
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,771,806 – 15,772,006 |
| Length | 200 |
| Max. P | 0.955396 |

| Location | 15,771,806 – 15,771,926 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.94 |
| Mean single sequence MFE | -27.91 |
| Consensus MFE | -20.66 |
| Energy contribution | -22.73 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.09 |
| Mean z-score | -3.30 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.928752 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15771806 120 + 22224390 GAUGAUAGCUAGCGAAAGUGUGUUAAAACACAACAAAAUAAUAAUGAAAAGGUGGUAACUUUGGCGAGCCAAAGUUCAAAUACCGAUGCAGUUAGAAAAGAAAAAGUGCAGGUUGUUAUA .(((((((((.(((...((((......)))).............((....((((..((((((((....))))))))....))))....))................))).))))))))). ( -30.60) >DroSec_CAF1 2390 120 + 1 GAUGAUAGCUAGCGAAAUUGUGUUAUAACACAAAAAAAUAAUAAUGAAAAGGUGGUAACUUCGGCGAGCCAAAGUUCAAAUACCGAUGCAGUUAGGAAAGAAAAAGUGCAGUUUGUUAUA ....(((((.(((....((((((....)))))).................((((..(((((.((....)).)))))....))))..((((.((..........)).))))))).))))). ( -23.50) >DroEre_CAF1 2542 116 + 1 GAUGAUAGCUAGCACAAGCGUGUUAUAACAGAA-AAAAUAAUAACGAAAAGG---UAACUUUGGCGAGCCAAAGUUCAAGUACCGAUGCAGUUAAAAAUGAAAAAGUGCAGGUUGUUAUA .(((((((((.((((..(((((((((.......-......))))).....((---(((((((((....))))))).....)))).)))).........(....).)))).))))))))). ( -30.42) >DroYak_CAF1 2519 116 + 1 GUUGAUAGCUAGCAAAAGCGUGUUAUAACACAA-AAAAUAAUAACGAAAAGG---UAACUUCGGCGAGCCAAAGUUCAAAUACCGAUGCAGUUAGAAAAGAAAAAGUGCAGGUUGCCUUA ((((...(((......)))((((....))))..-.......))))...((((---(((((((((.((((....)))).....))))((((.((..........)).))))))))))))). ( -27.10) >consensus GAUGAUAGCUAGCAAAAGCGUGUUAUAACACAA_AAAAUAAUAACGAAAAGG___UAACUUCGGCGAGCCAAAGUUCAAAUACCGAUGCAGUUAGAAAAGAAAAAGUGCAGGUUGUUAUA .(((((((((.(((.....((((....))))..........((((.....((((..((((((((....))))))))....))))......))))............))).))))))))). (-20.66 = -22.73 + 2.06)


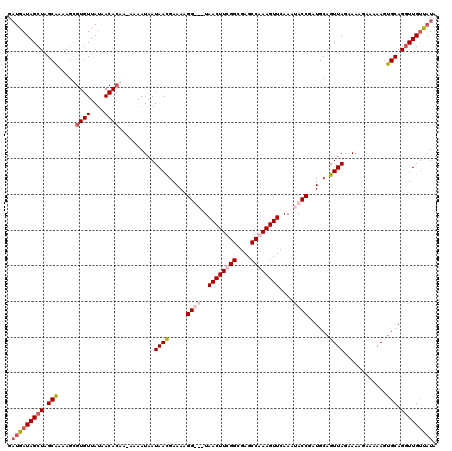
| Location | 15,771,806 – 15,771,926 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.94 |
| Mean single sequence MFE | -20.18 |
| Consensus MFE | -16.02 |
| Energy contribution | -16.40 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.852926 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15771806 120 - 22224390 UAUAACAACCUGCACUUUUUCUUUUCUAACUGCAUCGGUAUUUGAACUUUGGCUCGCCAAAGUUACCACCUUUUCAUUAUUAUUUUGUUGUGUUUUAACACACUUUCGCUAGCUAUCAUC ((((((((..((((.((..........)).))))..(((.....((((((((....))))))))...)))..............))))))))............................ ( -19.10) >DroSec_CAF1 2390 120 - 1 UAUAACAAACUGCACUUUUUCUUUCCUAACUGCAUCGGUAUUUGAACUUUGGCUCGCCGAAGUUACCACCUUUUCAUUAUUAUUUUUUUGUGUUAUAACACAAUUUCGCUAGCUAUCAUC .............................((((...(((.....((((((((....))))))))...))).................((((((....))))))....)).))........ ( -18.90) >DroEre_CAF1 2542 116 - 1 UAUAACAACCUGCACUUUUUCAUUUUUAACUGCAUCGGUACUUGAACUUUGGCUCGCCAAAGUUA---CCUUUUCGUUAUUAUUUU-UUCUGUUAUAACACGCUUGUGCUAGCUAUCAUC ((((((((((((((.((..........)).))))..))).....((((((((....)))))))).---..................-...)))))))....(((......)))....... ( -21.00) >DroYak_CAF1 2519 116 - 1 UAAGGCAACCUGCACUUUUUCUUUUCUAACUGCAUCGGUAUUUGAACUUUGGCUCGCCGAAGUUA---CCUUUUCGUUAUUAUUUU-UUGUGUUAUAACACGCUUUUGCUAGCUAUCAAC ...(((((...((.......................((((.....(((((((....)))))))))---))................-..((((....))))))..))))).......... ( -21.70) >consensus UAUAACAACCUGCACUUUUUCUUUUCUAACUGCAUCGGUAUUUGAACUUUGGCUCGCCAAAGUUA___CCUUUUCAUUAUUAUUUU_UUGUGUUAUAACACACUUUCGCUAGCUAUCAUC (((((((...((((.((..........)).))))..(((.....((((((((....))))))))...)))....................)))))))....................... (-16.02 = -16.40 + 0.38)



| Location | 15,771,846 – 15,771,966 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.05 |
| Mean single sequence MFE | -27.98 |
| Consensus MFE | -22.51 |
| Energy contribution | -23.70 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.691154 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15771846 120 + 22224390 AUAAUGAAAAGGUGGUAACUUUGGCGAGCCAAAGUUCAAAUACCGAUGCAGUUAGAAAAGAAAAAGUGCAGGUUGUUAUAGUCGUUCGAUUUUCCCACAAUUUGCAAUCUUGUCCUAAAA ....((....((((..((((((((....))))))))....))))....)).((((..((((.....((((((((((...((((....)))).....)))))))))).))))...)))).. ( -31.10) >DroSec_CAF1 2430 120 + 1 AUAAUGAAAAGGUGGUAACUUCGGCGAGCCAAAGUUCAAAUACCGAUGCAGUUAGGAAAGAAAAAGUGCAGUUUGUUAUAGUCGUUCGAUUUUCCCACAAAUUGCAAUCUUGUCCUAAAA ....((....((((..(((((.((....)).)))))....))))....)).((((((((((.....((((((((((...((((....)))).....)))))))))).)))).)))))).. ( -32.10) >DroEre_CAF1 2581 117 + 1 AUAACGAAAAGG---UAACUUUGGCGAGCCAAAGUUCAAGUACCGAUGCAGUUAAAAAUGAAAAAGUGCAGGUUGUUAUAAUCGUCCGAUUUUCCCACAAUUUGCAAUCUUGUCCUAAAA .((((.....((---(((((((((....))))))).....))))......)))).....((.....((((((((((...((((....)))).....))))))))))......))...... ( -27.20) >DroYak_CAF1 2558 117 + 1 AUAACGAAAAGG---UAACUUCGGCGAGCCAAAGUUCAAAUACCGAUGCAGUUAGAAAAGAAAAAGUGCAGGUUGCCUUAGACGUCCGAUUUUCCCACAAUUUGCAAUCUUGUCCUAAAA ........((((---(((((((((.((((....)))).....))))((((.((..........)).))))))))))))).((((...((((..............)))).))))...... ( -21.54) >consensus AUAACGAAAAGG___UAACUUCGGCGAGCCAAAGUUCAAAUACCGAUGCAGUUAGAAAAGAAAAAGUGCAGGUUGUUAUAGUCGUCCGAUUUUCCCACAAUUUGCAAUCUUGUCCUAAAA .((((.....((((..((((((((....))))))))....))))......)))).....((.....((((((((((...((((....)))).....))))))))))......))...... (-22.51 = -23.70 + 1.19)

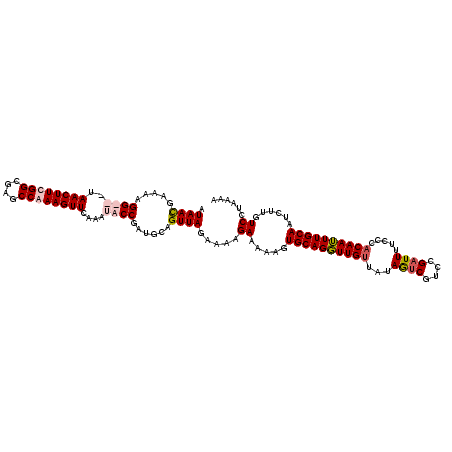

| Location | 15,771,886 – 15,772,006 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.90 |
| Mean single sequence MFE | -29.55 |
| Consensus MFE | -25.04 |
| Energy contribution | -26.72 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.955396 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15771886 120 + 22224390 UACCGAUGCAGUUAGAAAAGAAAAAGUGCAGGUUGUUAUAGUCGUUCGAUUUUCCCACAAUUUGCAAUCUUGUCCUAAAAUCGCCCUGAAAAUUCCCAUAUAAUAUAUGGGCAACGGAAA ..(((.(((..((((..((((.....((((((((((...((((....)))).....)))))))))).))))...)))).................((((((...))))))))).)))... ( -29.20) >DroSec_CAF1 2470 120 + 1 UACCGAUGCAGUUAGGAAAGAAAAAGUGCAGUUUGUUAUAGUCGUUCGAUUUUCCCACAAAUUGCAAUCUUGUCCUAAAAUCGCCCUGAAAAUUCCCAUAUAAUAUAUGGGCAUCGGAAA ..(((((((..((((((((((.....((((((((((...((((....)))).....)))))))))).)))).)))))).................((((((...)))))))))))))... ( -38.90) >DroEre_CAF1 2618 118 + 1 UACCGAUGCAGUUAAAAAUGAAAAAGUGCAGGUUGUUAUAAUCGUCCGAUUUUCCCACAAUUUGCAAUCUUGUCCUAAAAUCGGUCUGAAAAUUCCCAUAUAAUAUA--GGCAUCGGAAA ..(((((((.................((((((((((...((((....)))).....)))))))))).((..(.((.......)).).))..................--.)))))))... ( -27.30) >DroYak_CAF1 2595 118 + 1 UACCGAUGCAGUUAGAAAAGAAAAAGUGCAGGUUGCCUUAGACGUCCGAUUUUCCCACAAUUUGCAAUCUUGUCCUAAAAUCGCCCUGAAAAUUCCCAUAUAAUAUA--GGCAUCGGAAA ..(((((((..((((..((((.....(((((((((....((((....).))).....))))))))).))))...))))..((.....))..................--.)))))))... ( -22.80) >consensus UACCGAUGCAGUUAGAAAAGAAAAAGUGCAGGUUGUUAUAGUCGUCCGAUUUUCCCACAAUUUGCAAUCUUGUCCUAAAAUCGCCCUGAAAAUUCCCAUAUAAUAUA__GGCAUCGGAAA ..(((((((..((((..((((.....((((((((((...((((....)))).....)))))))))).))))...)))).................((((((...)))))))))))))... (-25.04 = -26.72 + 1.69)


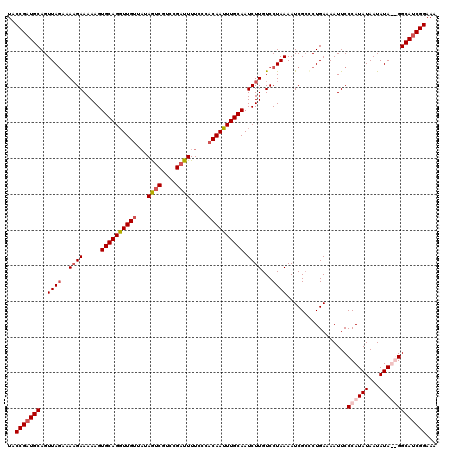
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:06:38 2006