
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,547,648 – 15,547,738 |
| Length | 90 |
| Max. P | 0.932194 |

| Location | 15,547,648 – 15,547,738 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 88.18 |
| Mean single sequence MFE | -21.16 |
| Consensus MFE | -15.07 |
| Energy contribution | -17.32 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.793869 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15547648 90 + 22224390 AAAAAGAGACUGAGAAUAUU--CGAGGAAACGAGCGCGACGGAGAACCUUAAAAUCAGAAUUGAAACUCUCAAUCGACGCGUAGCAAACGCA .......((.(((((...((--((((....)((.......((....))......))....)))))..))))).))...((((.....)))). ( -16.82) >DroSec_CAF1 2621 92 + 1 AAAAAGAGACUGACCGUUUCCGCGAGGAAACGAACGCGGCGGAGAACCUUAAAAUAAGAAUUGAAACUCUCAAUCGAAGCGUAGCAAACGCA ..............(((((((....))))))).((((..((((((...((((........))))...))))...))..)))).((....)). ( -23.40) >DroSim_CAF1 2629 92 + 1 AAAAAGAGACUGAGCGUUUCCGCGAGGAAACGAACGCGGCGGAGAACCUUAAAAUAAGAAUUGAAACUCUCAAUCGAAGCGUAGCAAACGCA .......((.((((.(((((((((.(....)...))))..((....))..............))))).)))).))...((((.....)))). ( -24.90) >DroEre_CAF1 4451 90 + 1 AAAACGAGACUGAGAAUGUU--CAAGAGAACGAGCGCGACGGAGAACCCAAAAAUUAGAAUUGAAAUUCUCAAUCGAAGCGUAGCAAACGCA ....(((...(((((((.((--(((.....((....))..((.....))...........)))))))))))).)))..((((.....)))). ( -19.50) >consensus AAAAAGAGACUGAGAAUUUC__CGAGGAAACGAACGCGACGGAGAACCUUAAAAUAAGAAUUGAAACUCUCAAUCGAAGCGUAGCAAACGCA .......((.((((((((((((((.(....)...))))..((....))..............)))))))))).))...((((.....)))). (-15.07 = -17.32 + 2.25)

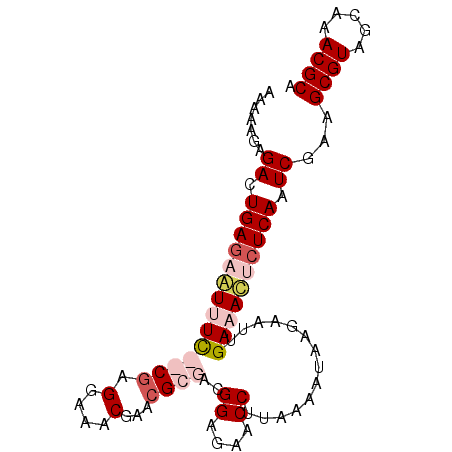

| Location | 15,547,648 – 15,547,738 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 88.18 |
| Mean single sequence MFE | -23.28 |
| Consensus MFE | -18.00 |
| Energy contribution | -19.12 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.22 |
| SVM RNA-class probability | 0.932194 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 15547648 90 - 22224390 UGCGUUUGCUACGCGUCGAUUGAGAGUUUCAAUUCUGAUUUUAAGGUUCUCCGUCGCGCUCGUUUCCUCG--AAUAUUCUCAGUCUCUUUUU .((((.....))))...(((((((((((((..............((....))(..((....))..)...)--)).))))))))))....... ( -19.60) >DroSec_CAF1 2621 92 - 1 UGCGUUUGCUACGCUUCGAUUGAGAGUUUCAAUUCUUAUUUUAAGGUUCUCCGCCGCGUUCGUUUCCUCGCGGAAACGGUCAGUCUCUUUUU .((((.....))))...((((((..(((((..............((....))..((((..........)))))))))..))))))....... ( -23.90) >DroSim_CAF1 2629 92 - 1 UGCGUUUGCUACGCUUCGAUUGAGAGUUUCAAUUCUUAUUUUAAGGUUCUCCGCCGCGUUCGUUUCCUCGCGGAAACGCUCAGUCUCUUUUU .((((.....))))...(((((((.(((((..............((....))..((((..........))))))))).)))))))....... ( -26.40) >DroEre_CAF1 4451 90 - 1 UGCGUUUGCUACGCUUCGAUUGAGAAUUUCAAUUCUAAUUUUUGGGUUCUCCGUCGCGCUCGUUCUCUUG--AACAUUCUCAGUCUCGUUUU .((((.....))))...((((((((((................((((.(......).))))((((....)--)))))))))))))....... ( -23.20) >consensus UGCGUUUGCUACGCUUCGAUUGAGAGUUUCAAUUCUUAUUUUAAGGUUCUCCGCCGCGCUCGUUUCCUCG__AAAACGCUCAGUCUCUUUUU .((((.....))))...((((((((((((...............((....)).(((((..........)))))))))))))))))....... (-18.00 = -19.12 + 1.12)

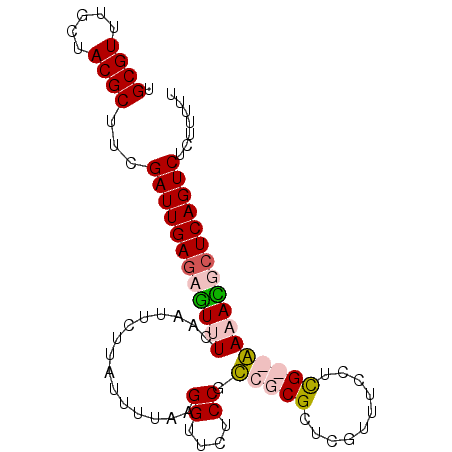

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:03:07 2006