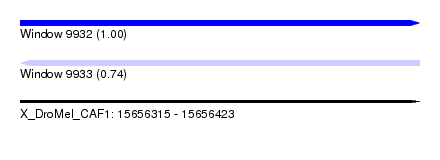

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,656,315 – 15,656,423 |
| Length | 108 |
| Max. P | 0.999345 |

| Location | 15,656,315 – 15,656,423 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 72.51 |
| Mean single sequence MFE | -26.80 |
| Consensus MFE | -16.49 |
| Energy contribution | -18.17 |
| Covariance contribution | 1.68 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.62 |
| SVM decision value | 3.53 |
| SVM RNA-class probability | 0.999345 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
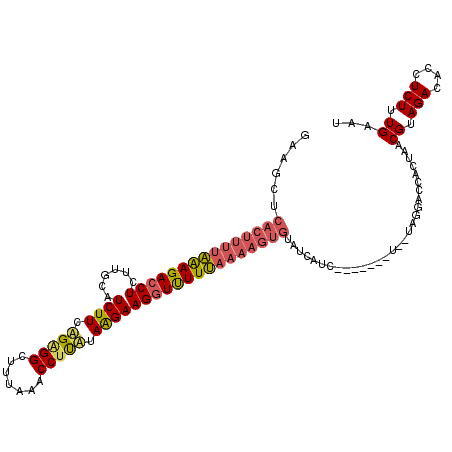
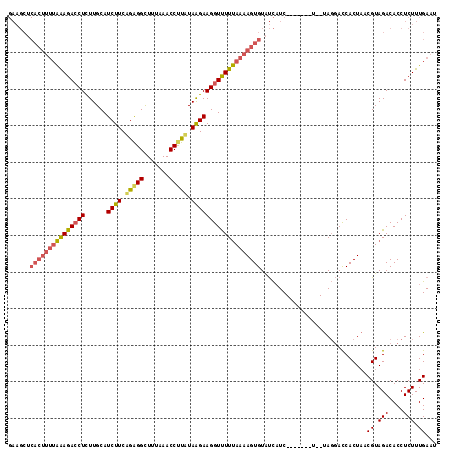
>X_DroMel_CAF1 15656315 108 + 22224390 GAAGCUCACUUUUAGAGACCUUUUGCAUCUUCUGAGGCUUUAAACCUUAUAAGAAGGUUUUUAAAAGUGUAUCAUC-------U--UAGGACCACUAACGUAGACACCUCUUUGAAU ......(((((((((((((((......((((.(((((.......))))).)))))))))))))))))))..(((..-------.--.(((.(.((....)).)...)))...))).. ( -27.00) >DroSec_CAF1 12817 108 + 1 GAAGCUCACUUUUAAAGACCUCUUGCAUCUUCAGGGGCUUUAAACCUCAUAGGAAGGUCUUUAAAAGUGUAUCAUC-------U--UAGGACCACUAACGUAGACACCUCUUUGAAU ......(((((((((((((((......((((..((((.......))))..)))))))))))))))))))..(((..-------.--.(((.(.((....)).)...)))...))).. ( -27.80) >DroEre_CAF1 12449 102 + 1 -------------GAAGAGCUCUUGCAUCUUCAGAGGUUUGAAACCCUUUAAGAAGGUUUUCAACAGUGUGCUUUCGAUAAUUUGCCAUCAUCACU--CGUAGACACCUCUUUGAAU -------------(((((((....)).)))))((((((.(((((.((((....)))).)))))((((((((....(((....)))....)).))))--.))....))))))...... ( -25.60) >consensus GAAGCUCACUUUUAAAGACCUCUUGCAUCUUCAGAGGCUUUAAACCUUAUAAGAAGGUUUUUAAAAGUGUAUCAUC_______U__UAGGACCACUAACGUAGACACCUCUUUGAAU ......(((((((((((((((......((((.(((((.......))))).))))))))))))))))))).............................((.(((....))).))... (-16.49 = -18.17 + 1.68)

| Location | 15,656,315 – 15,656,423 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 72.51 |
| Mean single sequence MFE | -27.13 |
| Consensus MFE | -14.43 |
| Energy contribution | -14.67 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.737844 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
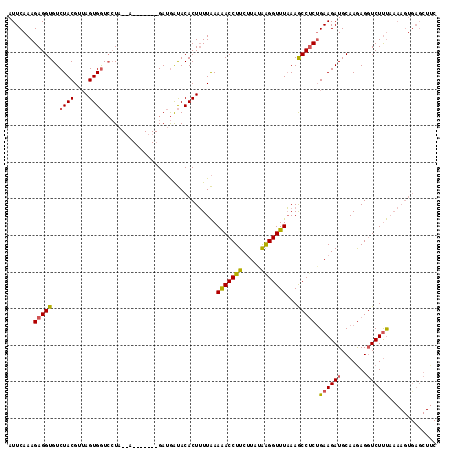
>X_DroMel_CAF1 15656315 108 - 22224390 AUUCAAAGAGGUGUCUACGUUAGUGGUCCUA--A-------GAUGAUACACUUUUAAAAACCUUCUUAUAAGGUUUAAAGCCUCAGAAGAUGCAAAAGGUCUCUAAAAGUGAGCUUC ..(((...(((...((((....)))).))).--.-------..)))..((((((((.(.((((((((...((((.....))))...))))......)))).).))))))))...... ( -24.60) >DroSec_CAF1 12817 108 - 1 AUUCAAAGAGGUGUCUACGUUAGUGGUCCUA--A-------GAUGAUACACUUUUAAAGACCUUCCUAUGAGGUUUAAAGCCCCUGAAGAUGCAAGAGGUCUUUAAAAGUGAGCUUC ..(((...(((...((((....)))).))).--.-------..)))..((((((((((((((((((.....))......((..(....)..))..))))))))))))))))...... ( -30.70) >DroEre_CAF1 12449 102 - 1 AUUCAAAGAGGUGUCUACG--AGUGAUGAUGGCAAAUUAUCGAAAGCACACUGUUGAAAACCUUCUUAAAGGGUUUCAAACCUCUGAAGAUGCAAGAGCUCUUC------------- ......((((((((((...--..(((((((.....)))))))..)).))....((((((.((((....)))).))))))))))))(((((.((....)))))))------------- ( -26.10) >consensus AUUCAAAGAGGUGUCUACGUUAGUGGUCCUA__A_______GAUGAUACACUUUUAAAAACCUUCUUAUAAGGUUUAAAGCCUCUGAAGAUGCAAGAGGUCUUUAAAAGUGAGCUUC .......(((((..((((....))))...............................(((((((.....)))))))...))))).((((((.......))))))............. (-14.43 = -14.67 + 0.23)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:04:29 2006