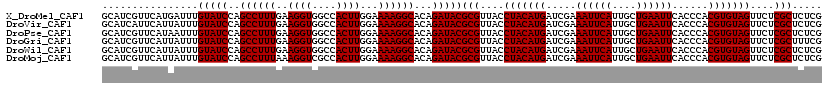
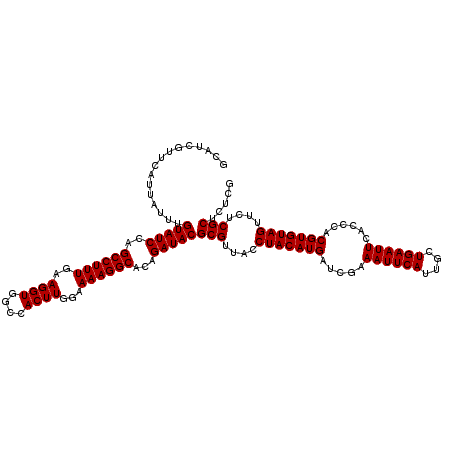
>X_DroMel_CAF1 16308399 120 + 22224390 GCAUCGUUCAUGAUUUGUAUCCAGCCUUUGAAGGUGGCCACUUGGAAAAGGCACAGAUACGCGUUACCUACAUGAUCGAAAUUCAUUGCUGAAUUCACCCACGUGUAGUUCUCGCUCUCG ................(((((..((((((..((((....))))...))))))...)))))(((....(((((((.....((((((....))))))......)))))))....)))..... ( -30.50) >DroVir_CAF1 11630 120 + 1 GCAUCAUUCAUUAUUUGUAUCCAGCCUUUGAAGGUGGCCACUUGGAAAAGGCACAGAUACGCGUUACCUACAUGAUCGAAAUUCAUUGCUGAAUUCACCCACGUGUAGUUCUCGCUCUCG ................(((((..((((((..((((....))))...))))))...)))))(((....(((((((.....((((((....))))))......)))))))....)))..... ( -30.50) >DroPse_CAF1 12501 120 + 1 GCAUCGUUCAUAAUUUGUAUCCAGCCUUUGAAGGUGGCCACUUGGAAAAGGCACAGAUACGCGUUACCUACAUGAUCGAAAUUCAUUGCUGAAUUCACCCACGUGUAGUUCUCGCUCUCG ................(((((..((((((..((((....))))...))))))...)))))(((....(((((((.....((((((....))))))......)))))))....)))..... ( -30.50) >DroGri_CAF1 12160 120 + 1 GCAUCGUUCAUUAUUUGUAUCCAGCCUUUGAAGGUGGCCACUUGGAAAAGGCACAGAUACGCGUUACCUACAUGAUCGAAAUUCAUUGCUGAAUUCACCCACGUGUAGUUCUCGCUUUCG ................(((((..((((((..((((....))))...))))))...)))))(((....(((((((.....((((((....))))))......)))))))....)))..... ( -30.50) >DroWil_CAF1 13515 120 + 1 GCAUCGUUCAUUAUUUGUAUCCAGCCUUUGAAGGUGGCCACUUGGAAAAGGCACAGAUACGCGUUACCUACAUGAUCGAAAUUCAUUGCUGAAUUCACCCACGUGUAGUUCUCGCUCUCG ................(((((..((((((..((((....))))...))))))...)))))(((....(((((((.....((((((....))))))......)))))))....)))..... ( -30.50) >DroMoj_CAF1 12270 120 + 1 GCAUCGUUCAUUAUUUGUAUCCAGCCUUUAAAGGUCGCCACUUGGAAAAGGCACAGAUACGCGUUACCUACAUGAUCGAAAUUCAUUGCUGAAUUCACCCACGUGUAGUUCUCGCUCUCG ................(((((..((((((..((((....))))...))))))...)))))(((....(((((((.....((((((....))))))......)))))))....)))..... ( -28.30) >consensus GCAUCGUUCAUUAUUUGUAUCCAGCCUUUGAAGGUGGCCACUUGGAAAAGGCACAGAUACGCGUUACCUACAUGAUCGAAAUUCAUUGCUGAAUUCACCCACGUGUAGUUCUCGCUCUCG ................(((((..((((((..((((....))))...))))))...)))))(((....(((((((.....((((((....))))))......)))))))....)))..... (-30.13 = -30.13 + 0.00)

| Location | 16,308,399 – 16,308,519 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.39 |
| Mean single sequence MFE | -32.78 |
| Consensus MFE | -32.67 |
| Energy contribution | -32.53 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.03 |
| Mean z-score | -0.90 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.896625 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
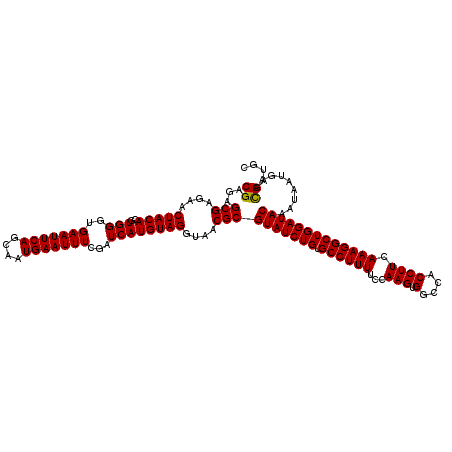
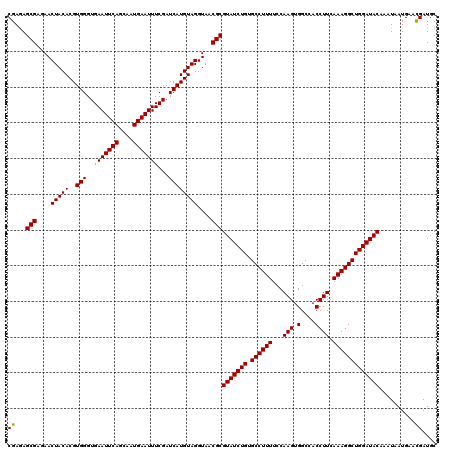
>X_DroMel_CAF1 16308399 120 - 22224390 CGAGAGCGAGAACUACACGUGGGUGAAUUCAGCAAUGAAUUUCGAUCAUGUAGGUAACGCGUAUCUGUGCCUUUUCCAAGUGGCCACCUUCAAAGGCUGGAUACAAAUCAUGAACGAUGC ((...(((....(((((..(((..(((((((....)))))))...))))))))....)))(((((((.((((((...(((.(....)))).)))))))))))))..........)).... ( -32.50) >DroVir_CAF1 11630 120 - 1 CGAGAGCGAGAACUACACGUGGGUGAAUUCAGCAAUGAAUUUCGAUCAUGUAGGUAACGCGUAUCUGUGCCUUUUCCAAGUGGCCACCUUCAAAGGCUGGAUACAAAUAAUGAAUGAUGC .....(((....(((((..(((..(((((((....)))))))...))))))))....)))(((((((.((((((...(((.(....)))).)))))))))))))................ ( -32.00) >DroPse_CAF1 12501 120 - 1 CGAGAGCGAGAACUACACGUGGGUGAAUUCAGCAAUGAAUUUCGAUCAUGUAGGUAACGCGUAUCUGUGCCUUUUCCAAGUGGCCACCUUCAAAGGCUGGAUACAAAUUAUGAACGAUGC ((...(((....(((((..(((..(((((((....)))))))...))))))))....)))(((((((.((((((...(((.(....)))).)))))))))))))..........)).... ( -32.50) >DroGri_CAF1 12160 120 - 1 CGAAAGCGAGAACUACACGUGGGUGAAUUCAGCAAUGAAUUUCGAUCAUGUAGGUAACGCGUAUCUGUGCCUUUUCCAAGUGGCCACCUUCAAAGGCUGGAUACAAAUAAUGAACGAUGC ((...(((....(((((..(((..(((((((....)))))))...))))))))....)))(((((((.((((((...(((.(....)))).)))))))))))))..........)).... ( -32.50) >DroWil_CAF1 13515 120 - 1 CGAGAGCGAGAACUACACGUGGGUGAAUUCAGCAAUGAAUUUCGAUCAUGUAGGUAACGCGUAUCUGUGCCUUUUCCAAGUGGCCACCUUCAAAGGCUGGAUACAAAUAAUGAACGAUGC ((...(((....(((((..(((..(((((((....)))))))...))))))))....)))(((((((.((((((...(((.(....)))).)))))))))))))..........)).... ( -32.50) >DroMoj_CAF1 12270 120 - 1 CGAGAGCGAGAACUACACGUGGGUGAAUUCAGCAAUGAAUUUCGAUCAUGUAGGUAACGCGUAUCUGUGCCUUUUCCAAGUGGCGACCUUUAAAGGCUGGAUACAAAUAAUGAACGAUGC ((...(((....(((((..(((..(((((((....)))))))...))))))))....)))(((((((.((((((...(((.(....)))).)))))))))))))..........)).... ( -34.70) >consensus CGAGAGCGAGAACUACACGUGGGUGAAUUCAGCAAUGAAUUUCGAUCAUGUAGGUAACGCGUAUCUGUGCCUUUUCCAAGUGGCCACCUUCAAAGGCUGGAUACAAAUAAUGAACGAUGC ((...(((....(((((..(((..(((((((....)))))))...))))))))....)))(((((((.((((((...(((.(....)))).)))))))))))))..........)).... (-32.67 = -32.53 + -0.14)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:15:30 2006