
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,620,557 – 15,620,712 |
| Length | 155 |
| Max. P | 0.991523 |

| Location | 15,620,557 – 15,620,675 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.68 |
| Mean single sequence MFE | -43.58 |
| Consensus MFE | -30.10 |
| Energy contribution | -32.34 |
| Covariance contribution | 2.24 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.82 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.59 |
| SVM RNA-class probability | 0.966319 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15620557 118 + 22224390 CUACUACAAGUGGGUAC--AACGAGGUCCGCUUUCAACGAACCAAGUUCUCCGGCGAGUUUGUGGACAGUAGUUGGUCCAACUCACUUAAGAACUCGGAUUGGUUGGAGUGGUCCCAGUU ((((.....))))....--(((..((.((((((.((((.((((.((((((..((.((((...(((((........))))))))).))..)))))).)).)).)))))))))).))..))) ( -43.60) >DroSec_CAF1 450 118 + 1 CUACUACAAGUGGGUAC--AACGAGGUCGUCUUCCAACGAACCAAGUUCUCCGGCGAGUUUGCGGACAGUAGCUUGUCCAACUCACUUAAGAACUCGGGUUGGUUGGAGUGGCCCUAGUU ((((.....))))....--(((..(((((.((.(((((.((((.((((((..((.(((((...((((((....))))))))))).))..))))))..)))).))))))))))))...))) ( -45.10) >DroSim_CAF1 450 118 + 1 CUACUACAAGUGGGUAC--AACGAGGUCCGCUUCCAACGAACCAAGUUCUCCGGCGAGUUUGCGGACAGUCGCUUGUCCAACUCACCUAAGAACUCGGAUUGGUUGGAGUGGCCCUAGUU ((((.....))))....--(((.(((.(((((.(((((.((((.((((((..((.(((((...((((((....))))))))))).))..)))))).)).)).)))))))))).))).))) ( -49.40) >DroEre_CAF1 439 118 + 1 CUACUACAAGUGGGUAA--AAUGAGGCCCCCUUUCAACAAACCGAGUUCUCAUGCGAGUUUGUGGGCAGUGACUUCGCAAACUCACUUAAGAACUCUGAUUGGUUGGAGUGGCUCGAGUU ...((((.((.((((..--......)))).))((((((...(.(((((((...(.(((((((((((.......))))))))))).)...))))))).)....))))))))))........ ( -36.90) >DroYak_CAF1 438 120 + 1 CUACUACAAGUGGGUAAAAAAUGAGGCCCACUCUCAUCGAACCGAGUUCUCAUGCGUGUUGGUGGGCAGAAGCUUCCCAAACUCACUUAAGAACUCUAAUUGGUUGAAGUGGGCCGAGAU ((((.....))))...........((((((((.(((((((...(((((((...(.(..((((..(((....)))..))))...).)...)))))))...)))).)))))))))))..... ( -42.90) >consensus CUACUACAAGUGGGUAC__AACGAGGUCCGCUUUCAACGAACCAAGUUCUCCGGCGAGUUUGUGGACAGUAGCUUGUCCAACUCACUUAAGAACUCGGAUUGGUUGGAGUGGCCCGAGUU ((((.....))))...........((.(((((.(((((.((((.((((((..((.(((((...((((((....))))))))))).))..)))))).)).)).)))))))))).))..... (-30.10 = -32.34 + 2.24)



| Location | 15,620,557 – 15,620,675 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.68 |
| Mean single sequence MFE | -43.66 |
| Consensus MFE | -30.28 |
| Energy contribution | -31.32 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.20 |
| Mean z-score | -3.40 |
| Structure conservation index | 0.69 |
| SVM decision value | 2.27 |
| SVM RNA-class probability | 0.991523 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15620557 118 - 22224390 AACUGGGACCACUCCAACCAAUCCGAGUUCUUAAGUGAGUUGGACCAACUACUGUCCACAAACUCGCCGGAGAACUUGGUUCGUUGAAAGCGGACCUCGUU--GUACCCACUUGUAGUAG (((.(((.((.((.((((.((.((((((((((..(((((((((((........)))))...))))))..)))))))))))).))))..)).)).))).)))--................. ( -40.60) >DroSec_CAF1 450 118 - 1 AACUAGGGCCACUCCAACCAACCCGAGUUCUUAAGUGAGUUGGACAAGCUACUGUCCGCAAACUCGCCGGAGAACUUGGUUCGUUGGAAGACGACCUCGUU--GUACCCACUUGUAGUAG .....(((..((((((((.((((.((((((((..((((((((((((......)))))...)))))))..)))))))))))).)))))).((((....))))--)).)))........... ( -46.20) >DroSim_CAF1 450 118 - 1 AACUAGGGCCACUCCAACCAAUCCGAGUUCUUAGGUGAGUUGGACAAGCGACUGUCCGCAAACUCGCCGGAGAACUUGGUUCGUUGGAAGCGGACCUCGUU--GUACCCACUUGUAGUAG (((.(((.((.(((((((.((.((((((((((.(((((((((((((......)))))...)))))))).)))))))))))).))))).)).)).))).)))--................. ( -50.40) >DroEre_CAF1 439 118 - 1 AACUCGAGCCACUCCAACCAAUCAGAGUUCUUAAGUGAGUUUGCGAAGUCACUGCCCACAAACUCGCAUGAGAACUCGGUUUGUUGAAAGGGGGCCUCAUU--UUACCCACUUGUAGUAG .(((((((......((((.((((.(((((((((.(((((((((.(...........).))))))))).))))))))))))).))))...(((.........--...))).)))).))).. ( -39.70) >DroYak_CAF1 438 120 - 1 AUCUCGGCCCACUUCAACCAAUUAGAGUUCUUAAGUGAGUUUGGGAAGCUUCUGCCCACCAACACGCAUGAGAACUCGGUUCGAUGAGAGUGGGCCUCAUUUUUUACCCACUUGUAGUAG .....(((((((((((.(.((((.(((((((((.(((.(((.((...((....))...))))).))).))))))))))))).).))).))))))))........................ ( -41.40) >consensus AACUAGGGCCACUCCAACCAAUCCGAGUUCUUAAGUGAGUUGGACAAGCUACUGUCCACAAACUCGCCGGAGAACUUGGUUCGUUGAAAGCGGACCUCGUU__GUACCCACUUGUAGUAG .....((.((.(((((((.((((.((((((((..((((((((((((......))))))...))))))..)))))))))))).)))))).).)).))........................ (-30.28 = -31.32 + 1.04)



| Location | 15,620,595 – 15,620,712 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 84.73 |
| Mean single sequence MFE | -32.30 |
| Consensus MFE | -21.46 |
| Energy contribution | -22.38 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.735477 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15620595 117 - 22224390 GAGAAUGAAUACAUUAAUAUCAGACGGUCGAACUUGUAACUGGGACCACUCCAACCAAUCCGAGUUCUUAAGUGAGUUGGACCAACUACUGUCCACAAACUCGCCGGAGAACUUGGU (((((((....))))..........((((...(........).)))).)))........((((((((((..(((((((((((........)))))...))))))..)))))))))). ( -35.60) >DroSec_CAF1 488 113 - 1 GAGAA----UACAUUAAUAUCAGACGACCGAACUUGUAACUAGGGCCACUCCAACCAACCCGAGUUCUUAAGUGAGUUGGACAAGCUACUGUCCGCAAACUCGCCGGAGAACUUGGU (((..----..........((....))..(..((((....))))..).)))........((((((((((..((((((((((((......)))))...)))))))..)))))))))). ( -32.00) >DroSim_CAF1 488 113 - 1 GAGAA----UAAAUUAAUAUCAGACGACCCAACUUGUAACUAGGGCCACUCCAACCAAUCCGAGUUCUUAGGUGAGUUGGACAAGCGACUGUCCGCAAACUCGCCGGAGAACUUGGU (((..----..........((....))(((............)))...)))........((((((((((.(((((((((((((......)))))...)))))))).)))))))))). ( -36.00) >DroEre_CAF1 477 113 - 1 GAGAA----UACAUUAAUACCAGACGACCGUACUUGUAACUCGAGCCACUCCAACCAAUCAGAGUUCUUAAGUGAGUUUGCGAAGUCACUGCCCACAAACUCGCAUGAGAACUCGGU .....----.................(((((.((((.....))))..))............(((((((((.(((((((((.(...........).))))))))).)))))))))))) ( -29.70) >DroYak_CAF1 478 113 - 1 GAGAA----UGCAUUAAUACCAGACCACCGAACUUGUAUCUCGGCCCACUUCAACCAAUUAGAGUUCUUAAGUGAGUUUGGGAAGCUUCUGCCCACCAACACGCAUGAGAACUCGGU .....----..............(((.((((.........)))).................(((((((((.(((.(((.((...((....))...))))).))).)))))))))))) ( -28.20) >consensus GAGAA____UACAUUAAUAUCAGACGACCGAACUUGUAACUAGGGCCACUCCAACCAAUCCGAGUUCUUAAGUGAGUUGGACAAGCUACUGUCCACAAACUCGCCGGAGAACUUGGU ...........................................................((((((((((..((((((((((((......))))))...))))))..)))))))))). (-21.46 = -22.38 + 0.92)
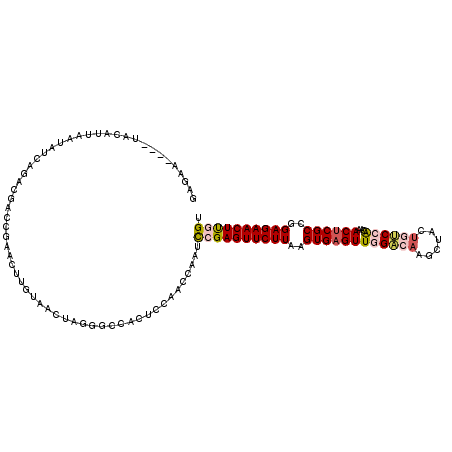


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:03:57 2006