
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,687,046 – 15,687,204 |
| Length | 158 |
| Max. P | 0.692133 |

| Location | 15,687,046 – 15,687,165 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.29 |
| Mean single sequence MFE | -29.62 |
| Consensus MFE | -24.87 |
| Energy contribution | -25.91 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.692133 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15687046 119 - 22224390 UAUACCGACCAGUGGAGAACGCCUCGAAUUGGCUGUUCACGUCACUC-GCACGCUGGUCACACCGAUUUUGUAAAUGCAAAAAUUGAGUAUGUAUAAGUAUUUACCAUCGUGUCGUUAUA ......((((((((..(((((((.......))).)))).((.....)-)..))))))))((((.(((...((((((((...................)))))))).)))))))....... ( -32.51) >DroSec_CAF1 2713 118 - 1 UAUACCGAUCAGUGGAGAACGCCUUGAAUUGGCUGUUCACGCCAUUC-GCACGCUGGUCACACCGAUUUUGUAAAUGCAA-AAUUGAGUAUGUAUAAGUAUUUACCAUCGUGUCGUUAUA ......((((((((..(((((((.......))).))))..((.....-)).))))))))(((((((((((((....))))-))))).(((.(((....))).)))....))))....... ( -32.90) >DroSim_CAF1 2805 119 - 1 UAUACCGACCAGUGGAGAACGCCUUGAAUUGGCUGUUCACGCCACUC-GCACGCUGGUCACACCGAUUUUGUAAAUGCAAAAAUUGAGUAUGUAUAAGUAUUUACCAUCGUGUCGUUAUA ......((((((((..(((((((.......))).))))..((.....-)).))))))))((((.(((...((((((((...................)))))))).)))))))....... ( -34.01) >DroEre_CAF1 4040 119 - 1 UAUACUGACAAGUGGAGAACGCCUCAAAUUGCCUGUUUUCGCCACUC-GCACCCUGGUCACACUGAUUUUAUAAAUGCAAAAAUUGAGUAUGUAUAAGUAUUUACCAUCGUGUCGUUAUA .....((((.(((((.(((.((............)).))).))))).-((((..((((......(((((((((.((((.........)))).)))))).))).))))..)))).)))).. ( -20.10) >DroYak_CAF1 2587 120 - 1 UAUACUGACCAUUGGUGAACGCCUCAAAUUGGCUGUUUACGCCAUUUGGCACCCUGGCCACACUGAUUUUAUAAAUGCAAAAAUUGAGUAUGUAUAAGUAUUUACCAUCGUGUCGUUAUA ((((.(((((..(((((((..(.(((...(((((......(((....))).....)))))...)))..(((((.((((.........)))).))))))..)))))))..).)))).)))) ( -28.60) >consensus UAUACCGACCAGUGGAGAACGCCUCGAAUUGGCUGUUCACGCCACUC_GCACGCUGGUCACACCGAUUUUGUAAAUGCAAAAAUUGAGUAUGUAUAAGUAUUUACCAUCGUGUCGUUAUA ......((((((((..(((((((.......))).))))..((......)).))))))))((((.(((...((((((((...................)))))))).)))))))....... (-24.87 = -25.91 + 1.04)



| Location | 15,687,086 – 15,687,204 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 88.68 |
| Mean single sequence MFE | -32.94 |
| Consensus MFE | -24.42 |
| Energy contribution | -25.02 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.559594 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15687086 118 - 22224390 UAUCAGCUGUUUGAAUGCACAACCUGUGCAAUGCGAAAAUAUACCGACCAGUGGAGAACGCCUCGAAUUGGCUGUUCACGUCACUC-GCACGCUGGUCACACCGAUUUUGUAAAUGCAA .....((........(((((.....))))).(((((((.......((((((((..(((((((.......))).)))).((.....)-)..)))))))).......)))))))...)).. ( -34.74) >DroSec_CAF1 2752 118 - 1 UAUCAGCUGUUUGAAUGCACAACCUGUGCAAUACGAAAAUAUACCGAUCAGUGGAGAACGCCUUGAAUUGGCUGUUCACGCCAUUC-GCACGCUGGUCACACCGAUUUUGUAAAUGCAA .....((........(((((.....))))).(((((((.......((((((((..(((((((.......))).))))..((.....-)).)))))))).......)))))))...)).. ( -33.64) >DroSim_CAF1 2845 118 - 1 UAUCAGCUGUUUGAAUGCACAACCUGUGCAAUGCGAAAAUAUACCGACCAGUGGAGAACGCCUUGAAUUGGCUGUUCACGCCACUC-GCACGCUGGUCACACCGAUUUUGUAAAUGCAA .....((........(((((.....))))).(((((((.......((((((((..(((((((.......))).))))..((.....-)).)))))))).......)))))))...)).. ( -36.24) >DroEre_CAF1 4080 118 - 1 UAUCAGCUGUUUGGAUGCUCAGUCUGUGCAACGCGAAAAUAUACUGACAAGUGGAGAACGCCUCAAAUUGCCUGUUUUCGCCACUC-GCACCCUGGUCACACUGAUUUUAUAAAUGCAA .(((((.(((...(((...(((..((((....(((((((((((((....))))(((.....)))........)))))))))....)-)))..))))))))))))))............. ( -26.60) >DroYak_CAF1 2627 119 - 1 UAUCAGCUGUUUGGGUGAACAGCCUGUGCCACGCAAAAAUAUACUGACCAUUGGUGAACGCCUCAAAUUGGCUGUUUACGCCAUUUGGCACCCUGGCCACACUGAUUUUAUAAAUGCAA .(((((.(((.(((((((((((((((((...)))).................(((....))).......)))))))))).)))...(((......)))))))))))............. ( -33.50) >consensus UAUCAGCUGUUUGAAUGCACAACCUGUGCAAUGCGAAAAUAUACCGACCAGUGGAGAACGCCUCGAAUUGGCUGUUCACGCCACUC_GCACGCUGGUCACACCGAUUUUGUAAAUGCAA .....((..(((((.(((((.....))))).).))))........((((((((..(((((((.......))).))))..((......)).)))))))).................)).. (-24.42 = -25.02 + 0.60)

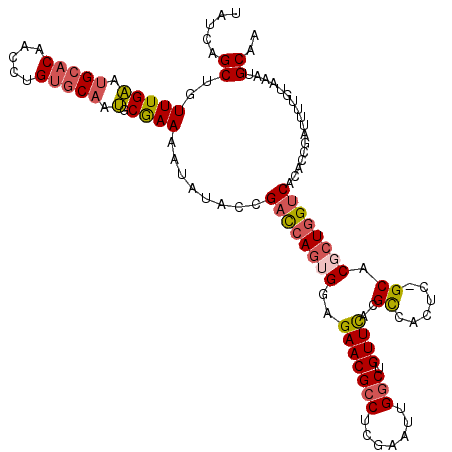

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:04:54 2006