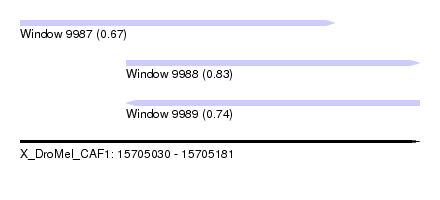
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,705,030 – 15,705,181 |
| Length | 151 |
| Max. P | 0.828279 |

| Location | 15,705,030 – 15,705,149 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.39 |
| Mean single sequence MFE | -23.44 |
| Consensus MFE | -15.05 |
| Energy contribution | -14.62 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.666373 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15705030 119 + 22224390 CUUUUGCUUAACAAAUUCCCUUCAAAUGCAAAGCAAAUGCUUGCUGUUACAGCUUUAAAAUGUGAUUUUCUUAU-UUUUCUAAGUGGGAAUGUCCCAUUUAGUUUCAAAUAUAUACAUAC .....((((((((..............((((.((....))))))))))).)))......(((((.......(((-((..(((((((((.....)))))))))....)))))..))))).. ( -22.83) >DroSec_CAF1 16217 116 + 1 CUUUUGUUUAACAAAUUCGUUUCCAAUGCAAAUU---UAUUUGCUGCAACAGCUUUAAAAUUUGAUUUUCUUAU-UUUUCUAAGUGGGAAUGUCCCAUUUAGUUUCAAAUAUAUACAUAC ..((((.....)))).............((((((---(....((((...))))....))))))).......(((-((..(((((((((.....)))))))))....)))))......... ( -21.20) >DroEre_CAF1 16018 116 + 1 CUU-UGUGUAAGAAAUUUGUUACCAAUGCAAAUU---UAUGUGUUGCAACAGCUUUAAAAUUUGAGUUUCUUUUUUUUUCUAAGUGGGAAUGUCCCAUUUAGUUCCAUAUAUAUACAUAC ...-((((((..((((((((.......)))))))---)(((((..((...(((((........)))))............((((((((.....))))))))))..))))).))))))... ( -26.30) >consensus CUUUUGUUUAACAAAUUCGUUUCCAAUGCAAAUU___UAUUUGCUGCAACAGCUUUAAAAUUUGAUUUUCUUAU_UUUUCUAAGUGGGAAUGUCCCAUUUAGUUUCAAAUAUAUACAUAC ..(((((....................))))).....(((((((((...))))..........................(((((((((.....)))))))))....)))))......... (-15.05 = -14.62 + -0.44)


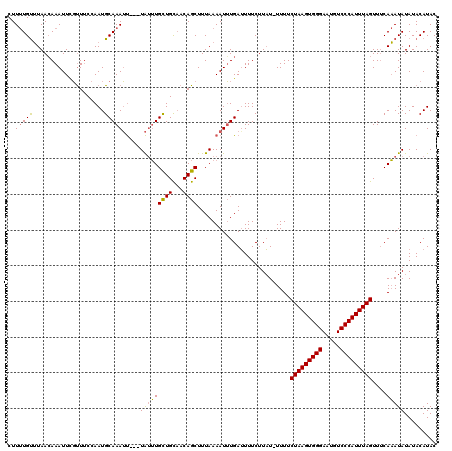
| Location | 15,705,070 – 15,705,181 |
|---|---|
| Length | 111 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 92.28 |
| Mean single sequence MFE | -27.53 |
| Consensus MFE | -21.41 |
| Energy contribution | -21.30 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.12 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.828279 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15705070 111 + 22224390 UUGCUGUUACAGCUUUAAAAUGUGAUUUUCUUAU-UUUUCUAAGUGGGAAUGUCCCAUUUAGUUUCAAAUAUAUACAUACAUAUGUGGGUCAAUGUAGCACCUACUCAA-AAA ..((((...))))......(((((.......(((-((..(((((((((.....)))))))))....)))))......)))))..(((((((......).))))))....-... ( -25.42) >DroSec_CAF1 16254 111 + 1 UUGCUGCAACAGCUUUAAAAUUUGAUUUUCUUAU-UUUUCUAAGUGGGAAUGUCCCAUUUAGUUUCAAAUAUAUACAUACAUAUGUGGGUCAAUGUAGCAUCUAUUCAA-AAA .(((((((...(((.....((((((.........-....(((((((((.....)))))))))..))))))...(((((....))))))))...))))))).........-... ( -26.16) >DroEre_CAF1 16054 113 + 1 GUGUUGCAACAGCUUUAAAAUUUGAGUUUCUUUUUUUUUCUAAGUGGGAAUGUCCCAUUUAGUUCCAUAUAUAUACAUACAUAUGUGGGUCAAUGUAGCACCUACUCAAAAAA ((((((((..(((((........)))))...........(((((((((.....))))))))).(((((((((........)))))))))....))))))))............ ( -31.00) >consensus UUGCUGCAACAGCUUUAAAAUUUGAUUUUCUUAU_UUUUCUAAGUGGGAAUGUCCCAUUUAGUUUCAAAUAUAUACAUACAUAUGUGGGUCAAUGUAGCACCUACUCAA_AAA .(((((((...(((.........................(((((((((.....)))))))))...........(((((....))))))))...)))))))............. (-21.41 = -21.30 + -0.11)


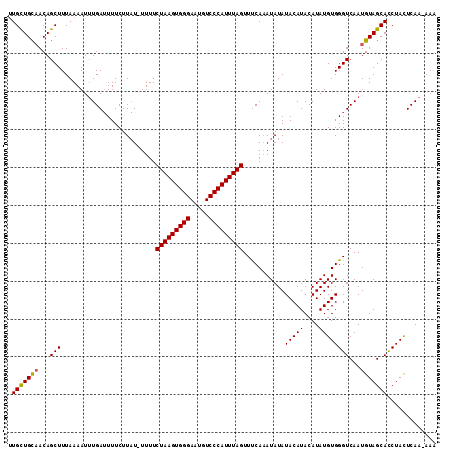
| Location | 15,705,070 – 15,705,181 |
|---|---|
| Length | 111 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 92.28 |
| Mean single sequence MFE | -24.50 |
| Consensus MFE | -19.23 |
| Energy contribution | -19.23 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.738513 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15705070 111 - 22224390 UUU-UUGAGUAGGUGCUACAUUGACCCACAUAUGUAUGUAUAUAUUUGAAACUAAAUGGGACAUUCCCACUUAGAAAA-AUAAGAAAAUCACAUUUUAAAGCUGUAACAGCAA .((-(..((((.(((((((((((.....)).))))).)))).))))..)))((((.((((.....)))).))))....-.....................((((...)))).. ( -24.30) >DroSec_CAF1 16254 111 - 1 UUU-UUGAAUAGAUGCUACAUUGACCCACAUAUGUAUGUAUAUAUUUGAAACUAAAUGGGACAUUCCCACUUAGAAAA-AUAAGAAAAUCAAAUUUUAAAGCUGUUGCAGCAA .((-(..((((.(((((((((((.....)).))))).)))).))))..)))((((.((((.....)))).))))....-.....................((((...)))).. ( -24.00) >DroEre_CAF1 16054 113 - 1 UUUUUUGAGUAGGUGCUACAUUGACCCACAUAUGUAUGUAUAUAUAUGGAACUAAAUGGGACAUUCCCACUUAGAAAAAAAAAGAAACUCAAAUUUUAAAGCUGUUGCAACAC ...(((((((.((..(......)..)).((((((((....))))))))...((((.((((.....)))).))))............))))))).......((....))..... ( -25.20) >consensus UUU_UUGAGUAGGUGCUACAUUGACCCACAUAUGUAUGUAUAUAUUUGAAACUAAAUGGGACAUUCCCACUUAGAAAA_AUAAGAAAAUCAAAUUUUAAAGCUGUUGCAGCAA ....(((((((.(((((((((((.....)).))))).)))).)))))))..((((.((((.....)))).))))..........................((((...)))).. (-19.23 = -19.23 + 0.01)


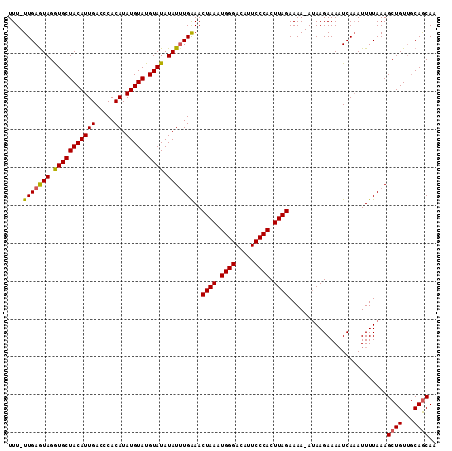
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:05:20 2006