
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,701,612 – 15,701,718 |
| Length | 106 |
| Max. P | 0.950674 |

| Location | 15,701,612 – 15,701,718 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.29 |
| Mean single sequence MFE | -25.30 |
| Consensus MFE | -20.06 |
| Energy contribution | -19.86 |
| Covariance contribution | -0.21 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.950674 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15701612 106 + 22224390 -------UAAUGAAAUUGAUGAAUAUGUAUUUAUGUGGGGAUGU------UAGGAUUUUUCCAUUUGCAUUUACUUAUGUACAUAUUUAUGUACAAUAUAACCAAUUGAUGUUCAUCA-G -------.........((((((((((.((......(((((((((------..(((....)))....)))))).....(((((((....)))))))......)))..))))))))))))-. ( -25.00) >DroSec_CAF1 12761 113 + 1 -------UAAUGAAAUUGAUGAAUAUAUAUUUAUGUAGGGAUGUACGUAUAAGGAUUUUUCCAUUUGCAUUUACAUAUGUACGUAUUUAUGUACAAUAUAACCAAUUGAUGUUCAUCAAG -------........(((((((((((.....((((((..((((((.((....(((....))))).))))))))))))(((((((....))))))).............))))))))))). ( -27.40) >DroEre_CAF1 12475 119 + 1 GUACAUAUAUUUAAAUUGAUGAAUACGUAUUUAUGUUGGGAUGUACAUAUAAGGAUUUUUCCAUUUGCAUUUAUGU-CAUAUAUAUUUAUGUACAAUAUGACCAAUUGAUGUUCAUCAAG ...............((((((((((((...((((((((..............(((....)))...(((((..((((-.....))))..))))))))))))).....)).)))))))))). ( -23.50) >consensus _______UAAUGAAAUUGAUGAAUAUGUAUUUAUGUAGGGAUGUAC_UAUAAGGAUUUUUCCAUUUGCAUUUACAUAUGUACAUAUUUAUGUACAAUAUAACCAAUUGAUGUUCAUCAAG ...............((((((((((((((..((((((.(.(((((.......(((....))).........))))).).))))))..))))).((((.......))))..))))))))). (-20.06 = -19.86 + -0.21)


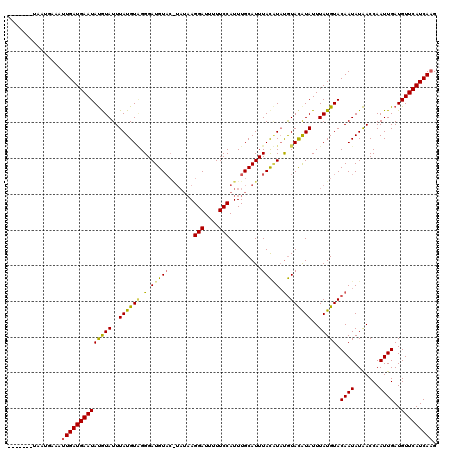
| Location | 15,701,612 – 15,701,718 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.29 |
| Mean single sequence MFE | -17.90 |
| Consensus MFE | -11.93 |
| Energy contribution | -11.93 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.821794 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
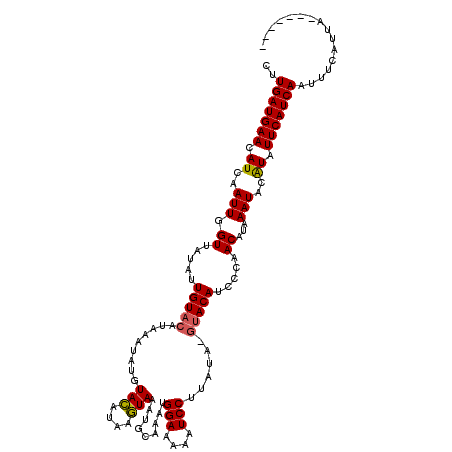
>X_DroMel_CAF1 15701612 106 - 22224390 C-UGAUGAACAUCAAUUGGUUAUAUUGUACAUAAAUAUGUACAUAAGUAAAUGCAAAUGGAAAAAUCCUA------ACAUCCCCACAUAAAUACAUAUUCAUCAAUUUCAUUA------- .-(((((((......(((.((((..(((((((....)))))))...))))...)))..(((....)))..------.....................))))))).........------- ( -16.50) >DroSec_CAF1 12761 113 - 1 CUUGAUGAACAUCAAUUGGUUAUAUUGUACAUAAAUACGUACAUAUGUAAAUGCAAAUGGAAAAAUCCUUAUACGUACAUCCCUACAUAAAUAUAUAUUCAUCAAUUUCAUUA------- .((((((((..(((.(((.(((((((((((........))))).))))))...))).)))..............(((......)))...........))))))))........------- ( -17.80) >DroEre_CAF1 12475 119 - 1 CUUGAUGAACAUCAAUUGGUCAUAUUGUACAUAAAUAUAUAUG-ACAUAAAUGCAAAUGGAAAAAUCCUUAUAUGUACAUCCCAACAUAAAUACGUAUUCAUCAAUUUAAAUAUAUGUAC .((((((((.........((((((((((......))).)))))-)).....(((....(((....)))...(((((........))))).....)))))))))))............... ( -19.40) >consensus CUUGAUGAACAUCAAUUGGUUAUAUUGUACAUAAAUAUGUACAUAAGUAAAUGCAAAUGGAAAAAUCCUUAUA_GUACAUCCCAACAUAAAUACAUAUUCAUCAAUUUCAUUA_______ ..(((((((.((..(((.((.....(((((.........(((....))).........(((....)))......))))).....))...)))..)).)))))))................ (-11.93 = -11.93 + 0.00)

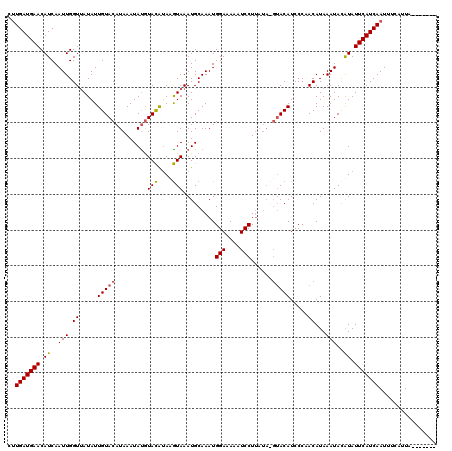
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:05:10 2006