
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,645,752 – 15,645,845 |
| Length | 93 |
| Max. P | 0.992824 |

| Location | 15,645,752 – 15,645,845 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 71.90 |
| Mean single sequence MFE | -19.84 |
| Consensus MFE | -10.43 |
| Energy contribution | -10.76 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.589095 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15645752 93 + 22224390 UUUGUAUUUUAAACUGGCAGAGCGACGCUCGCUCGGUCGCU--CUCUCUCUCUGGCGCUCUCACGCUCUUGCUCUGCUUCUAGCCCCACAAUUCU .((((..........((((((((((.((......((.((((--..........)))))).....))..)))))))))).........)))).... ( -21.41) >DroEre_CAF1 2171 85 + 1 UAUGUAUUUUACACUGGGAGAGCGGCGCUCUGUCGCUCUCUUG----CACUCUCGCGCUCUCACGCU-UG-----UCUUCUAGCCCCACAAUUCU .............(((((((((((((.....)))))))))...----.......(((......))).-..-----....))))............ ( -21.40) >DroYak_CAF1 2609 70 + 1 UCUGUAUUUUACACUGGGAGAGCGACGCUCUU--------------------CCGCGCUCUCACGCUCUG-----UCUUCUAGCCCCACAAUUCU ..............((((((((((.((.....--------------------.)))))))))..(((..(-----....).))).)))....... ( -16.70) >consensus UAUGUAUUUUACACUGGGAGAGCGACGCUCUCUCG_UC_CU______C_CUCUCGCGCUCUCACGCUCUG_____UCUUCUAGCCCCACAAUUCU ..............((((((((((.(............................))))))))..(((..............))).)))....... (-10.43 = -10.76 + 0.33)


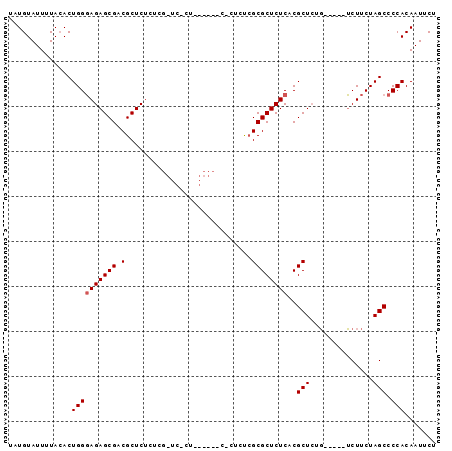
| Location | 15,645,752 – 15,645,845 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 71.90 |
| Mean single sequence MFE | -24.30 |
| Consensus MFE | -15.50 |
| Energy contribution | -15.83 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.64 |
| SVM decision value | 2.35 |
| SVM RNA-class probability | 0.992824 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15645752 93 - 22224390 AGAAUUGUGGGGCUAGAAGCAGAGCAAGAGCGUGAGAGCGCCAGAGAGAGAG--AGCGACCGAGCGAGCGUCGCUCUGCCAGUUUAAAAUACAAA ....(((((....((((.(((((((..(.((((....)))))...((..(..--.((......))...).)))))))))...))))...))))). ( -28.10) >DroEre_CAF1 2171 85 - 1 AGAAUUGUGGGGCUAGAAGA-----CA-AGCGUGAGAGCGCGAGAGUG----CAAGAGAGCGACAGAGCGCCGCUCUCCCAGUGUAAAAUACAUA .......(((.(((......-----..-)))......((((....)))----)..(((((((.(.....).))))))))))(((((...))))). ( -25.00) >DroYak_CAF1 2609 70 - 1 AGAAUUGUGGGGCUAGAAGA-----CAGAGCGUGAGAGCGCGG--------------------AAGAGCGUCGCUCUCCCAGUGUAAAAUACAGA ....(((((..((.......-----....((((....))))((--------------------.(((((...))))).))...))....))))). ( -19.80) >consensus AGAAUUGUGGGGCUAGAAGA_____CAGAGCGUGAGAGCGCGAGAG_G______AG_GA_CGAAAGAGCGUCGCUCUCCCAGUGUAAAAUACAAA .......(((((.................((((....))))........................((((...))))))))).............. (-15.50 = -15.83 + 0.33)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:04:13 2006