
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,731,930 – 15,732,083 |
| Length | 153 |
| Max. P | 0.733123 |

| Location | 15,731,930 – 15,732,043 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.79 |
| Mean single sequence MFE | -25.67 |
| Consensus MFE | -18.58 |
| Energy contribution | -18.86 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.32 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.655947 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15731930 113 - 22224390 AUGAGGGAUGCUGGAUGAAACCACCAGGCGACUGACUGGCACACAAUGAUUUGGAAGGAAACAGCGUCCUAAAAGUUCCGAAAAGGAAGGAAAUAGGUAGAGUUAU-------AAGCCCA ....(((((((((.......(((.(((....)))..)))....(((....)))........))))))))).....((((.....)))).......(((........-------..))).. ( -26.50) >DroSec_CAF1 4748 111 - 1 AUGAGGGAUGCUGGAUGAAA---------GGUCGACUGGUGUACAAUGAGUUGGAAGGAAACAGCAUCCUAAAAGUCCCGAAAAGGAAGGAAAUAGAUAGUCAUGUAUAGCUGAAGCCUA ....(((((...(((((...---------..((((((.((.....)).))))))..(....)..))))).....)))))....(((..((..(((.((....)).)))..))....))). ( -28.40) >DroSim_CAF1 5064 111 - 1 AUGAGGGAUGCUGGAUGAAA---------GGUCGGCUGGUUUACAAUGAGUUGGAAGGAAACAGCAUCCUAAAAGUCCCGAAAAGGAAGGAAAUAGAUAGUUAUGUAUAGCUGAAGCCCA ....(((((((((.......---------..((((((.((.....)).)))))).......))))))))).....(((......)))..........((((((....))))))....... ( -25.69) >DroEre_CAF1 4772 107 - 1 AUGAGGGAAUCUGGAUGAAA---------GGUCGACUGGCUUACAAUGAGUUGGAAGGAAACAACCUCUUAAAAGUCCCGAAAAGGAAGGAAAUAGGUAGUU----ACAGCUUACACCCA ....(((............(---------((((....)))))....(((((((...(....).((((........(((......))).......))))....----.)))))))..))). ( -22.66) >DroYak_CAF1 4902 107 - 1 AUGAGGGAAACUGGAUGAAA---------GAUCGACUGGUUUACAAUGAGUUGGAAGGAAACAACAUCCUAAAAGUCCCGAAAAGGAAGGAAAUAGAUAGCU----AUAGCUUACACCCA ....((((....(((((...---------..((((((.((.....)).))))))..(....)..)))))......)))).....((.(((..((((....))----))..)))....)). ( -25.10) >consensus AUGAGGGAUGCUGGAUGAAA_________GGUCGACUGGUUUACAAUGAGUUGGAAGGAAACAGCAUCCUAAAAGUCCCGAAAAGGAAGGAAAUAGAUAGUU_U_UAUAGCUGAAGCCCA ....(((((...(((((..............((((((.((.....)).))))))..(....)..))))).....)))))......................................... (-18.58 = -18.86 + 0.28)


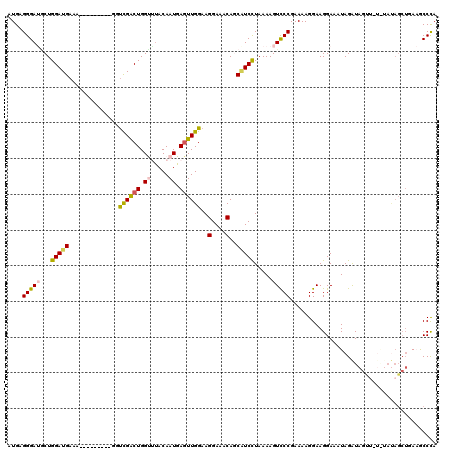
| Location | 15,731,963 – 15,732,083 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.65 |
| Mean single sequence MFE | -28.66 |
| Consensus MFE | -24.40 |
| Energy contribution | -24.60 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.733123 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15731963 120 - 22224390 GCUGUUCAAUUCAGAGGCAGGAAGGAAGUUUCUGGCAUAAAUGAGGGAUGCUGGAUGAAACCACCAGGCGACUGACUGGCACACAAUGAUUUGGAAGGAAACAGCGUCCUAAAAGUUCCG ((((((...(((...(.(((...((.....((..((((.........))))..)).....))..(((....))).))).)...(((....))))))...))))))............... ( -30.20) >DroSec_CAF1 4788 110 - 1 GCA-UAGUAUUCAGAGGCAGGAAGGAAGUUUCUGGCAUAAAUGAGGGAUGCUGGAUGAAA---------GGUCGACUGGUGUACAAUGAGUUGGAAGGAAACAGCAUCCUAAAAGUCCCG .((-(..(((((((((((.........)))))))).))).))).(((((...(((((...---------..((((((.((.....)).))))))..(....)..))))).....))))). ( -32.10) >DroSim_CAF1 5104 110 - 1 GCA-UAGUAUUCAGAGGCAGGAAGGAAGUUUCUGGCAUAAAUGAGGGAUGCUGGAUGAAA---------GGUCGGCUGGUUUACAAUGAGUUGGAAGGAAACAGCAUCCUAAAAGUCCCG .((-(..(((((((((((.........)))))))).))).))).(((((...(((((...---------..((((((.((.....)).))))))..(....)..))))).....))))). ( -30.50) >DroEre_CAF1 4808 110 - 1 GCA-AAGUAUUUAAAGGCAACAAGGAAAUUGCUGGCAUAUAUGAGGGAAUCUGGAUGAAA---------GGUCGACUGGCUUACAAUGAGUUGGAAGGAAACAACCUCUUAAAAGUCCCG ...-..((((.....(((((........))))).....))))..((((..((...(((.(---------(((...(..((((.....))))..)..(....).)))).)))..)))))). ( -23.40) >DroYak_CAF1 4938 110 - 1 GCA-AAGUAUUUAAAGGCAACAAGGAAAUUGCUGGCAUAAAUGAGGGAAACUGGAUGAAA---------GAUCGACUGGUUUACAAUGAGUUGGAAGGAAACAACAUCCUAAAAGUCCCG ...-...((((((..(((((........)))))....)))))).((((....(((((...---------..((((((.((.....)).))))))..(....)..)))))......)))). ( -27.10) >consensus GCA_UAGUAUUCAGAGGCAGGAAGGAAGUUUCUGGCAUAAAUGAGGGAUGCUGGAUGAAA_________GGUCGACUGGUUUACAAUGAGUUGGAAGGAAACAGCAUCCUAAAAGUCCCG ...............(.((((((.....)))))).)........(((((...(((((..............((((((.((.....)).))))))..(....)..))))).....))))). (-24.40 = -24.60 + 0.20)

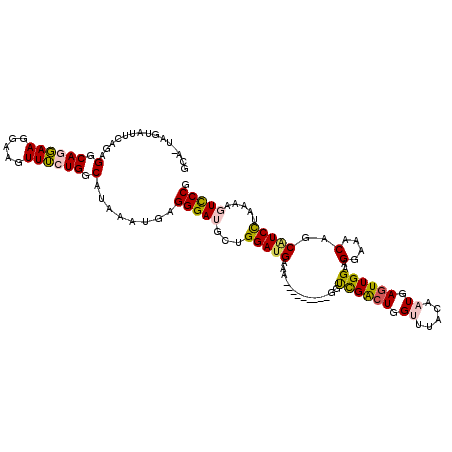

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:06:10 2006