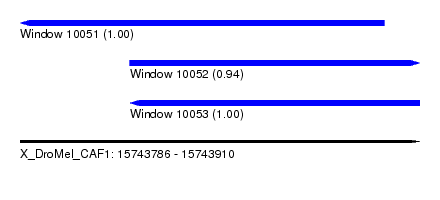

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,743,786 – 15,743,910 |
| Length | 124 |
| Max. P | 0.999597 |

| Location | 15,743,786 – 15,743,899 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 78.27 |
| Mean single sequence MFE | -24.73 |
| Consensus MFE | -15.19 |
| Energy contribution | -17.25 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.24 |
| Mean z-score | -3.51 |
| Structure conservation index | 0.61 |
| SVM decision value | 3.77 |
| SVM RNA-class probability | 0.999597 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
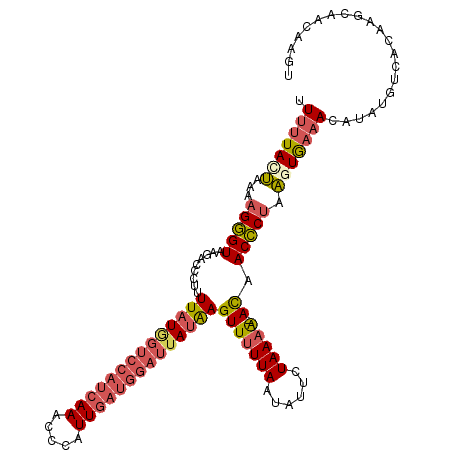
>X_DroMel_CAF1 15743786 113 - 22224390 UUUUUAUUAUAAGGGUAAGACCCUUUUAUAGUCCAUCAAACCCAUUGAUGGAUUAUAAGUUUUUAAUGUUCUAAUA-ACAACCCUAAGUGAAACAUAUUUCACAAGCAACAAGU ...........(((((.........((((((((((((((.....))))))))))))))(((.(((......))).)-)).)))))..((((((....))))))........... ( -29.10) >DroSec_CAF1 2783 113 - 1 UUUUUACUAAAAGGGUAAGACAGUUUUAUGGUCCAUCAAGCCCAUUGAUGGAUUAUAAGUUUUUAAUAUUCUAAAA-ACAACCCUAAGUGAAACAUAUGUUACAAGCAACAAGU .(((((((...(((((.........((((((((((((((.....))))))))))))))(((((((......)))))-)).))))).)))))))....((((......))))... ( -32.60) >DroSim_CAF1 2789 113 - 1 UUUUUAUUAAACGGGUAAGACCGUUUUAUGGUCCAUCAAAUCCAUUGAUGGAUUAUAAGUUUUUAAUAUUCUAAAA-ACAACCCUAAUUAAAACAUAUGUUACAAGCAACAAGU ............((((.........((((((((((((((.....))))))))))))))(((((((......)))))-)).)))).............((((......))))... ( -26.70) >DroYak_CAF1 2821 108 - 1 UUUUUACAAAAAGUGUUU-CCUCUUUUAUUGUGCAUCAAACCCAUUUU-----UAAGAGUUUUUAUUAUUCUAAAGAAUAACACUCUAUGCUAGACAUUUCACAAAAAAACAGC ..........((((((((-.(((((.....(((.........)))...-----.)))))......((((((....))))))...........)))))))).............. ( -10.50) >consensus UUUUUACUAAAAGGGUAAGACCCUUUUAUGGUCCAUCAAACCCAUUGAUGGAUUAUAAGUUUUUAAUAUUCUAAAA_ACAACCCUAAGUGAAACAUAUGUCACAAGCAACAAGU .(((((((...(((((.........((((((((((((((.....))))))))))))))(((((((......))))).)).))))).)))))))..................... (-15.19 = -17.25 + 2.06)


| Location | 15,743,820 – 15,743,910 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 68.88 |
| Mean single sequence MFE | -18.52 |
| Consensus MFE | -2.14 |
| Energy contribution | -4.38 |
| Covariance contribution | 2.24 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.12 |
| SVM decision value | 1.34 |
| SVM RNA-class probability | 0.943682 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
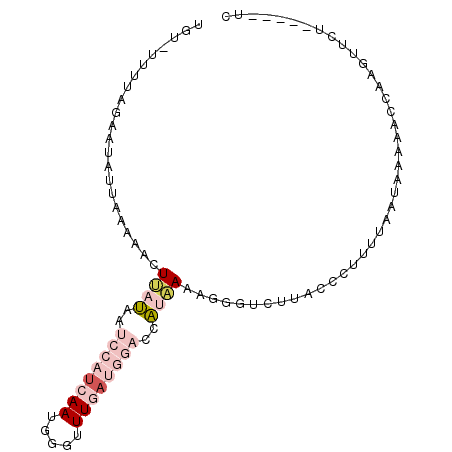
>X_DroMel_CAF1 15743820 90 + 22224390 UGU-UAUUAGAACAUUAAAAACUUAUAAUCCAUCAAUGGGUUUGAUGGACUAUAAAAGGGUCUUACCCUUAUAAUAAAAACAAAGUUCU-----UC ...-....(((((.........(((((.((((((((.....)))))))).)))))(((((.....)))))..............)))))-----.. ( -24.20) >DroSec_CAF1 2817 90 + 1 UGU-UUUUAGAAUAUUAAAAACUUAUAAUCCAUCAAUGGGCUUGAUGGACCAUAAAACUGUCUUACCCUUUUAGUAAAAACGAAGUUCU-----UC .((-((((..............((((..((((((((.....))))))))..)))).((((...........))))))))))........-----.. ( -13.80) >DroSim_CAF1 2823 90 + 1 UGU-UUUUAGAAUAUUAAAAACUUAUAAUCCAUCAAUGGAUUUGAUGGACCAUAAAACGGUCUUACCCGUUUAAUAAAAACGAAGUUCU-----UC .((-(((((..............(((..((((((((.....))))))))..)))((((((......))))))..)))))))........-----.. ( -20.80) >DroEre_CAF1 2849 78 + 1 UAUAUUUUAGAAUAAUAAAAACUCUCU-----UAAAUGGGUUUGAUGGGGUGUGAAAGGGG-UG-----------GAAACUCA-CUUCUUAGCUCC ..................((((((...-----.....))))))...(((((..(((..(((-(.-----------...)))).-.)))...))))) ( -18.20) >DroYak_CAF1 2855 89 + 1 UAUUCUUUAGAAUAAUAAAAACUCUUA-----AAAAUGGGUUUGAUGCACAAUAAAAGAGG-AAACACUUUUUGUAAAAACCA-AUUCUAAGCUUC .....((((((((.....((((((...-----.....))))))..((((....(((((.(.-...).))))))))).......-)))))))).... ( -15.60) >consensus UGU_UUUUAGAAUAUUAAAAACUUAUAAUCCAUCAAUGGGUUUGAUGGACCAUAAAAGGGUCUUACCCUUUUAAUAAAAACCAAGUUCU_____UC ......................((((..((((((((.....))))))))..))))......................................... ( -2.14 = -4.38 + 2.24)


| Location | 15,743,820 – 15,743,910 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 68.88 |
| Mean single sequence MFE | -19.44 |
| Consensus MFE | -6.64 |
| Energy contribution | -9.24 |
| Covariance contribution | 2.60 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.68 |
| Structure conservation index | 0.34 |
| SVM decision value | 2.74 |
| SVM RNA-class probability | 0.996734 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15743820 90 - 22224390 GA-----AGAACUUUGUUUUUAUUAUAAGGGUAAGACCCUUUUAUAGUCCAUCAAACCCAUUGAUGGAUUAUAAGUUUUUAAUGUUCUAAUA-ACA ..-----(((((..............(((((.....)))))((((((((((((((.....)))))))))))))).........)))))....-... ( -26.50) >DroSec_CAF1 2817 90 - 1 GA-----AGAACUUCGUUUUUACUAAAAGGGUAAGACAGUUUUAUGGUCCAUCAAGCCCAUUGAUGGAUUAUAAGUUUUUAAUAUUCUAAAA-ACA ((-----(....)))(((((.(((.....))))))))....((((((((((((((.....))))))))))))))(((((((......)))))-)). ( -25.00) >DroSim_CAF1 2823 90 - 1 GA-----AGAACUUCGUUUUUAUUAAACGGGUAAGACCGUUUUAUGGUCCAUCAAAUCCAUUGAUGGAUUAUAAGUUUUUAAUAUUCUAAAA-ACA ((-----(....)))(((((((...(((((......)))))((((((((((((((.....)))))))))))))).............)))))-)). ( -26.70) >DroEre_CAF1 2849 78 - 1 GGAGCUAAGAAG-UGAGUUUC-----------CA-CCCCUUUCACACCCCAUCAAACCCAUUUA-----AGAGAGUUUUUAUUAUUCUAAAAUAUA ((((((......-..))))))-----------..-.............................-----..(((((.......)))))........ ( -5.80) >DroYak_CAF1 2855 89 - 1 GAAGCUUAGAAU-UGGUUUUUACAAAAAGUGUUU-CCUCUUUUAUUGUGCAUCAAACCCAUUUU-----UAAGAGUUUUUAUUAUUCUAAAGAAUA ((((((((((((-.(((((.(((((((((.....-...))))..)))))....))))).)))))-----...)))))))...(((((....))))) ( -13.20) >consensus GA_____AGAACUUCGUUUUUACUAAAAGGGUAAGACCCUUUUAUAGUCCAUCAAACCCAUUGAUGGAUUAUAAGUUUUUAAUAUUCUAAAA_ACA ........................................(((((((((((((((.....)))))))))))))))..................... ( -6.64 = -9.24 + 2.60)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:06:19 2006