
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,834,968 – 15,835,128 |
| Length | 160 |
| Max. P | 0.848781 |

| Location | 15,834,968 – 15,835,088 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.22 |
| Mean single sequence MFE | -44.82 |
| Consensus MFE | -31.43 |
| Energy contribution | -30.30 |
| Covariance contribution | -1.13 |
| Combinations/Pair | 1.48 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.628975 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15834968 120 - 22224390 CACUUCACCGGACGCCUGUCCACGCAACGUGCGUUUCAUUUGCAGCUGUCUUCAGAAGGCUGUGAUGGCCAAAUGGCCGACAGAGCGGCUCGUUCGCACACGCGUCGUCUCUGGCUUUAU .........((((....))))((((.....)))).....(..(((((.((....)).)))))..)(((((....))))).(((((((((.(((......))).)))).)))))....... ( -44.20) >DroVir_CAF1 2202 120 - 1 CACAUCGCCGGACGCCUGUCCGCGCAACGUACGCUACAUAUGCAAUUGCCUGCAGAAGGCUGUCGUGGCCAAAUGGCCAACGGAACGCUUGGUGCGGACACGUGUCGUCUCUGGUUUUAU ......(((((((((.(((((((((..(((..((.......))....((((.....))))..((((((((....)))).)))).)))....))))))))).))))).....))))..... ( -46.30) >DroGri_CAF1 2487 120 - 1 CACAUCGCCGGAUGCCUGUCCGCGCAAUGUGCGCUACAUUUGCAACUGCCUGCAGAAGGCGGUCGUUGCCAAAUGGCAAACGGAGCGUUUGGUGCGUACGCGUGUCGUCUCCGGUUUUAU ......((((((.((.....(((((...((((((..((..(((.(((((((.....)))))))(((((((....))).))))..)))..))..)))))))))))..)).))))))..... ( -50.30) >DroWil_CAF1 2629 120 - 1 CACAUCCCCGGACGCCUGUCCGCGUAAUGUACGCUAUAUUUGCAAUUGCCUUCAGAAGGCGGUGGUCGCCAAAUGGCCAACGGAGCGUCUGGUGCGUACUAGGGUCGUUUCUGGUUUCAU .......(((((((((((((((((((...))))).............((((.....))))))((((((.....)))))))))).))))))))((...(((((((....)))))))..)). ( -40.60) >DroMoj_CAF1 585 120 - 1 CACGUCGCCGGACGCUUGUCCGCGCAACGUGCGCUACAUCUGCAACUGCUUGCAGAAGGCGGUCGUGGCCAAGUGGCCGACGGAGCGUUUGGUGCGCACGCGCGUCGUAUCCGGUUUUAU ......((((((..(.....(((((...((((((.((.(((((((....)))))))(((((.((((((((....)))).))))..))))).)))))))))))))..)..))))))..... ( -54.60) >DroAna_CAF1 5034 120 - 1 UACUUCACCAGAUGCUUGCCCGCGAAAUGUGCGCUAUAUUUGUAACUGCCUUCAGAAAGCUGUAAUGGCUAAAUGGCCAACAGAACGUUUGGUCCGUACCCGUGUUGUAUCUGGAUUUAU .......((((((((.....((((....(((((...((..(((..(((....)))....((((..(((((....))))))))).)))..))...))))).))))..))))))))...... ( -32.90) >consensus CACAUCGCCGGACGCCUGUCCGCGCAACGUGCGCUACAUUUGCAACUGCCUGCAGAAGGCGGUCGUGGCCAAAUGGCCAACGGAGCGUUUGGUGCGUACACGUGUCGUCUCUGGUUUUAU .......(((((.((......((((...((((((..((..(((.((.((((.....)))).))..(((((....))))).....)))..))..))))))..)))).)).)))))...... (-31.43 = -30.30 + -1.13)


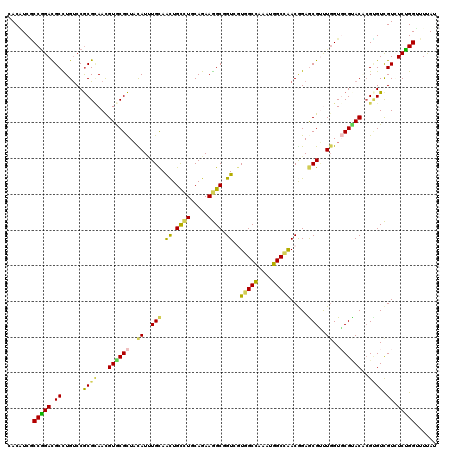
| Location | 15,835,008 – 15,835,128 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.61 |
| Mean single sequence MFE | -44.90 |
| Consensus MFE | -33.41 |
| Energy contribution | -36.75 |
| Covariance contribution | 3.34 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.848781 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
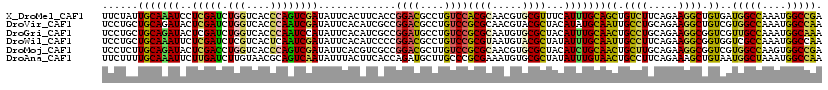
>X_DroMel_CAF1 15835008 120 + 22224390 UCGGCCAUUUGGCCAUCACAGCCUUCUGAAGACAGCUGCAAAUGAAACGCACGUUGCGUGGACAGGCGUCCGGUGAAGUGAAUAUCGACUGGGUGACCAGAUCGAGGAUUUGCAAUAGAA ..((((....)))).((((.(((((((...(.(((((((.........))).)))))..))).)))).....)))).(..(((.((((((((....)))).))))..)))..)....... ( -44.70) >DroVir_CAF1 2242 120 + 1 UUGGCCAUUUGGCCACGACAGCCUUCUGCAGGCAAUUGCAUAUGUAGCGUACGUUGCGCGGACAGGCGUCCGGCGAUGUGAAUAUCGAUUGGGUGACCAGAUCGAGUAUCUGCAGCAGGA .(((((....)))))......(((.((((((((....)).......((.((((((((.(((((....)))))))))))))....((((((((....))).)))))))..)))))).))). ( -53.80) >DroGri_CAF1 2527 120 + 1 UUUGCCAUUUGGCAACGACCGCCUUCUGCAGGCAGUUGCAAAUGUAGCGCACAUUGCGCGGACAGGCAUCCGGCGAUGUGAAUAUGGAUUGGGUGACCAGAUCGAGUAUCUGCAGCAGGA .(((((....)))))......(((.(((((((..(((((....))))).((((((((.((((......))))))))))))....(((.((((....)))).)))....))))))).))). ( -49.80) >DroWil_CAF1 2669 120 + 1 UUGGCCAUUUGGCGACCACCGCCUUCUGAAGGCAAUUGCAAAUAUAGCGUACAUUACGCGGACAGGCGUCCGGGGAUGUGAAUAUCGAUUGAGUGACGAGAUCGAGAAUUUGCAGCAGGA ...(((....))).....((((((.....))))..((((((((......((((((.(.(((((....))))).)))))))....((((((.........))))))..))))))))..)). ( -38.00) >DroMoj_CAF1 625 120 + 1 UCGGCCACUUGGCCACGACCGCCUUCUGCAAGCAGUUGCAGAUGUAGCGCACGUUGCGCGGACAAGCGUCCGGCGACGUGAAUAUCGACUGGGUGACCAGGUCGAGUAUCUGCAAGAGGA ..((((....)))).......(((((((....)))(((((((((.....((((((((.(((((....)))))))))))))....((((((((....))).))))).))))))))))))). ( -60.00) >DroAna_CAF1 5074 120 + 1 UUGGCCAUUUAGCCAUUACAGCUUUCUGAAGGCAGUUACAAAUAUAGCGCACAUUUCGCGGGCAAGCAUCUGGUGAAGUAAAUAUUGACUGCGUUACAAGAUCAAGAAUUUGCAAAAGAA .((((......)))).....((.((((.((.(((((((........(....)(((((((.((.......)).)))))))......))))))).)).........))))...))....... ( -23.10) >consensus UUGGCCAUUUGGCCACGACAGCCUUCUGAAGGCAGUUGCAAAUGUAGCGCACAUUGCGCGGACAGGCGUCCGGCGAUGUGAAUAUCGACUGGGUGACCAGAUCGAGUAUCUGCAACAGGA ..((((....))))......((((.....))))..((((((((......((((((((.(((((....)))))))))))))....((((((((....)))).))))..))))))))..... (-33.41 = -36.75 + 3.34)



| Location | 15,835,008 – 15,835,128 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.61 |
| Mean single sequence MFE | -36.98 |
| Consensus MFE | -29.07 |
| Energy contribution | -27.92 |
| Covariance contribution | -1.16 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.619392 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
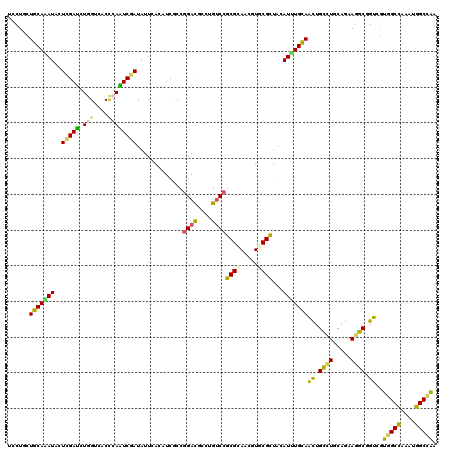
>X_DroMel_CAF1 15835008 120 - 22224390 UUCUAUUGCAAAUCCUCGAUCUGGUCACCCAGUCGAUAUUCACUUCACCGGACGCCUGUCCACGCAACGUGCGUUUCAUUUGCAGCUGUCUUCAGAAGGCUGUGAUGGCCAAAUGGCCGA .....((((......((((.((((....)))))))).............((((....))))..))))............(..(((((.((....)).)))))..)(((((....))))). ( -37.10) >DroVir_CAF1 2242 120 - 1 UCCUGCUGCAGAUACUCGAUCUGGUCACCCAAUCGAUAUUCACAUCGCCGGACGCCUGUCCGCGCAACGUACGCUACAUAUGCAAUUGCCUGCAGAAGGCUGUCGUGGCCAAAUGGCCAA .(((.((((((.((((((((.(((....))))))))..........(((((((....))))).))...))).((.......))......)))))).)))......(((((....))))). ( -41.50) >DroGri_CAF1 2527 120 - 1 UCCUGCUGCAGAUACUCGAUCUGGUCACCCAAUCCAUAUUCACAUCGCCGGAUGCCUGUCCGCGCAAUGUGCGCUACAUUUGCAACUGCCUGCAGAAGGCGGUCGUUGCCAAAUGGCAAA ......(((((((....(((.(((....))))))......(((((.(((((((....))))).)).)))))......)))))))(((((((.....)))))))..(((((....))))). ( -40.40) >DroWil_CAF1 2669 120 - 1 UCCUGCUGCAAAUUCUCGAUCUCGUCACUCAAUCGAUAUUCACAUCCCCGGACGCCUGUCCGCGUAAUGUACGCUAUAUUUGCAAUUGCCUUCAGAAGGCGGUGGUCGCCAAAUGGCCAA ......(((((((..(((((...........))))).....((((..((((((....))))).)..)))).......)))))))(((((((.....)))))))(((((.....))))).. ( -31.20) >DroMoj_CAF1 625 120 - 1 UCCUCUUGCAGAUACUCGACCUGGUCACCCAGUCGAUAUUCACGUCGCCGGACGCUUGUCCGCGCAACGUGCGCUACAUCUGCAACUGCUUGCAGAAGGCGGUCGUGGCCAAGUGGCCGA ......(((((((..(((((.(((....))))))))....(((((.(((((((....))))).)).)))))......)))))))(((((((.....)))))))..(((((....))))). ( -50.90) >DroAna_CAF1 5074 120 - 1 UUCUUUUGCAAAUUCUUGAUCUUGUAACGCAGUCAAUAUUUACUUCACCAGAUGCUUGCCCGCGAAAUGUGCGCUAUAUUUGUAACUGCCUUCAGAAAGCUGUAAUGGCUAAAUGGCCAA .....((((((..(....)..)))))).(((((...............((((((...((.(((.....))).))..))))))...(((....)))...)))))..(((((....))))). ( -20.80) >consensus UCCUGCUGCAAAUACUCGAUCUGGUCACCCAAUCGAUAUUCACAUCGCCGGACGCCUGUCCGCGCAACGUGCGCUACAUUUGCAACUGCCUGCAGAAGGCGGUCGUGGCCAAAUGGCCAA ......(((((((..(((((.(((....)))))))).............((((....))))((((.....))))...)))))))((.((((.....)))).))..(((((....))))). (-29.07 = -27.92 + -1.16)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:07:57 2006