
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,826,756 – 15,826,939 |
| Length | 183 |
| Max. P | 0.998179 |

| Location | 15,826,756 – 15,826,874 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.25 |
| Mean single sequence MFE | -26.28 |
| Consensus MFE | -21.89 |
| Energy contribution | -21.57 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.743009 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15826756 118 + 22224390 UGCAUUCAAAUUAAUGCUUAACCAUAACCUUGUGUGUAUGUAAAUAUAAACUUGGCAGGUUGAUUUUUUUU--UACCAUUCUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAU .(((((......)))))((((((....((..((.(((((....))))).))..))..))))))........--..(((......)))......((((((((((....))))))))))... ( -27.10) >DroSec_CAF1 5624 118 + 1 UGCAUUCAAAUUAGUGCUGAGCCAUAACCUUGUGUGUAUGUAAAUAUAAACCAGGCAGGUUGCUUAUUUUU--UACCAUUCUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAU ..........((((((((.(((.....(((.((.(((((....))))).)).)))..(((...........--.))).....)))))))))))((((((((((....))))))))))... ( -33.30) >DroSim_CAF1 5413 118 + 1 UGCAUGUAAAUUAUUGCUGAACCAUAACCUUGUGUGUAUGUAAAUAUAAACUAGGCAGGUUGCUUAUUUUU--UACCAUUCUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAU .(((.(((((..((.((..((((....(((.((.(((((....))))).)).)))..))))))..))..))--))).....))).........((((((((((....))))))))))... ( -26.40) >DroEre_CAF1 6134 117 + 1 UGUAUUAAAAUAAAUGUUGAACCAUAACCGUGUGUGUAUGUAAAUAUAAACUAGGCAGGUUGCUUUUUUU---UACCAUUCUGCUGUCACUAAACAUACGUACAAGUGUAUGUAUGUGAU .................(((.........((...(((((....))))).))..(((((..((........---...))..))))).)))....((((((((((....))))))))))... ( -21.20) >DroYak_CAF1 5427 120 + 1 UGUAUUGAAAUAAAUAUUGAACCAUAACCGUGUGUAUAUGUAAAUAUAAACUAGGCAGGUUAAUUUUUUUUUCUACCAUUCUGCUGGCACCAAACAUACGUACAAGUGUAUGUAUGUGAU ......((((.(((.(((((.((....((..((.(((((....))))).))..))..))))))).))).))))..(((......)))......((((((((((....))))))))))... ( -23.40) >consensus UGCAUUCAAAUUAAUGCUGAACCAUAACCUUGUGUGUAUGUAAAUAUAAACUAGGCAGGUUGCUUUUUUUU__UACCAUUCUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAU .(((((......)))))..............((((((((....))))......((((((....................)))))).))))...((((((((((....))))))))))... (-21.89 = -21.57 + -0.32)



| Location | 15,826,796 – 15,826,906 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.00 |
| Mean single sequence MFE | -27.06 |
| Consensus MFE | -24.67 |
| Energy contribution | -24.27 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.03 |
| SVM RNA-class probability | 0.998179 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15826796 110 + 22224390 UAAAUAUAAACUUGGCAGGUUGAUUUUUUUU--UACCAUUCUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCAUUUAUCUGU----A----CAUGUGUAA .............(((((..((.........--...))..))))).((((...((((((((((....))))))))))(((((.....)))))..........----.----...)))).. ( -28.90) >DroSec_CAF1 5664 114 + 1 UAAAUAUAAACCAGGCAGGUUGCUUAUUUUU--UACCAUUCUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCAUUUAUUUGUAGAUA----CAUAUGUAA ...(((((..((((.(((..((.(.......--.).))..)))))))..(((.((((((((((....))))))))))(((((.....))))).........)))...----..))))).. ( -28.50) >DroSim_CAF1 5453 114 + 1 UAAAUAUAAACUAGGCAGGUUGCUUAUUUUU--UACCAUUCUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCAUUUAUCUGUAGAUA----CAUAUGUAA ...(((((...(((((.....))))).....--......(((((.((......((((((((((....))))))))))(((((.....))))).....)).)))))..----..))))).. ( -28.20) >DroEre_CAF1 6174 113 + 1 UAAAUAUAAACUAGGCAGGUUGCUUUUUUU---UACCAUUCUGCUGUCACUAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCAUUUAUCUCUACAUAUG----UAUAUAA ....((((.((..(((((..((........---...))..)))))........((((((((((....))))))))))(((((.....)))))................)----).)))). ( -25.30) >DroYak_CAF1 5467 120 + 1 UAAAUAUAAACUAGGCAGGUUAAUUUUUUUUUCUACCAUUCUGCUGGCACCAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCUUUUAUCUCUACAUAUAUACAUAUAUAA .....(((((..((((.(((..(........)..))).....(((((((....((((((((((....))))))))))..)))...))))))))))))).......(((((....))))). ( -24.40) >consensus UAAAUAUAAACUAGGCAGGUUGCUUUUUUUU__UACCAUUCUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCAUUUAUCUGUACAUA____CAUAUGUAA ...(((((.....((((((....................))))))........((((((((((....))))))))))(((((.....))))).....................))))).. (-24.67 = -24.27 + -0.40)

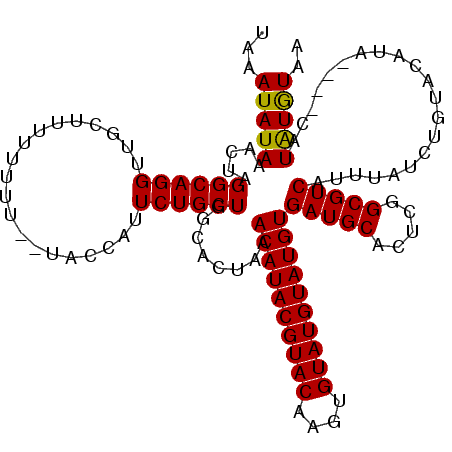

| Location | 15,826,834 – 15,826,939 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.91 |
| Mean single sequence MFE | -27.00 |
| Consensus MFE | -20.72 |
| Energy contribution | -20.50 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.96 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.92 |
| SVM RNA-class probability | 0.982618 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15826834 105 + 22224390 CUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCAUUUAUCUGU----A----CAUGUGUAAAAUGUAACGUUACCUGUGGUACACAGCAAUAAA------- .(((((((((...((((((((((....))))))))))(((((.....)))))..........----.----...))))..........((.(((...))))).))))).....------- ( -30.80) >DroEre_CAF1 6211 115 + 1 CUGCUGUCACUAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCAUUUAUCUCUACAUAUG----UAUAUAAAAUGUAAC-UAAACUGUGCCACCCAACAAUAAAUAAAUAA .(((.(((((...(((((((......)))))))..))))))))...((((.((((((....(((((...----........)))))..-)))).)))))).................... ( -21.80) >DroYak_CAF1 5507 119 + 1 CUGCUGGCACCAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCUUUUAUCUCUACAUAUAUACAUAUAUAAAAUGUAAC-UUAACUGUGCCACCCAACAAUAAAUAAAUAA ....((((((...((((((((((....))))))))))(((((.....))))).................(((((.......)))))..-......))))))................... ( -28.40) >consensus CUGCUGGCACUAAACAUACGUACAAGUGUAUGUAUGUGAUGCACUCGGCGUCAUUUAUCUCUACAUAU___CAUAUAUAAAAUGUAAC_UUAACUGUGCCACCCAACAAUAAAUAAAUAA ....((.(((...((((((((((....))))))))))(((((.....)))))...........................................))).))................... (-20.72 = -20.50 + -0.22)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:07:45 2006