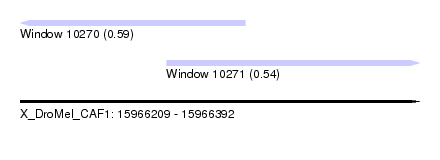
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,966,209 – 15,966,392 |
| Length | 183 |
| Max. P | 0.585966 |

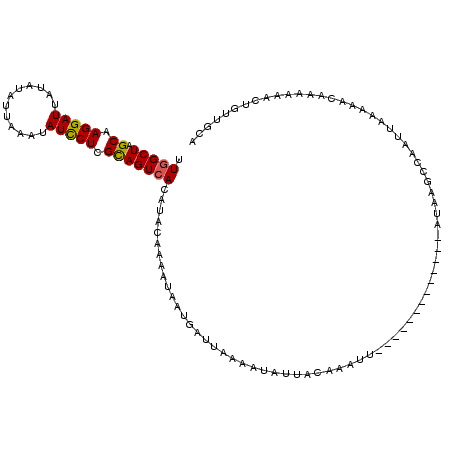
| Location | 15,966,209 – 15,966,312 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 76.56 |
| Mean single sequence MFE | -15.48 |
| Consensus MFE | -10.00 |
| Energy contribution | -9.88 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.585966 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15966209 103 - 22224390 UUUGCUAGCAAGGAUUAUAAAUUAAAUAUUCUCGCAGUCACAUACAAGAGAAUUAUUUAAAUAUUACAAACU-------------AUAAGCCAAUUAAAAACAAAAAACUGUUGCA .......((..((.(((((..((((((((((((...((.....))..))))).)))))))...........)-------------)))).)).......((((......)))))). ( -11.62) >DroSec_CAF1 34391 103 - 1 UUGGCUAGCAAGGAUUAUAUAUUAAAUAUCCUCGCAGUCACAUACAAAAUAAUGAUUAAAAUAUUACAAAUU-------------AUAAGCCAAUUUAAAACAAAAAACUGUUGCA ((((((.((.(((((............))))).))(((((............)))))...............-------------...)))))).....((((......))))... ( -16.20) >DroSim_CAF1 25426 103 - 1 UUGGCUAGCAAGGAUUAUAUAUUAAAUAUCCUCGCAGUCACAUAUAAGAAAAGGAUUAAAAUAUUACAAAUU-------------AUAAGCCAAUUAAAAACAGAAAACUGUUGCA ((((((.((.(((((............))))).)).......(((((......(((......))).....))-------------))))))))).....(((((....)))))... ( -19.00) >DroYak_CAF1 17464 108 - 1 UUGGCUAGCAAGGAUUACAUUUUAAAUAUCCUCGUAGUCACAUACAAAAUUAAGUUUAAC-----UCAAAUUGAUCAAGAACAAUGUAAAAAUAUUUGACACAGAUA---UUUGCA .((((((.(.(((((............))))).))))))).............((((...-----((.....))....))))..(((((..(((((((...))))))---)))))) ( -15.10) >consensus UUGGCUAGCAAGGAUUAUAUAUUAAAUAUCCUCGCAGUCACAUACAAAAUAAUGAUUAAAAUAUUACAAAUU_____________AUAAGCCAAUUAAAAACAAAAAACUGUUGCA .(((((.((.(((((............))))).)))))))............................................................................ (-10.00 = -9.88 + -0.12)



| Location | 15,966,276 – 15,966,392 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 90.95 |
| Mean single sequence MFE | -16.78 |
| Consensus MFE | -13.72 |
| Energy contribution | -14.10 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.536350 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
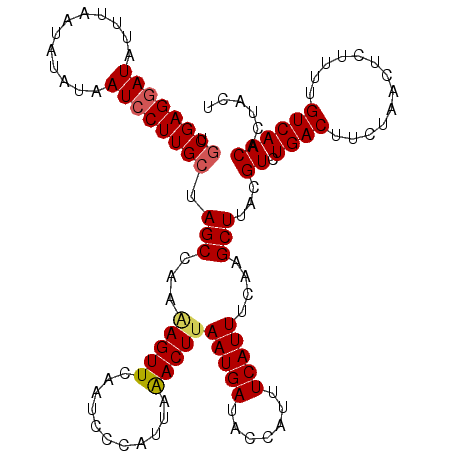
>X_DroMel_CAF1 15966276 116 + 22224390 UGCGAGAAUAUUUAAUUUAUAAUCCUUGCUAGCAAAAAGUUCAAUCGCAUUAAACUUAAUGAUACCAUUUCAUUUCAAGCUUACGUCUGACUUCUAACUCUUUUGUCAACACUACU .(((((..................))))).(((...(((((.(((...))).)))))(((((.......)))))....)))...((.((((.............))))))...... ( -11.79) >DroSec_CAF1 34458 115 + 1 UGCGAGGAUAUUUAAUAUAUAAUCCUUGCUAGCCAA-AGUUCAAUCCCAUUAGACUUAAUGAUACCAUUUCAUUUCAAGCUUACGUCUGACUUCUAACUCUUUUGUCAACACUACU .((((((((............)))))))).(((..(-(((..(((...)))..))))(((((.......)))))....)))...((.((((.............))))))...... ( -17.92) >DroSim_CAF1 25493 116 + 1 UGCGAGGAUAUUUAAUAUAUAAUCCUUGCUAGCCAAAAGUUCAAUCCCAUUAGACUUAAUGAUACCAUUUCAUUUUAAGCUUACGUCUGACUUCUAACUCUUUUGUCAACACUACU .((((((((............))))))))..(.((((((..........((((((..(((((.......)))))..........)))))).........)))))).)......... ( -19.71) >DroYak_CAF1 17536 113 + 1 UACGAGGAUAUUUAAAAUGUAAUCCUUGCUAGCCAAGAGUUCAAUCUCAACCAACUUAAUGAUACCAUUUCAUUAUCAGCUUACGUCUGACUUCCAACUCUUUUGUCAACAC---U ..(((((((............)))))))......(((((((...............((((((.......))))))(((((....).)))).....)))))))..........---. ( -17.70) >consensus UGCGAGGAUAUUUAAUAUAUAAUCCUUGCUAGCCAAAAGUUCAAUCCCAUUAAACUUAAUGAUACCAUUUCAUUUCAAGCUUACGUCUGACUUCUAACUCUUUUGUCAACACUACU .((((((((............)))))))).(((...(((((...........)))))(((((.......)))))....)))...((.((((.............))))))...... (-13.72 = -14.10 + 0.37)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:10:03 2006